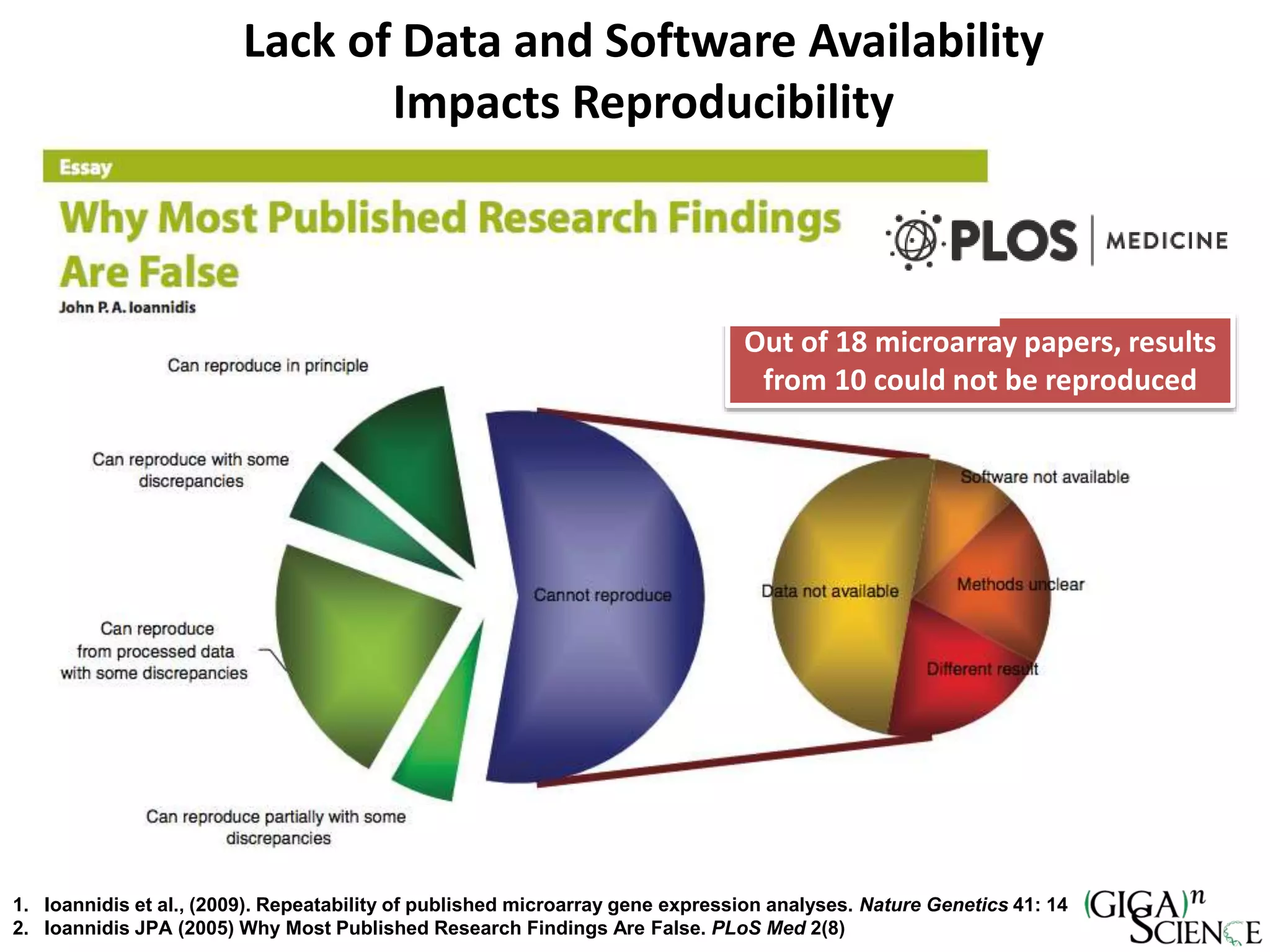
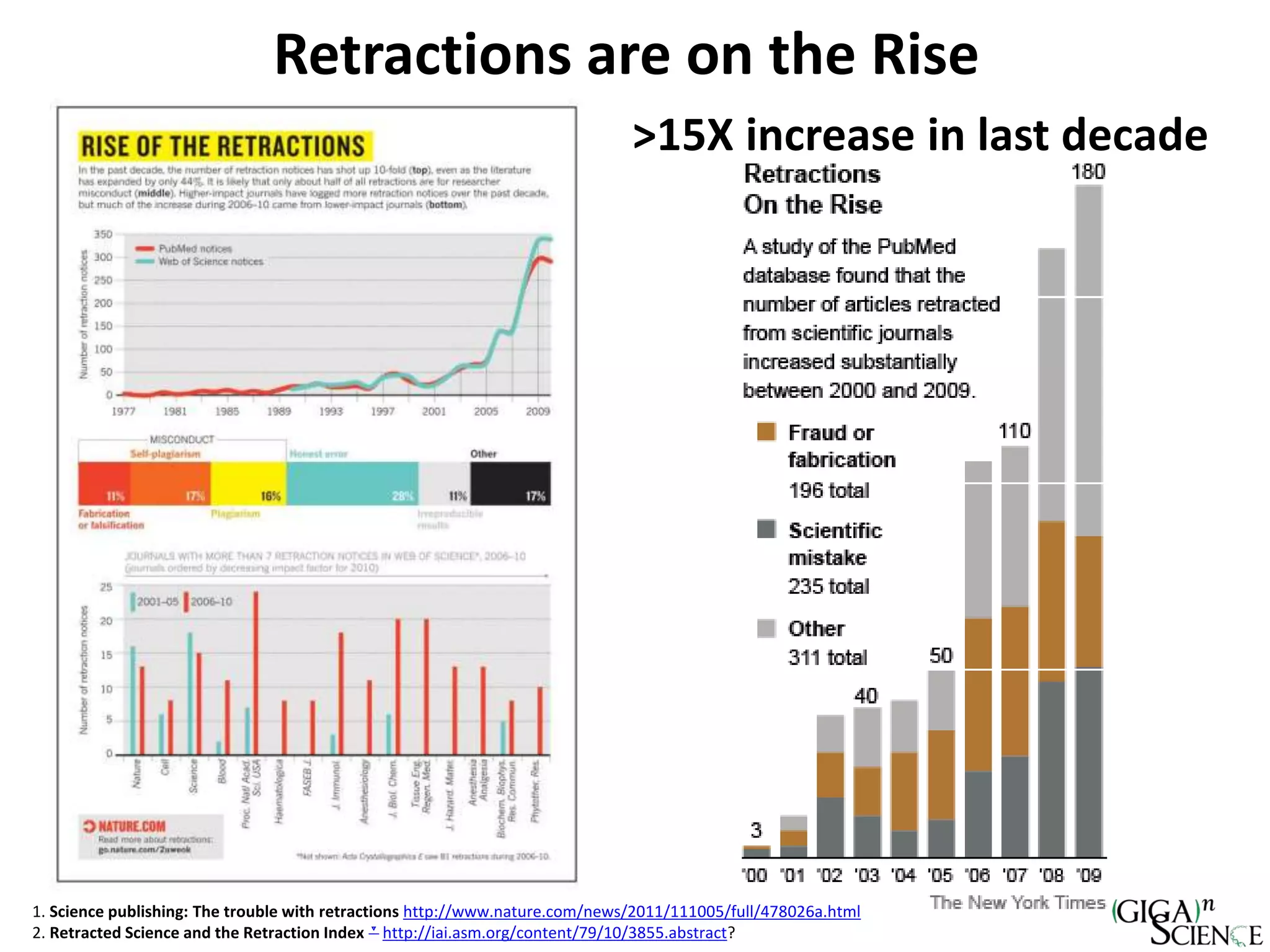
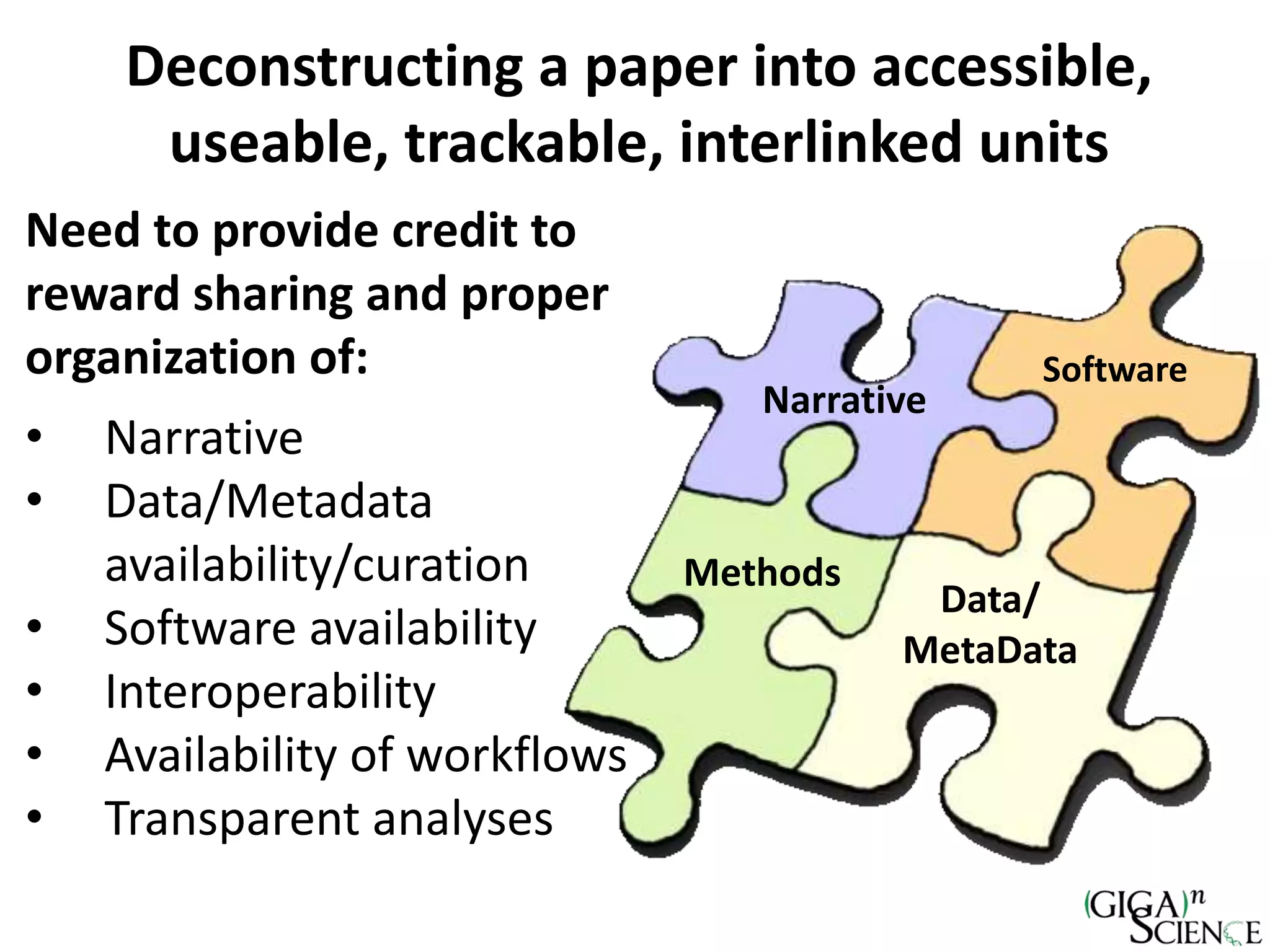
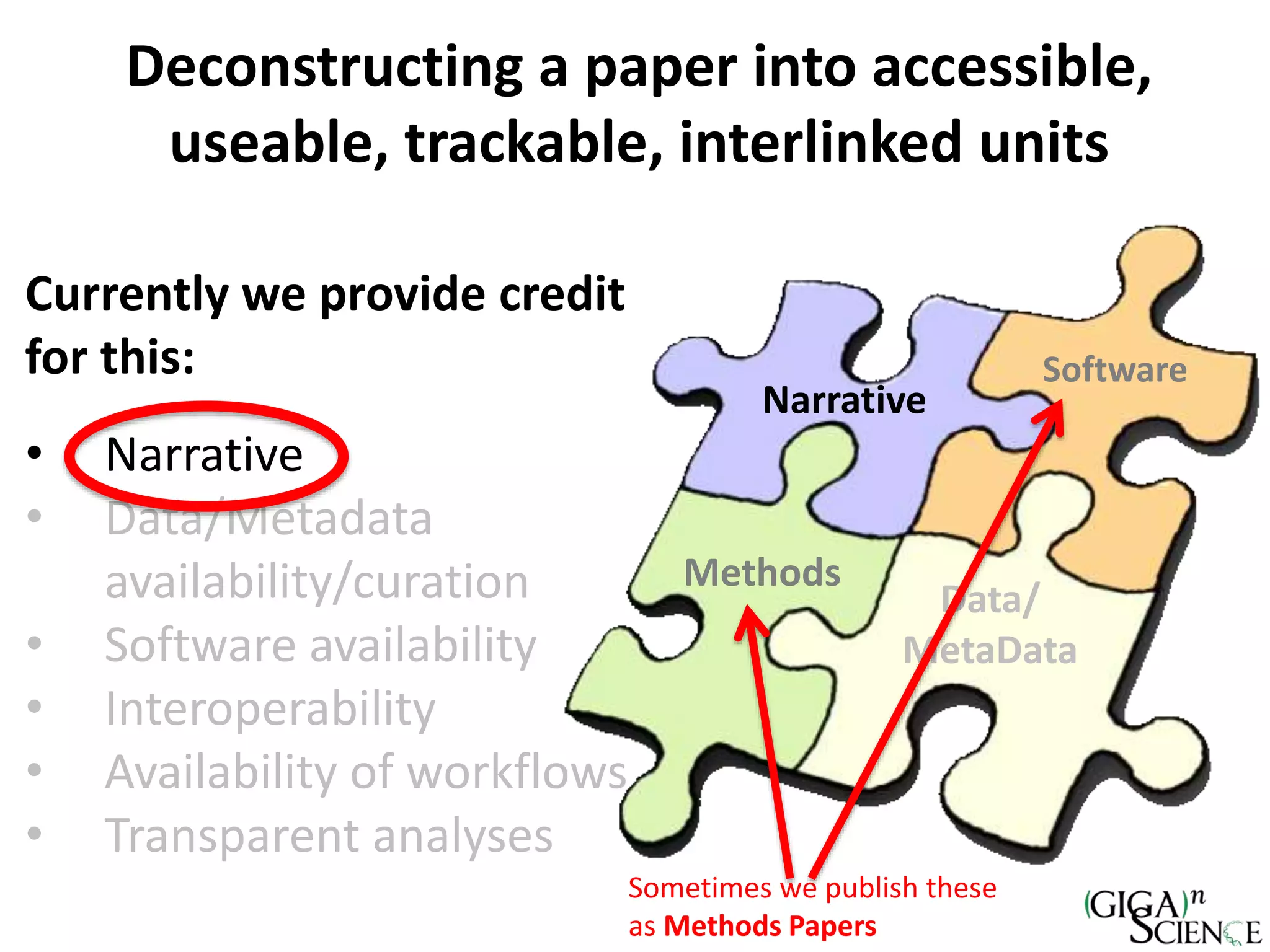
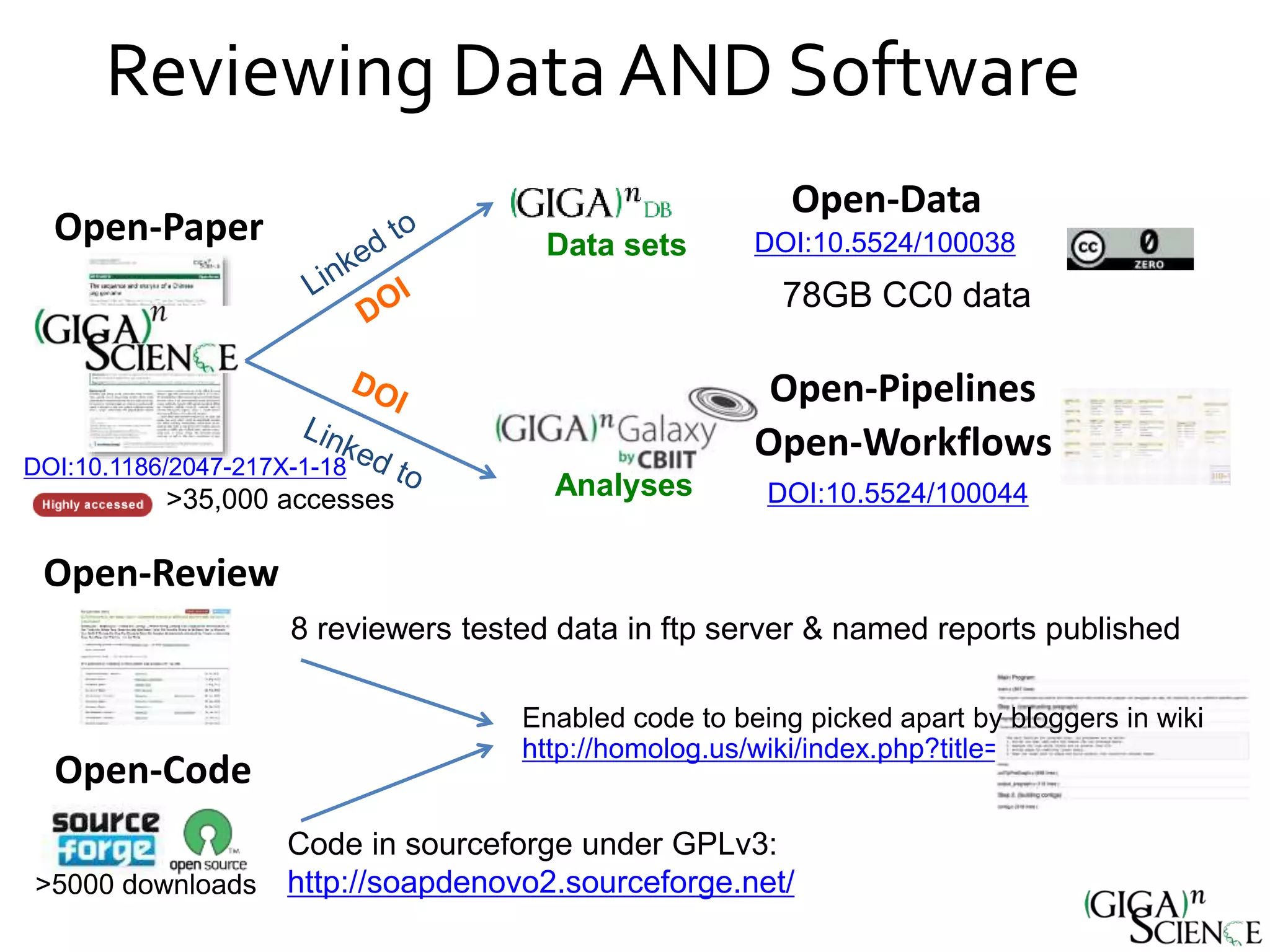
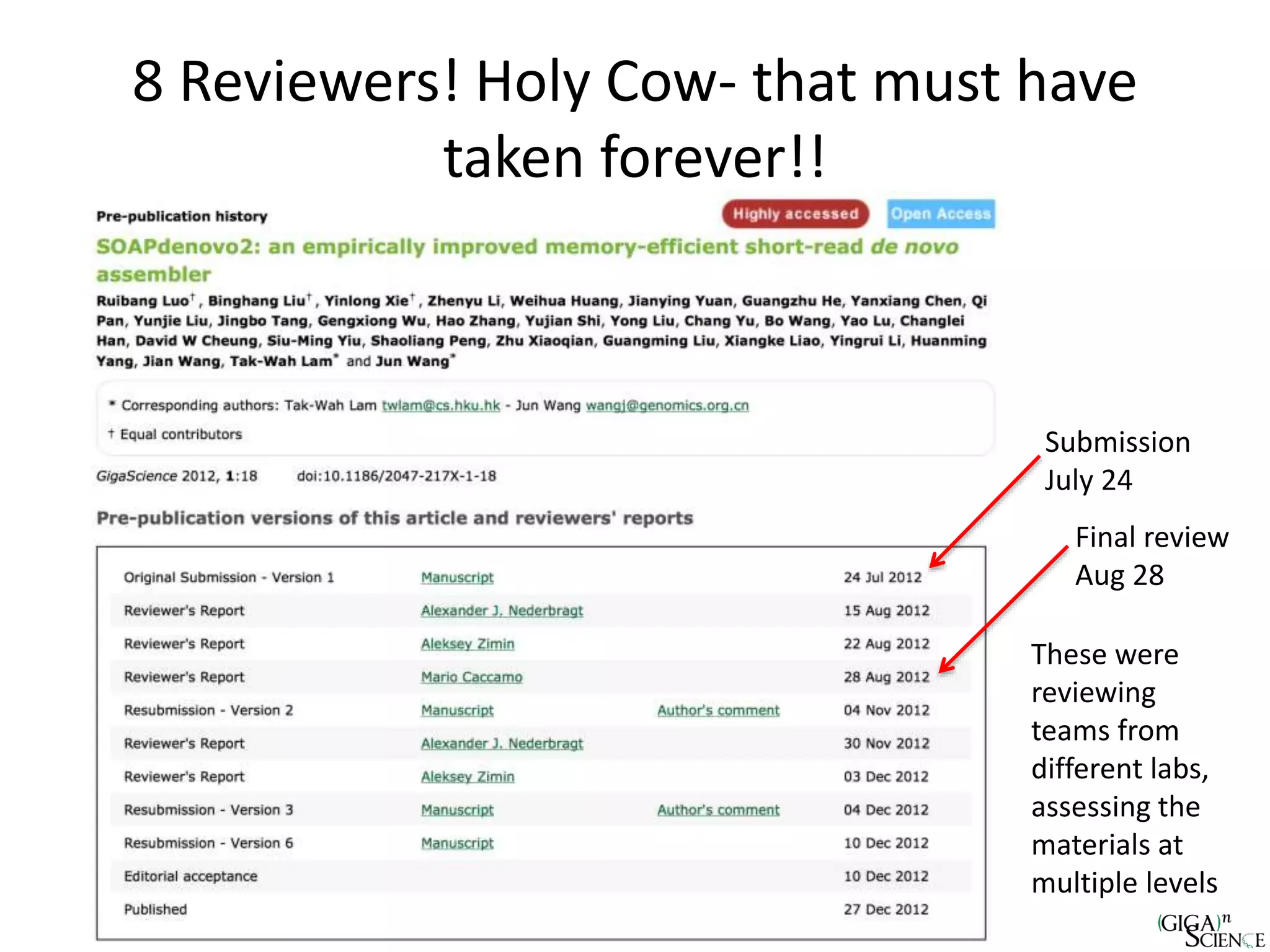
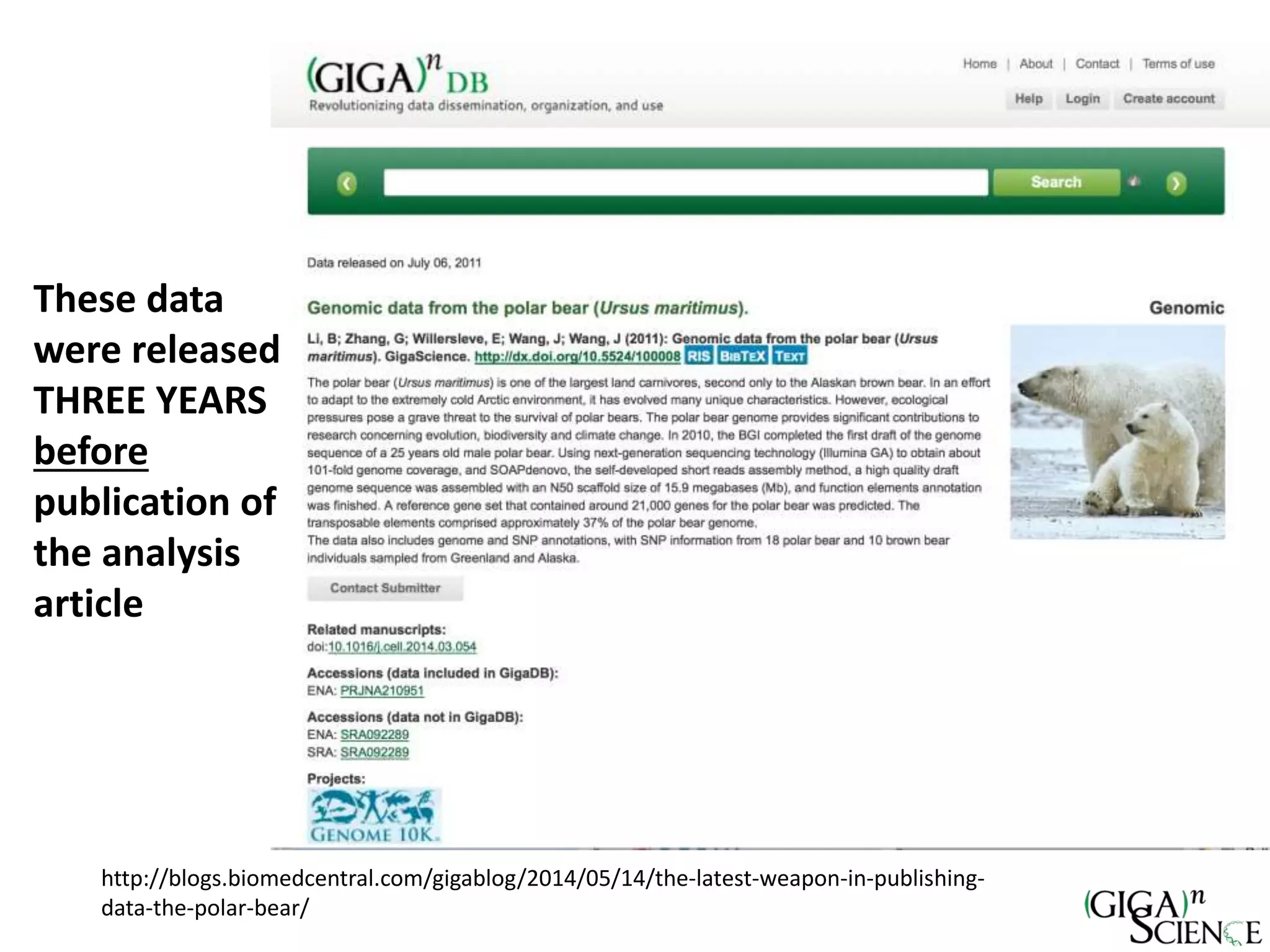
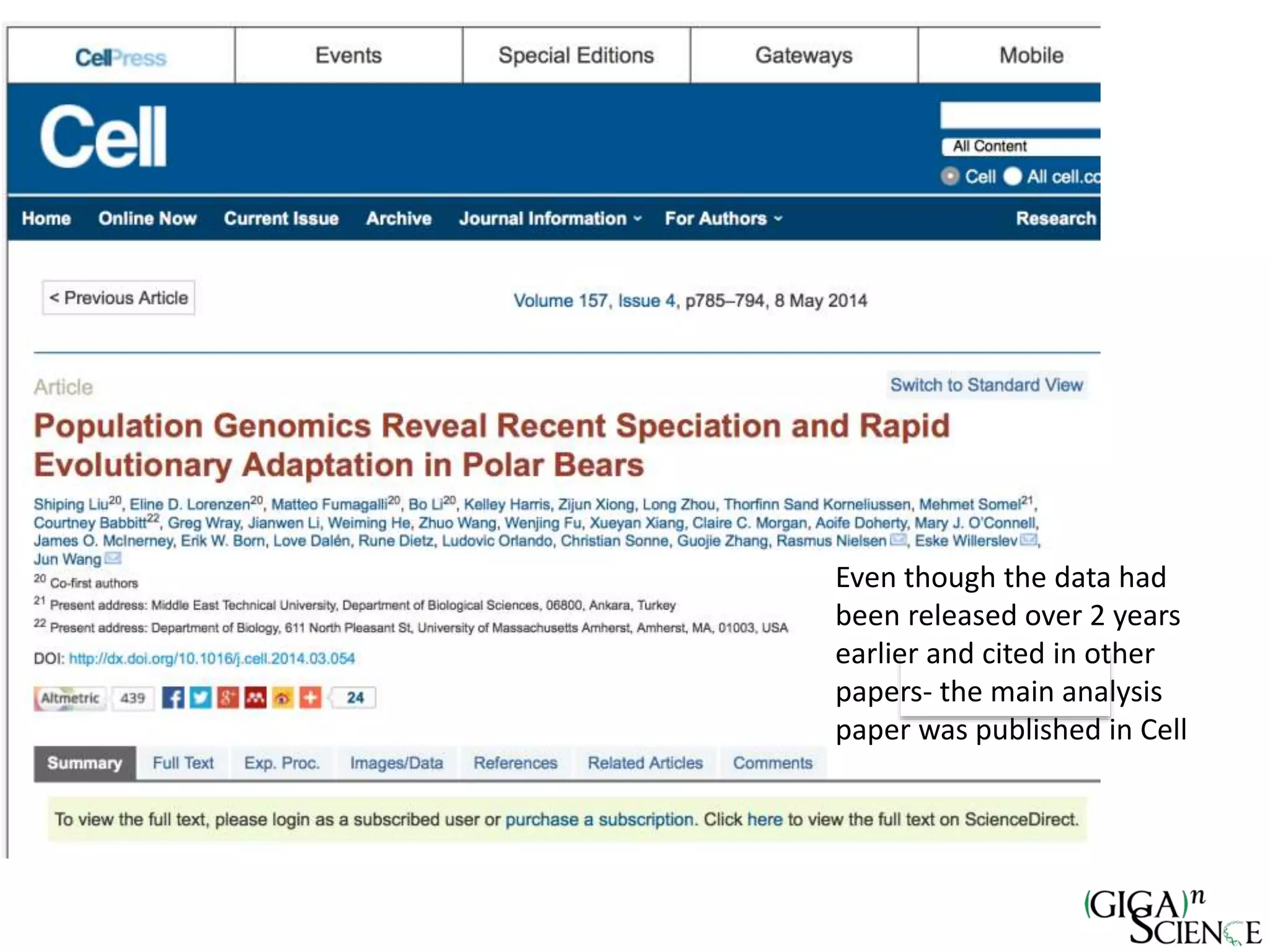
The document discusses the importance of publishing research beyond mere data release mandates, emphasizing the need for accessible and reproducible scientific communication. It outlines challenges such as salami slicing in publications, lack of data availability, and the rising issue of retractions, while promoting practices for proper data citation, software release, and organization in scientific research. The content also highlights initiatives by GigaScience and related repositories to enhance data sharing and interlinking with scholarly articles to improve reproducibility and credit for researchers.