1) The document discusses randomized algorithms for linear algebra problems like matrix decomposition and column subset selection.
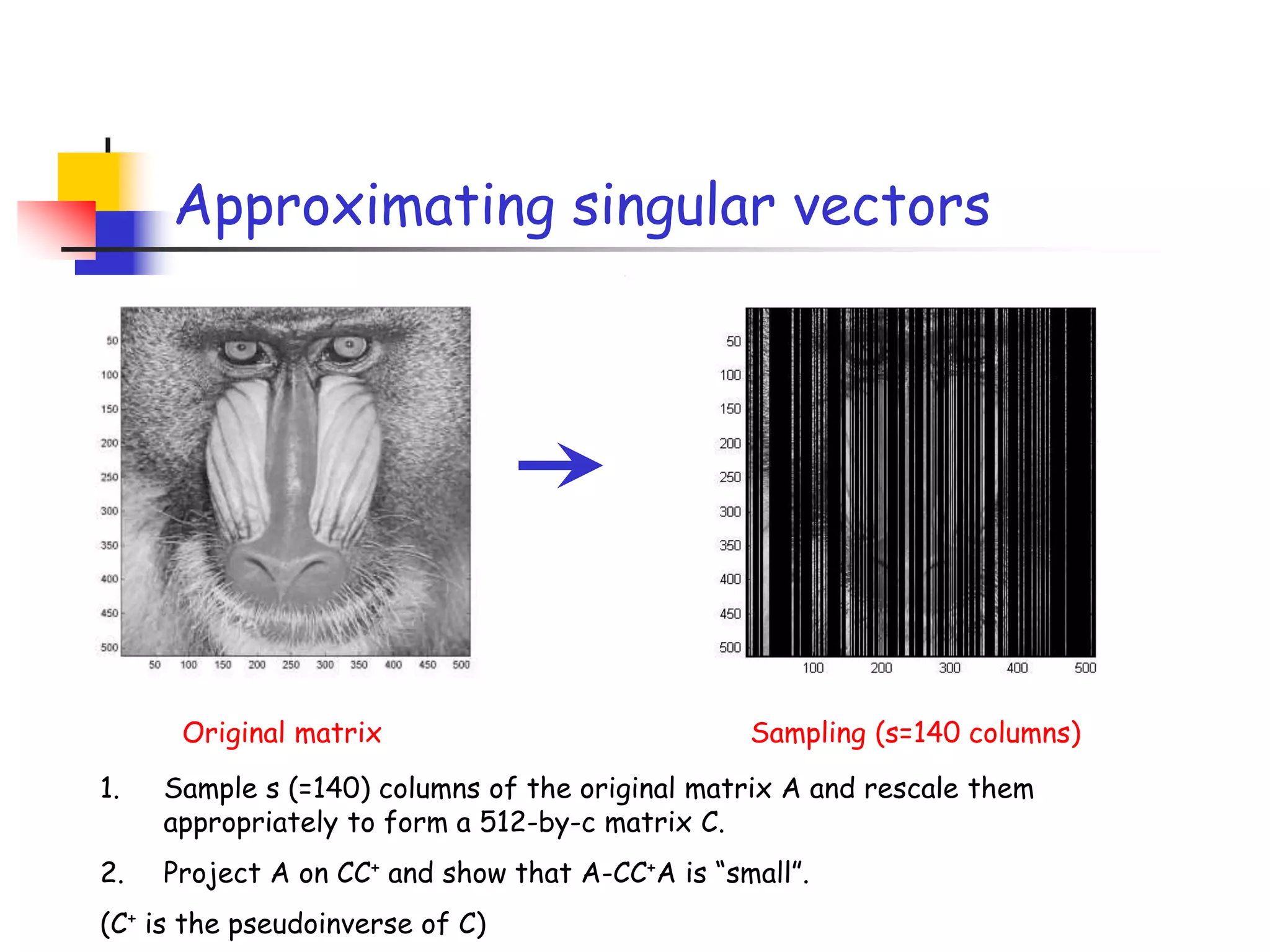
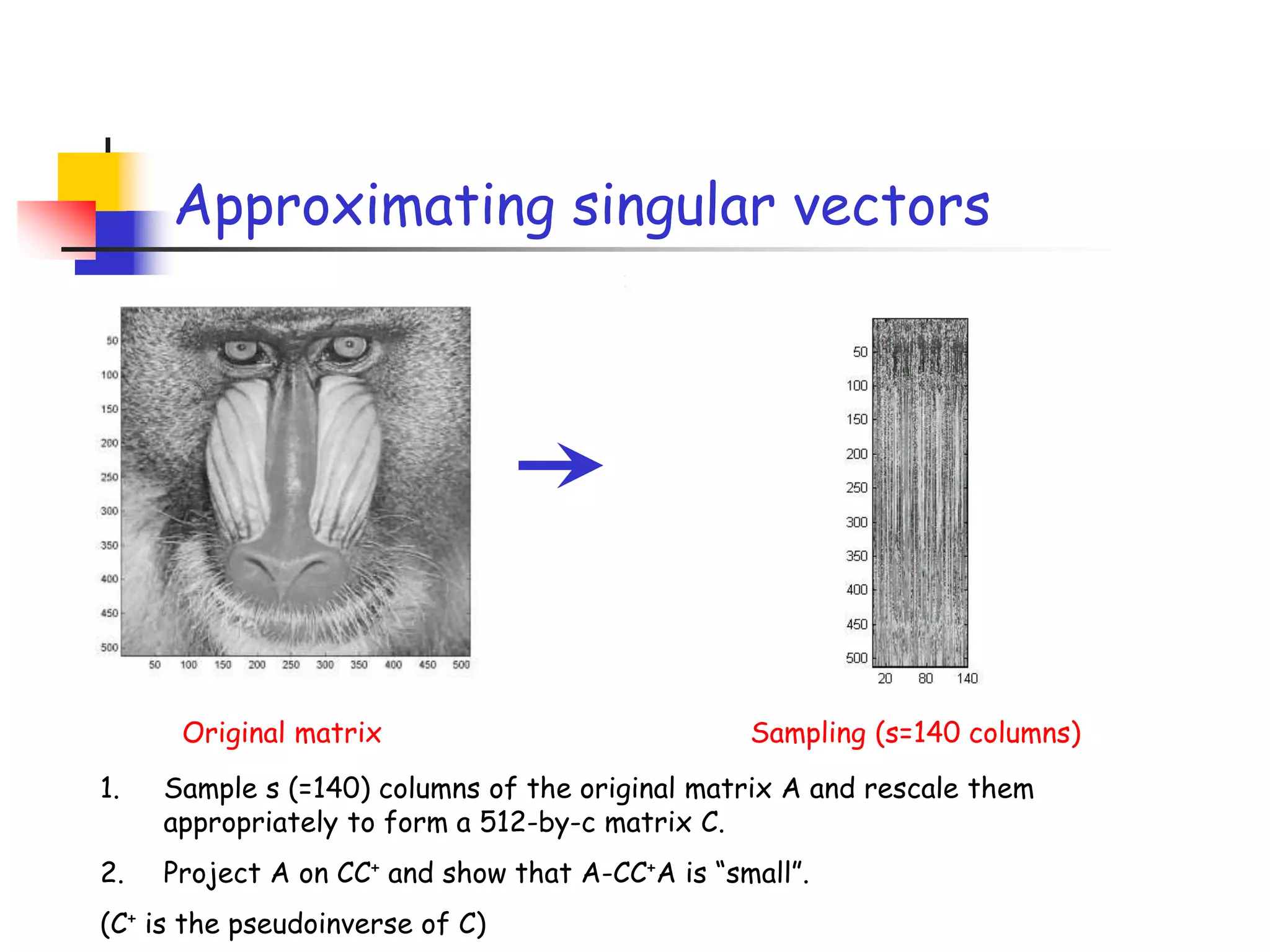
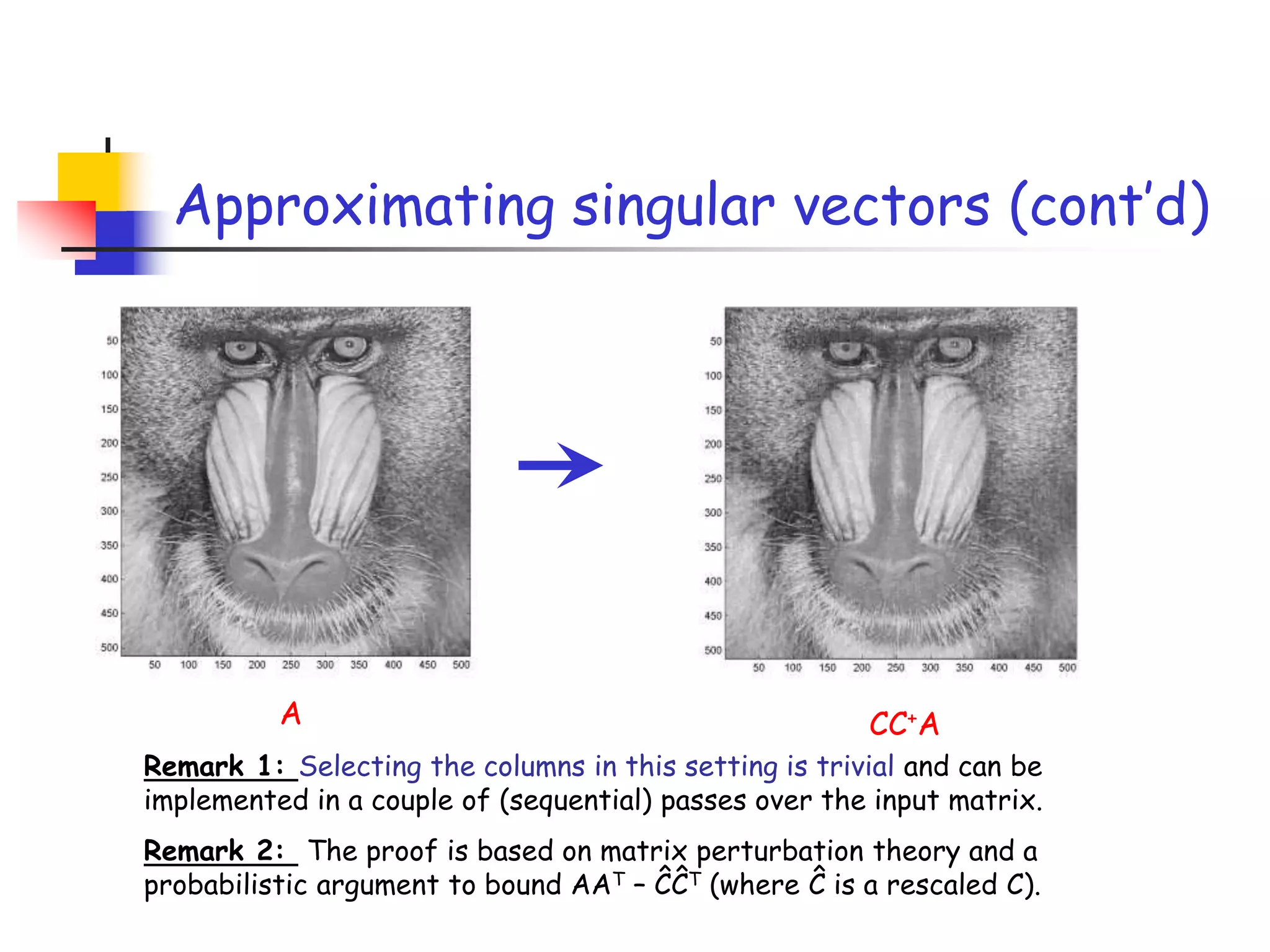
2) It describes how sampling rows/columns of a matrix using random projections can create smaller matrices that approximate the original well.
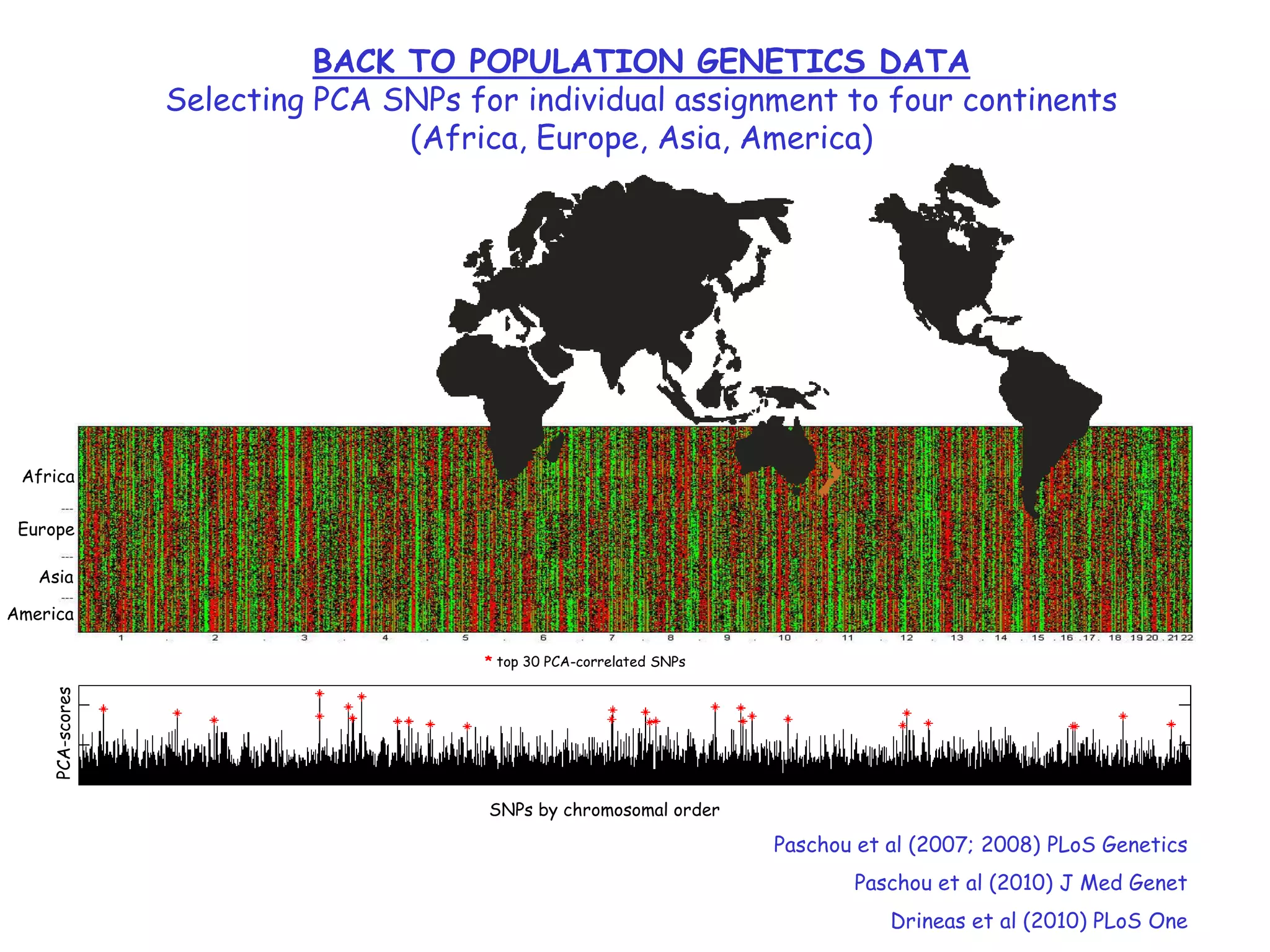
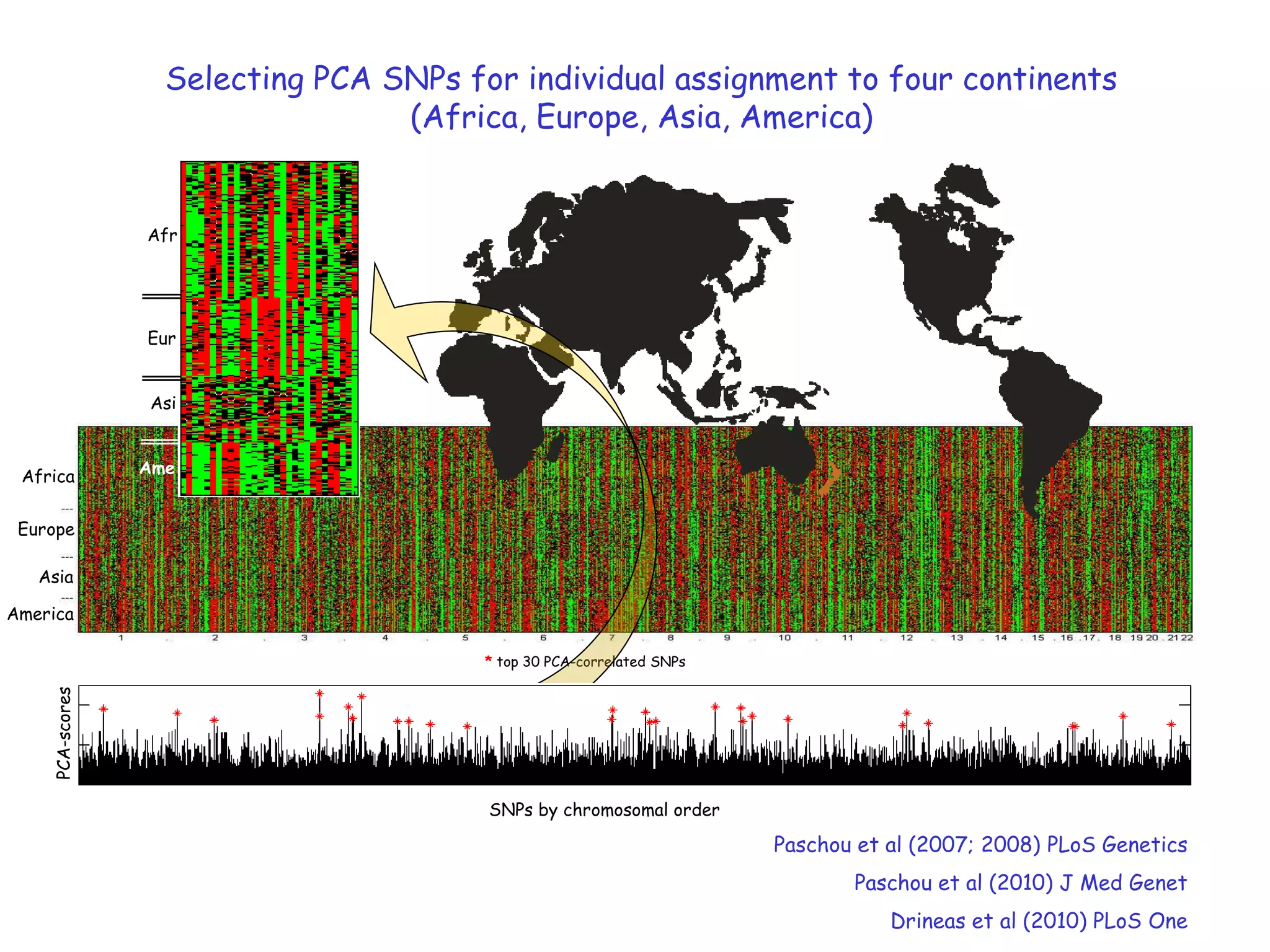
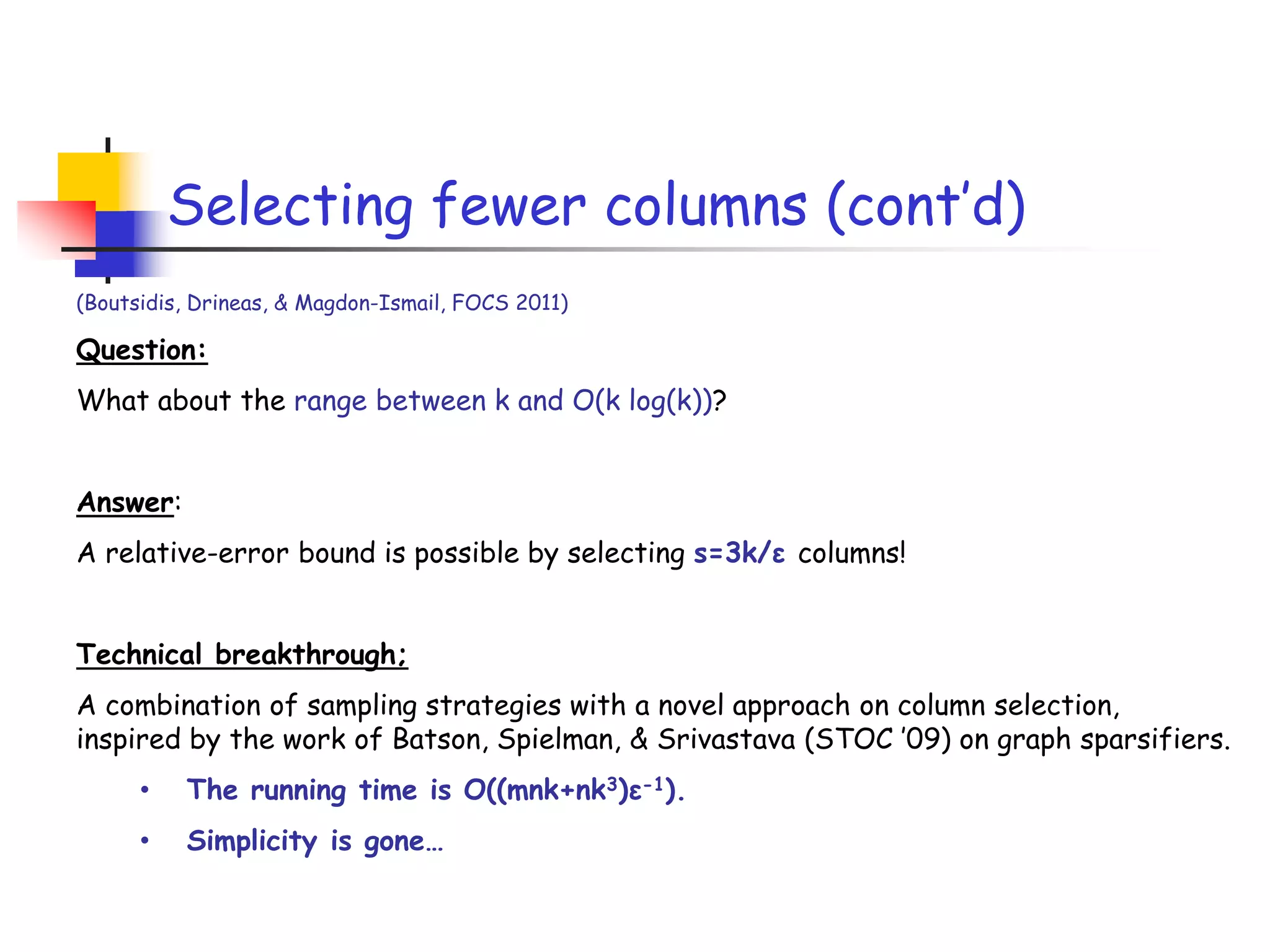
3) The talk will illustrate applications of these ideas to the column subset selection problem and approximating low-rank matrix decompositions.