This document proposes an approach called PTU (Provenance-To-Use) to improve the repeatability of scientific experiments by minimizing computation time during repeatability testing. PTU builds a package containing the software, input data, and provenance trace from a reference execution. Testers can then selectively replay parts of the provenance graph using the ptu-exec tool, reducing testing time compared to full re-execution. The document describes the PTU components, including tools for auditing reference runs, building provenance packages, and selectively replaying parts of the provenance graph. Examples applying PTU to the PEEL0 and TextAnalyzer applications show reductions in testing time.






![PTU Package
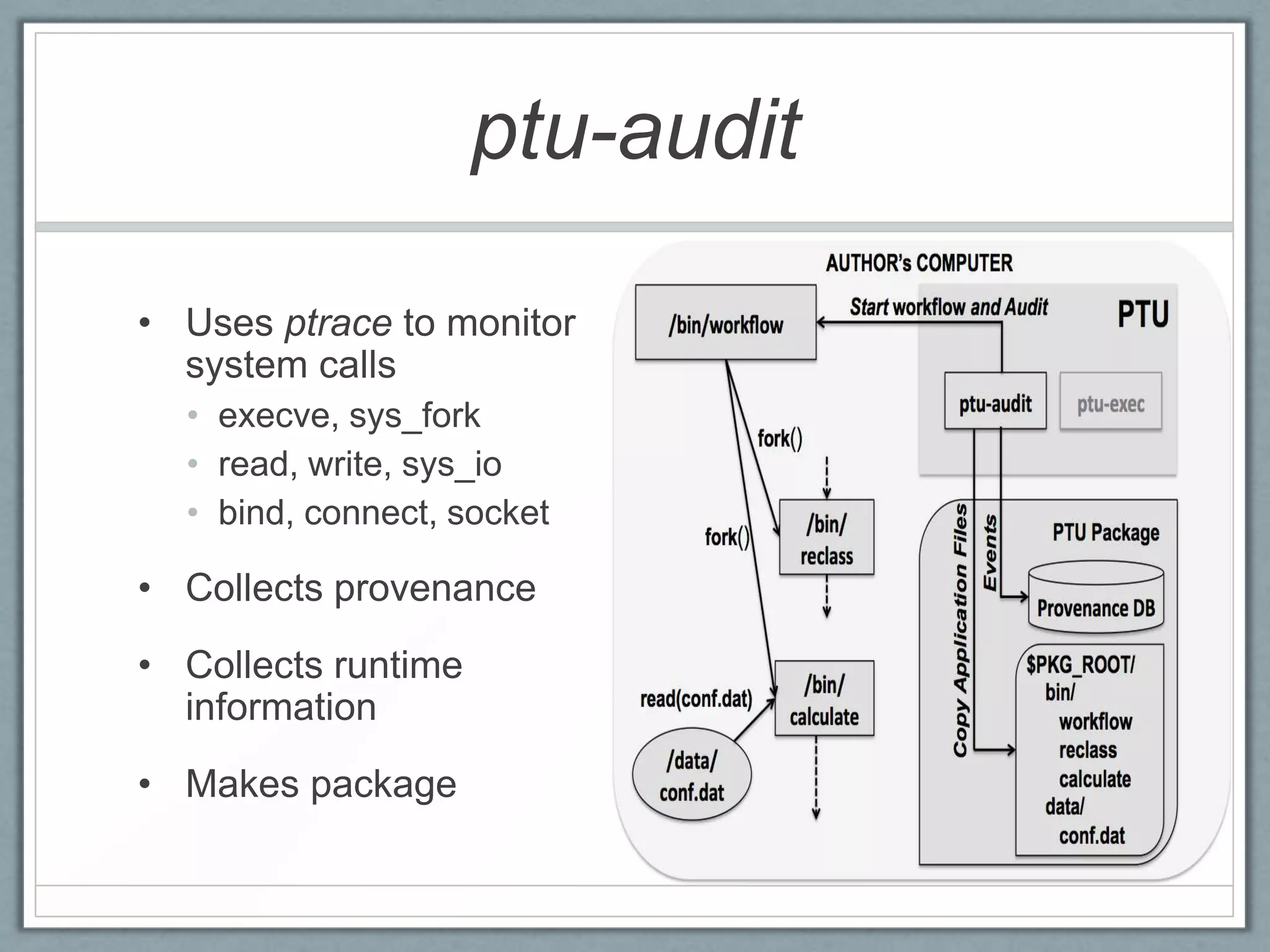
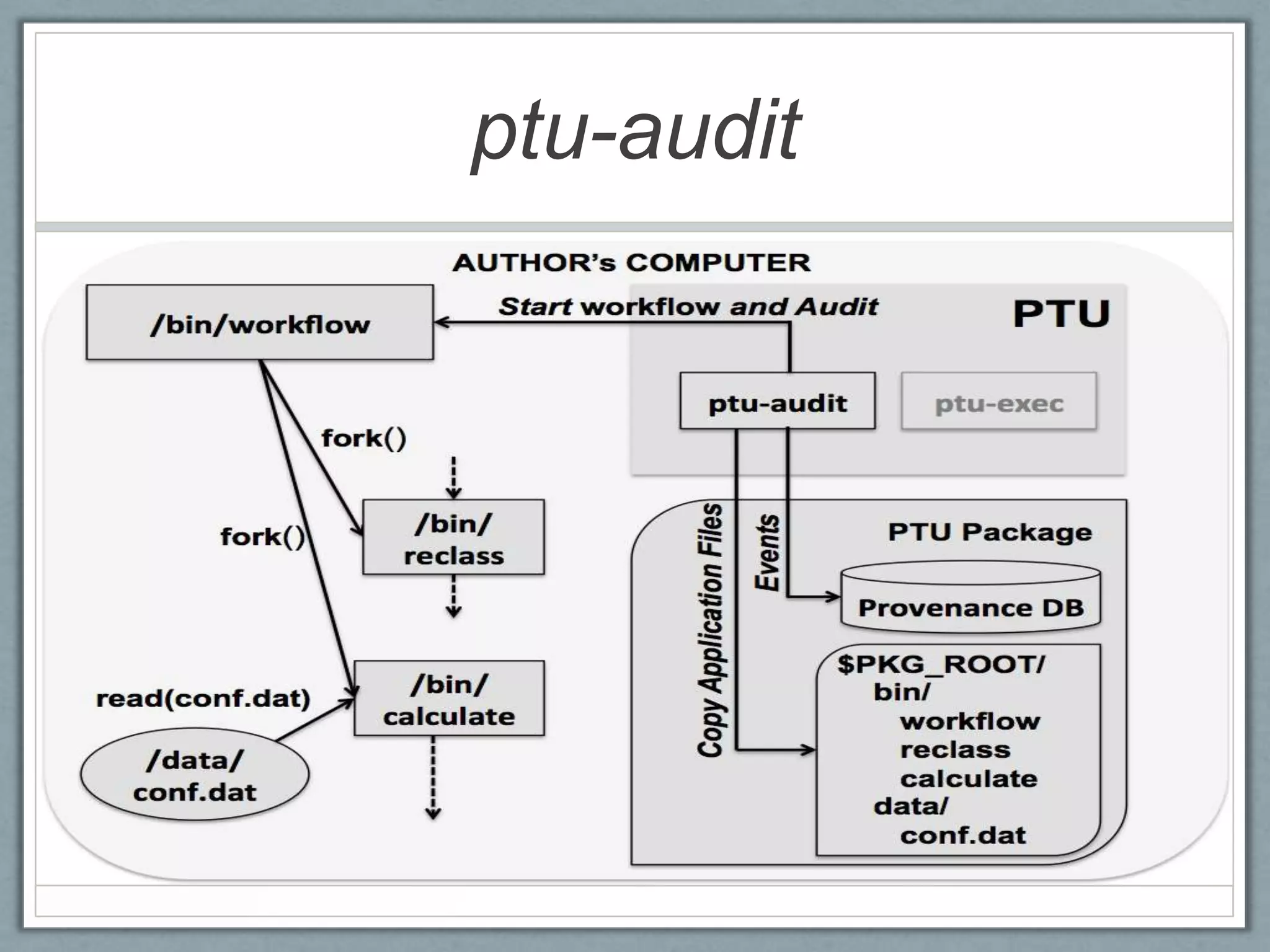
• [Figure 2. The PTU package. The tester chooses
to run the sub-graph rooted at /bin/calculate ]](https://image.slidesharecdn.com/ptu-tapp13-130402115338-phpapp01/75/PTU-Using-Provenance-for-Repeatability-10-2048.jpg)
![ptu-exec
• [Figure 3. ptu-exec re-runs part of the application
from /bin/calculate. It uses CDE to re-route file
dependencies]](https://image.slidesharecdn.com/ptu-tapp13-130402115338-phpapp01/75/PTU-Using-Provenance-for-Repeatability-11-2048.jpg)


![PEEL0
• [Figure 4: Time reduction in testing PEEL0 using
PTU]
• Or use the actual execution graph??](https://image.slidesharecdn.com/ptu-tapp13-130402115338-phpapp01/75/PTU-Using-Provenance-for-Repeatability-14-2048.jpg)

![TextAnalyzer
• [Figure 5. Time reduction in testing TextAnalyzer
using PTU]](https://image.slidesharecdn.com/ptu-tapp13-130402115338-phpapp01/75/PTU-Using-Provenance-for-Repeatability-16-2048.jpg)