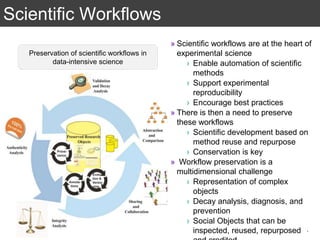
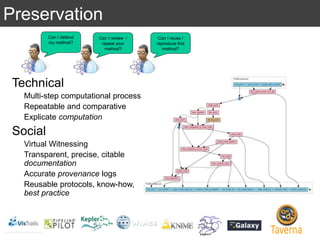
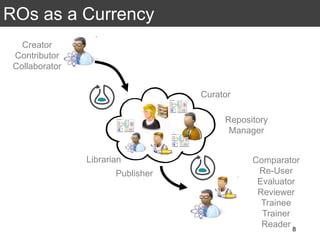
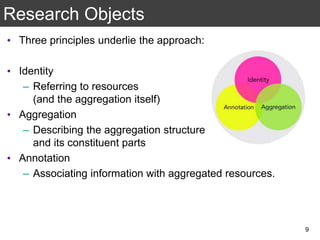
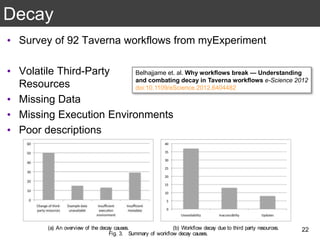
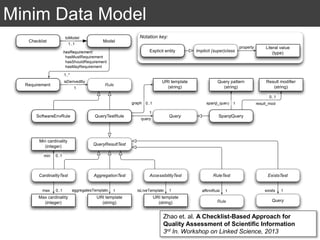
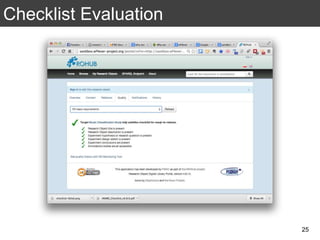
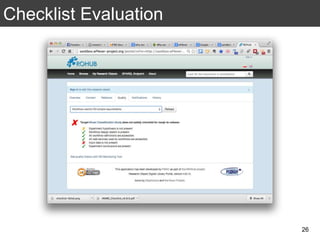
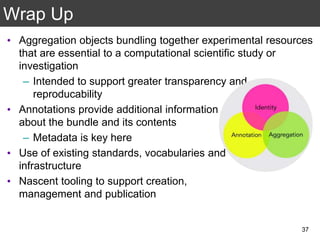
The document discusses the importance of reproducible research in scientific discovery, emphasizing the role of metadata and workflows in enabling transparency and reproducibility. It presents the concept of Research Objects (ROs) as aggregations of essential experimental resources, highlighting the need for proper annotation and identification mechanisms. Additionally, it addresses challenges in preserving scientific workflows and the tools available to manage, publish, and ensure the integrity of research data.