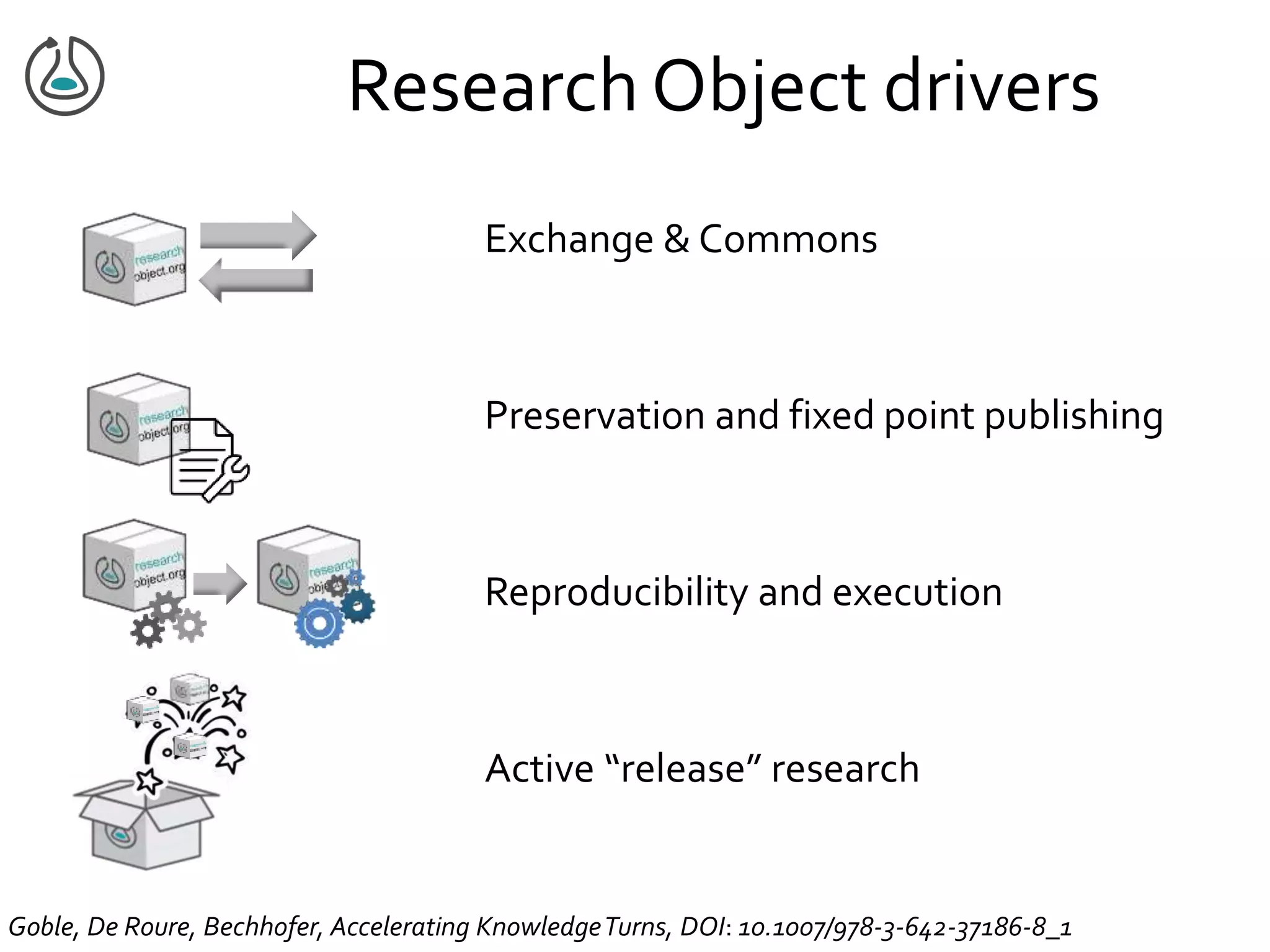
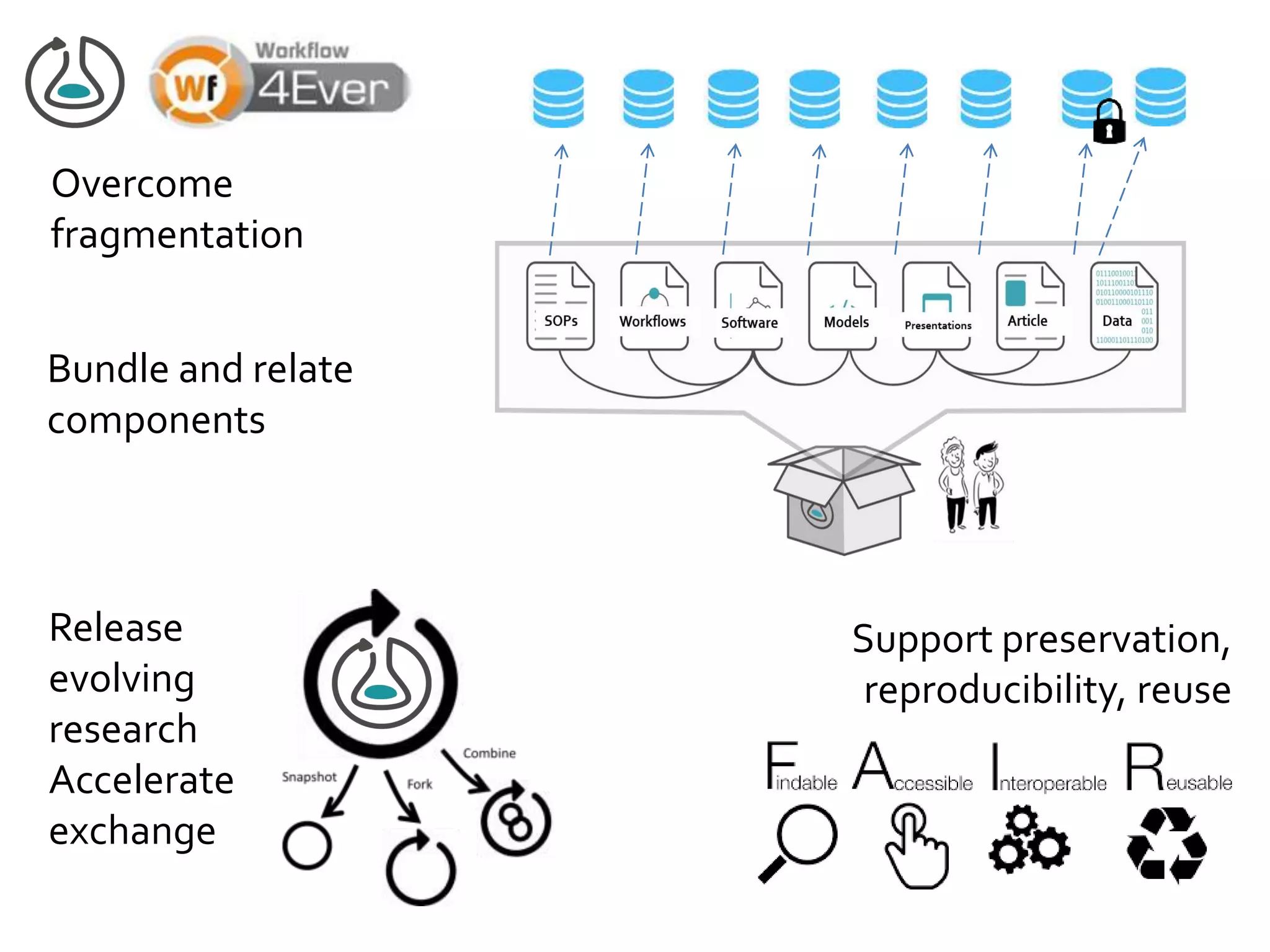
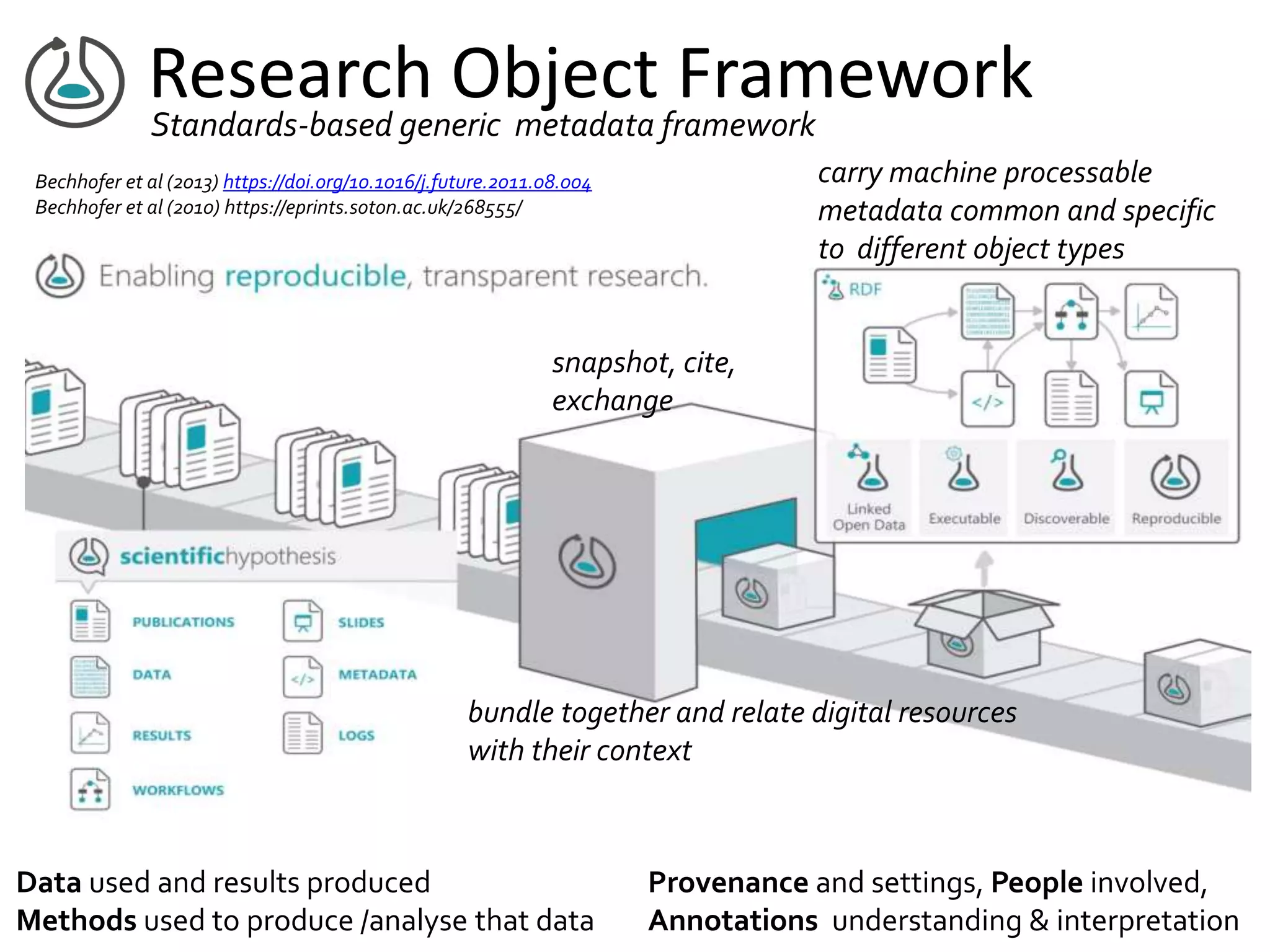
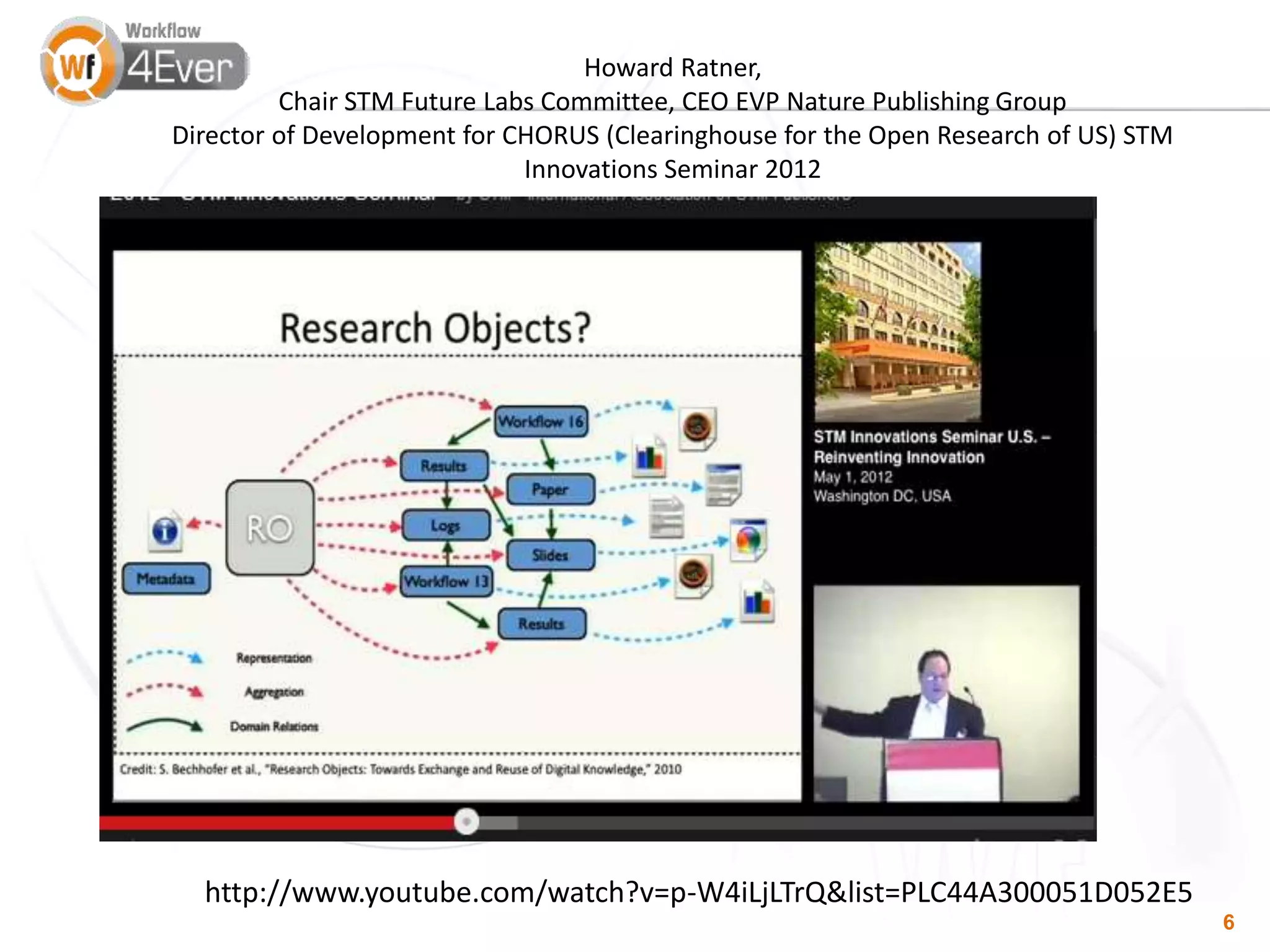
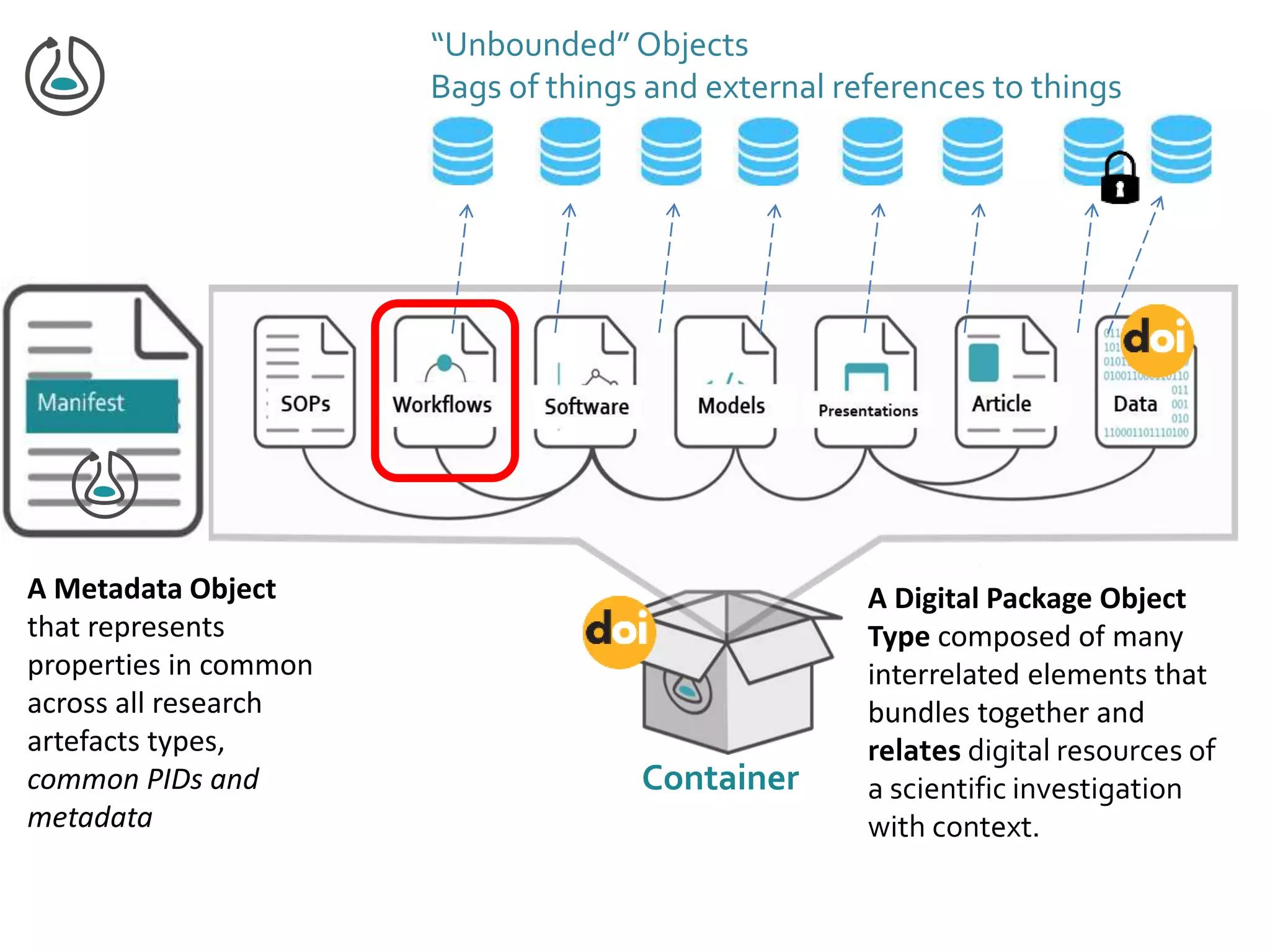
This document summarizes a presentation on Research Objects given on October 29, 2018 in Amsterdam. It discusses how Research Objects can bundle together different components of a research investigation, such as data, methods, provenance, and results, to facilitate exchange, reproducibility, and preservation of research. The Research Object framework carries machine-readable metadata about these components. Examples are given of Research Objects that bundle workflows and computational experiments. Challenges and opportunities are discussed around developing community tools and standards to work with Research Objects.



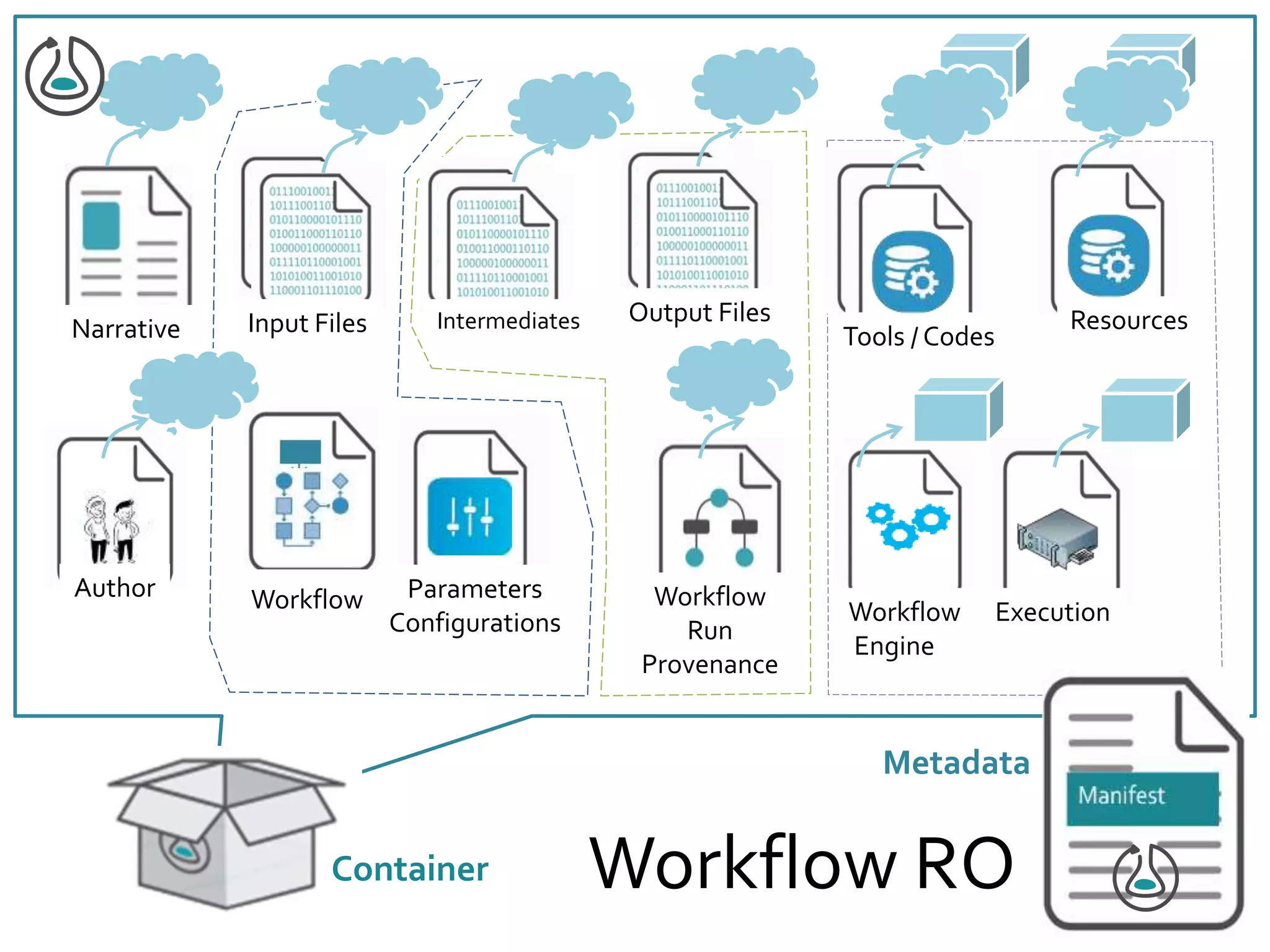
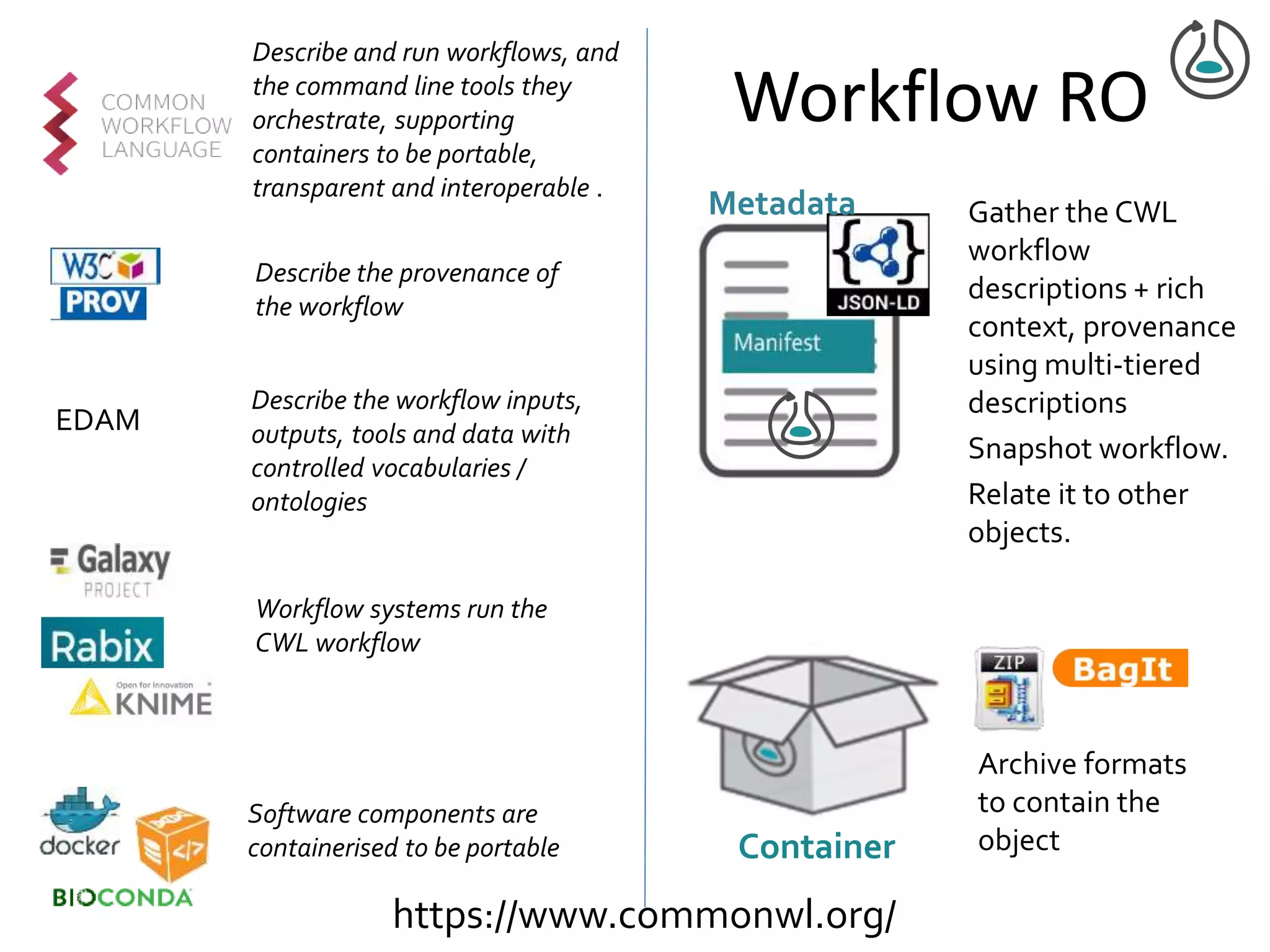
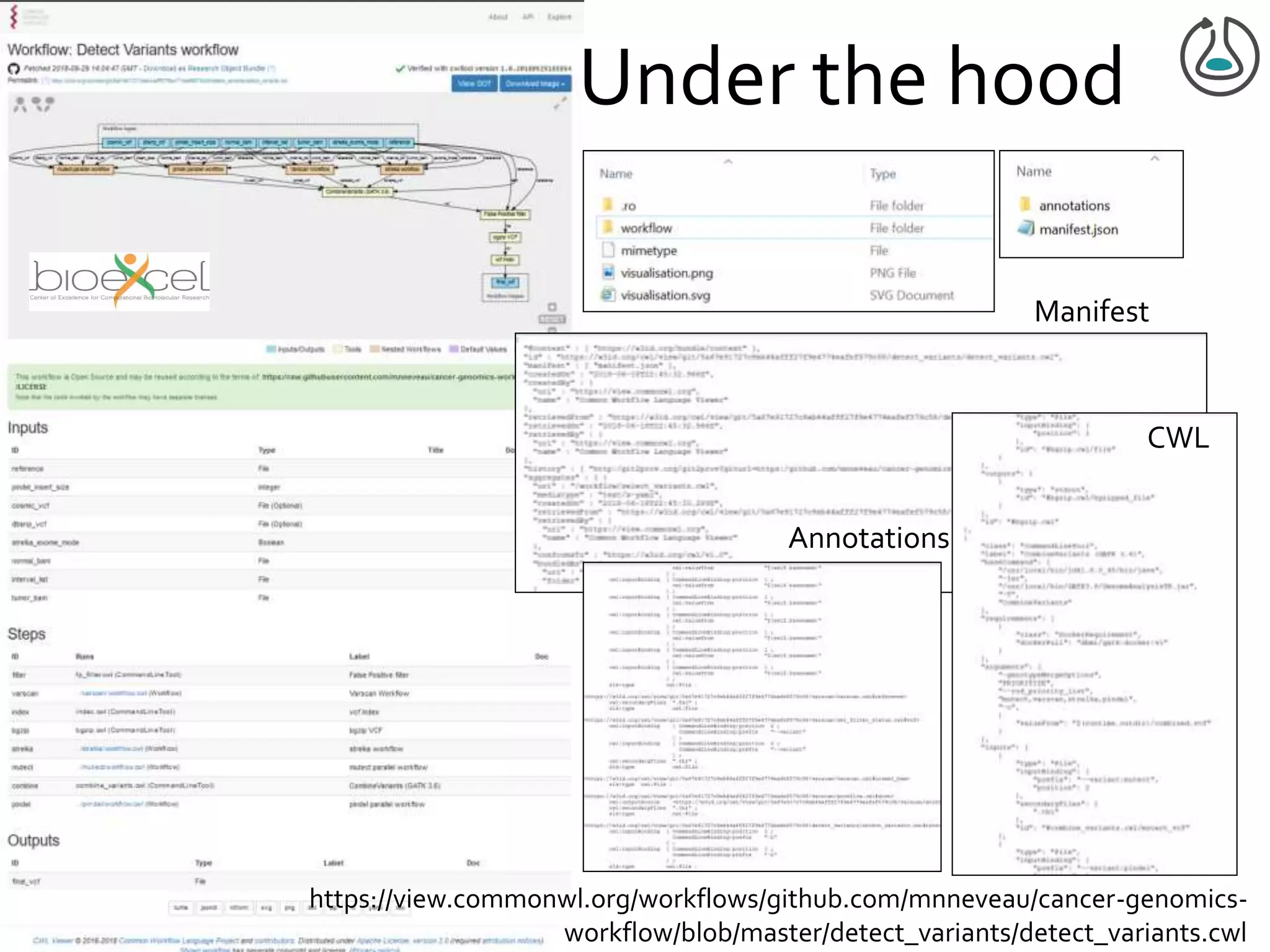
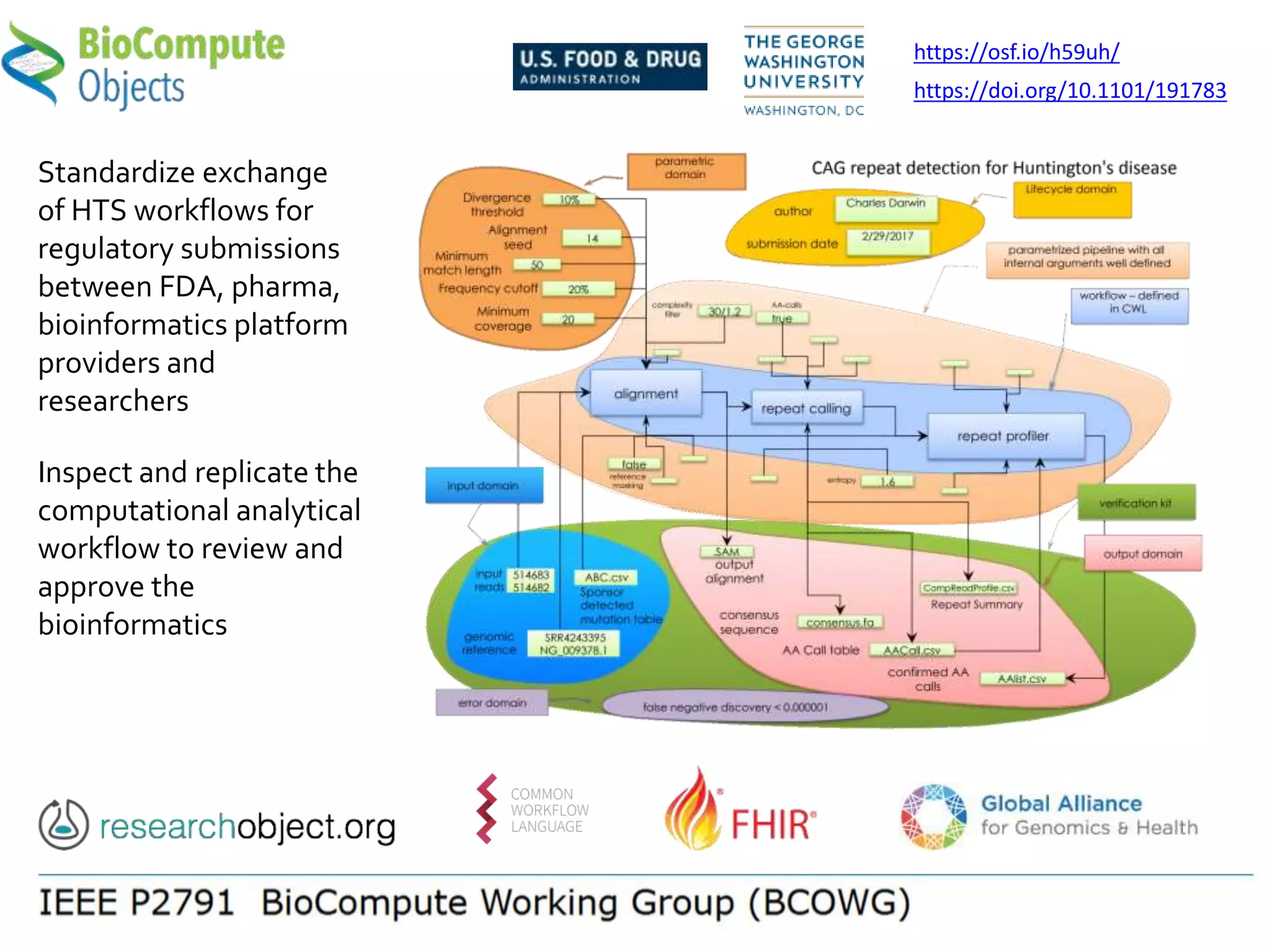
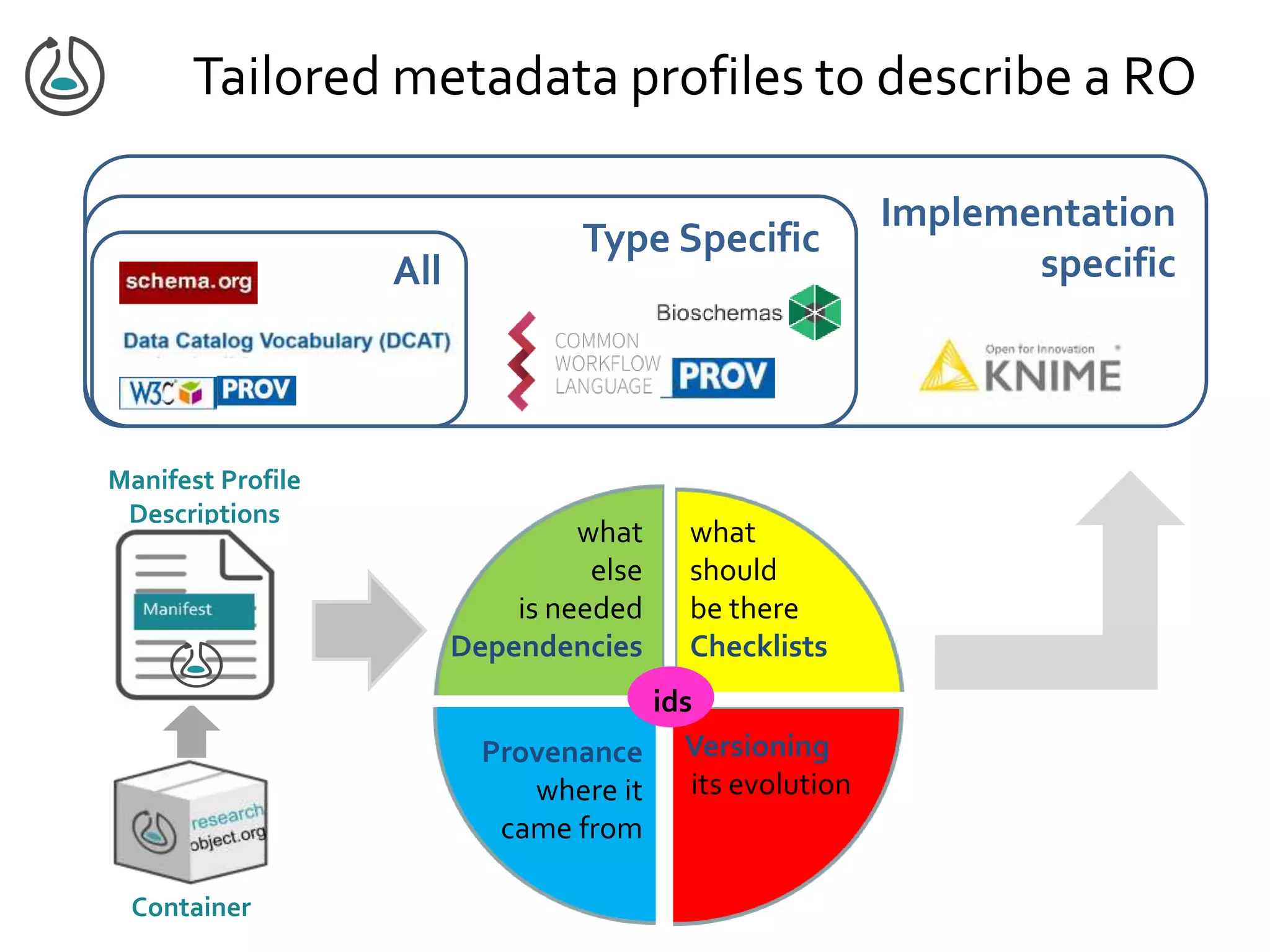
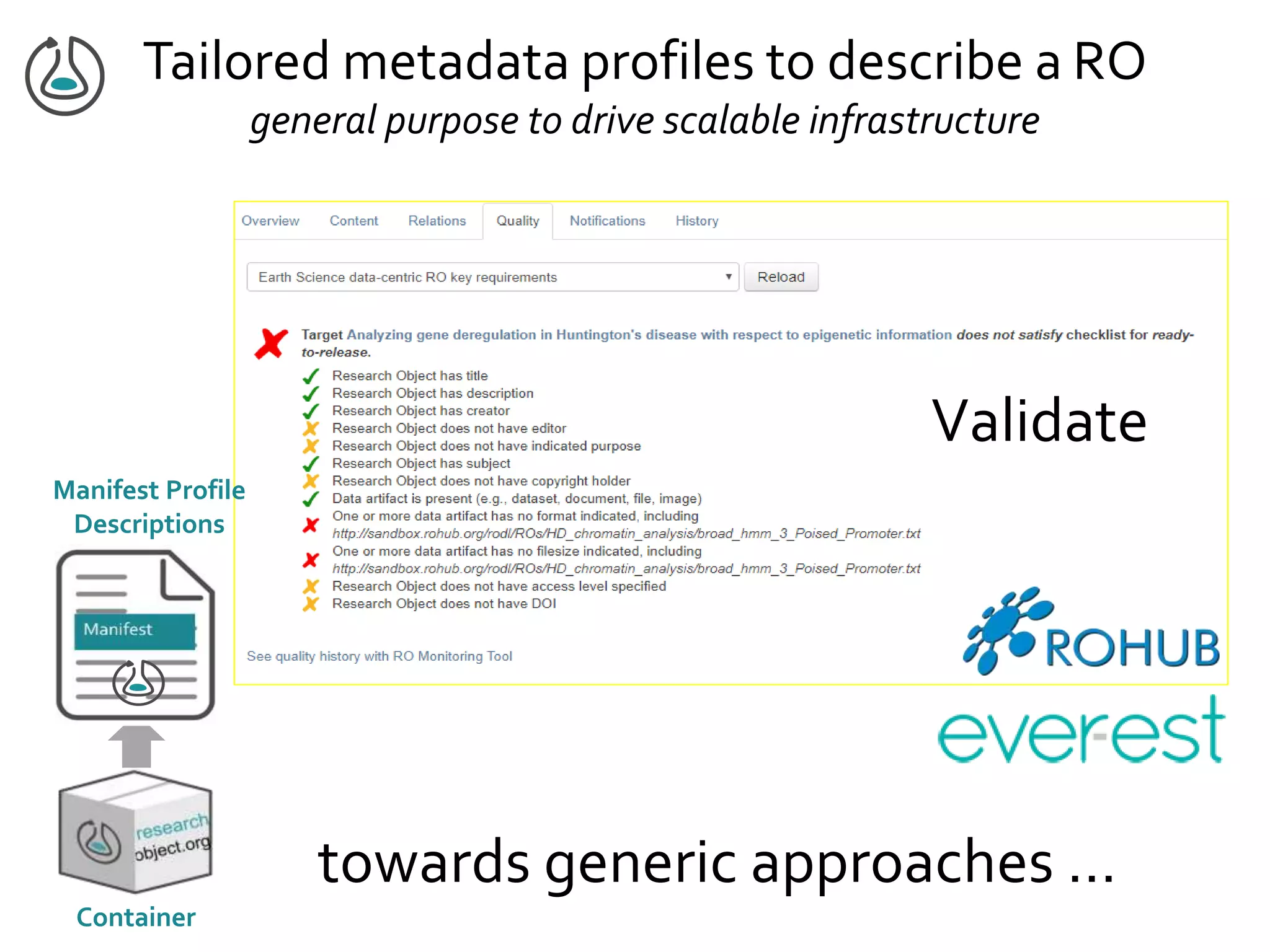
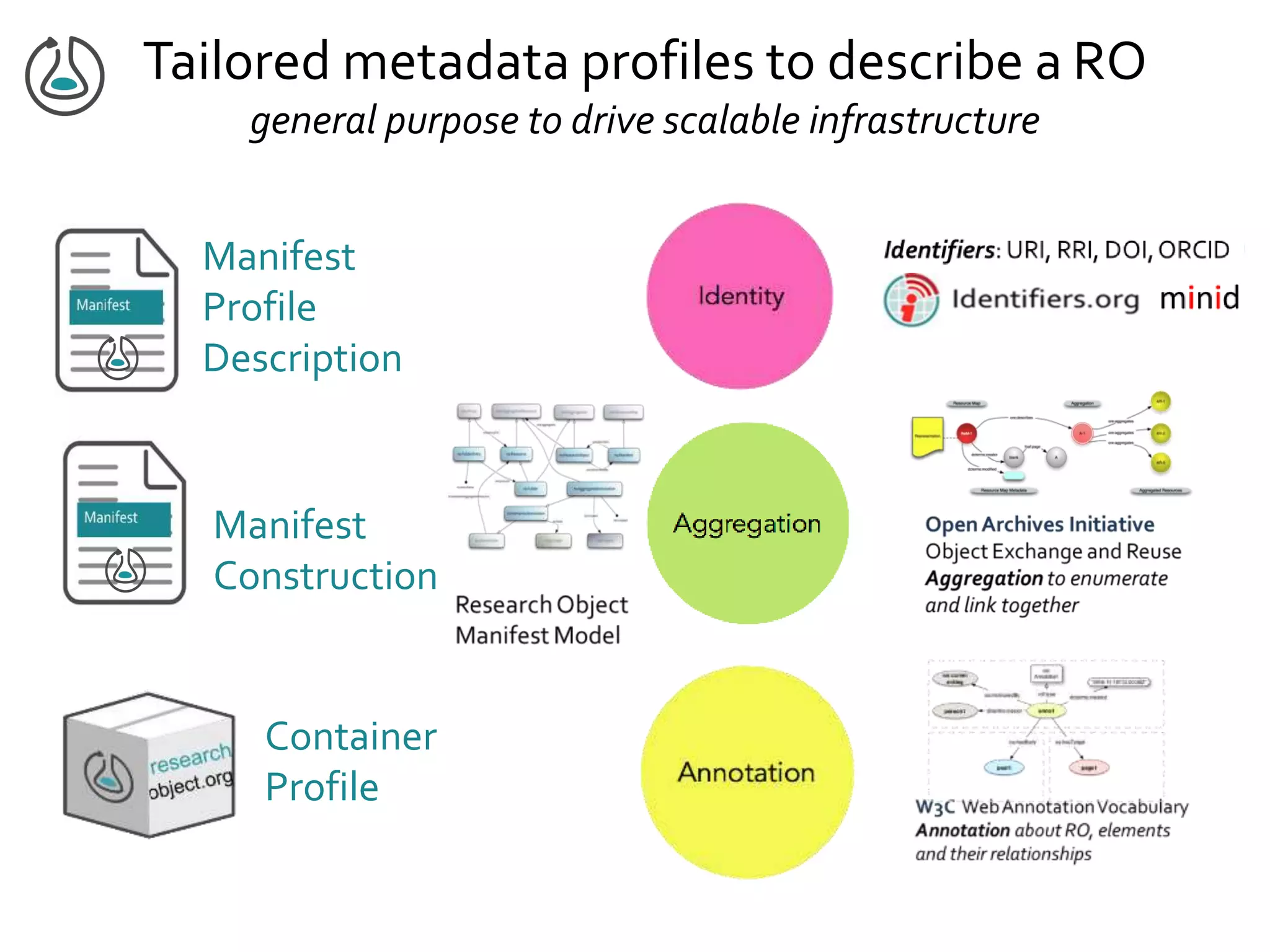

![Manifest Profile Description
general construction & validation tooling
Linked Data and RDF Shapes
Validate graph-based data against a set
of conditions
Shapes Constraint Language
Gamble,Zhao, Klyne,Goble. IEEE eScience 2012,
http://dx.doi.org/10.1109/eScience.2012.6404489
Minim model for defining checklists
ro-show
• RO pre-processing to merge to
single graph
• RDF Shape that indicates to
follow links
• Bespoke validators / unpackers
to iterate over the RO
[Lilian Gorea,Oluwatomide Fasugba 2018]](https://image.slidesharecdn.com/ro2018-goble-181109150205/75/Research-Object-Community-Update-18-2048.jpg)