The document discusses the impact of machine learning on cognitive neuroimaging, emphasizing its ability to address complex questions and improve models of brain activity through encoding and decoding approaches. It highlights the importance of rich encoding models, the challenges of interpreting overlapping activations, and the role of machine learning in enhancing the understanding of cognitive functions. Various statistical methods and models, including multi-voxel pattern analysis, are explored to improve predictions related to brain activity and its implications for cognitive neuroscience.
![Machine learning and cognitive neuroimaging:
new tools can answer new questions
Gaël Varoquaux
How machine learning is shaping cognitive neuroimaging
[Varoquaux and Thirion 2014]](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-1-2048.jpg)
![Cognitive neuroscience: linking psychology and
neuroscience (neural implementations)
Vision: A computational investigation into the human representation
and processing of visual information [Marr 1982]
G Varoquaux 2](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-2-2048.jpg)
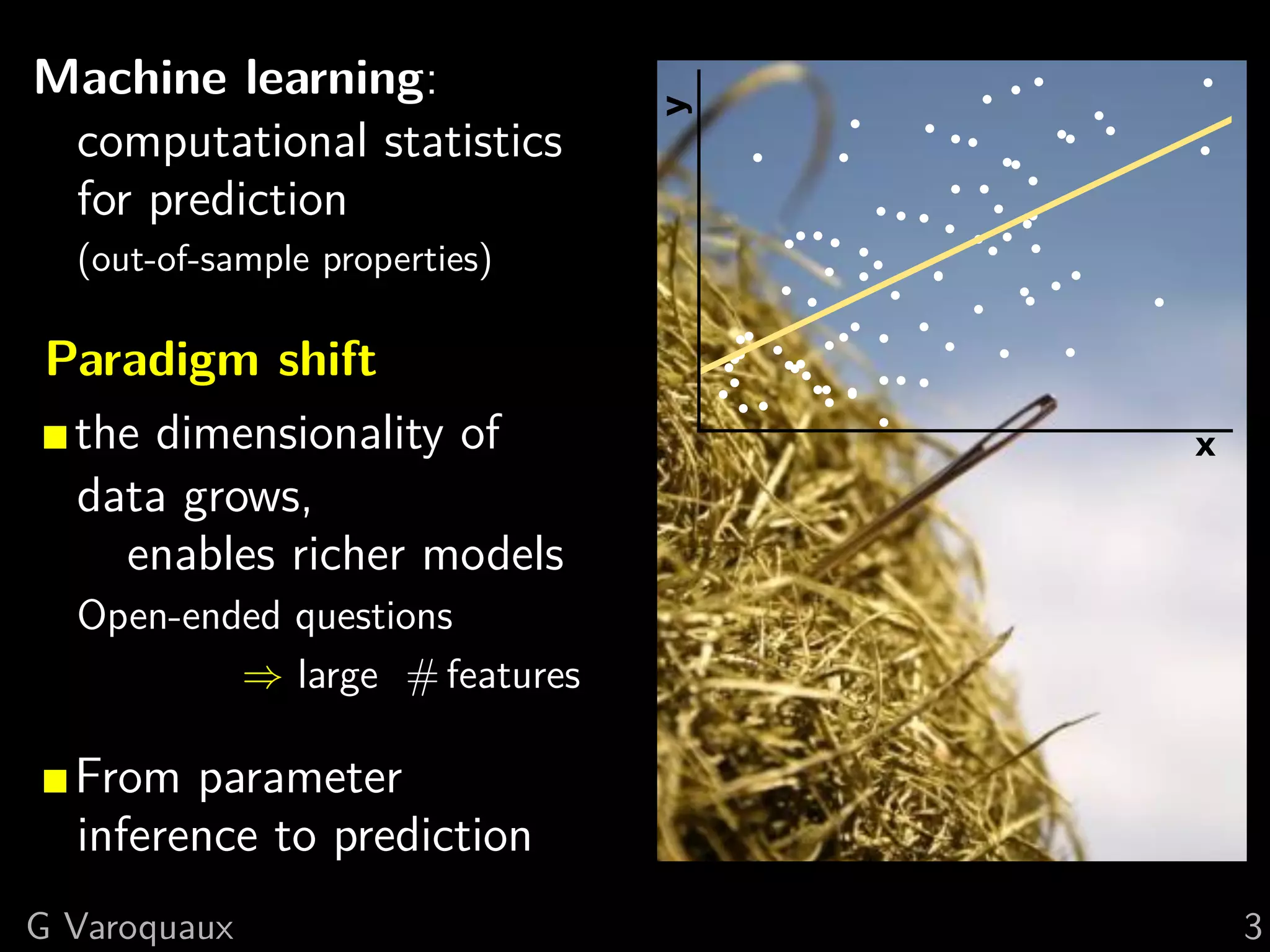







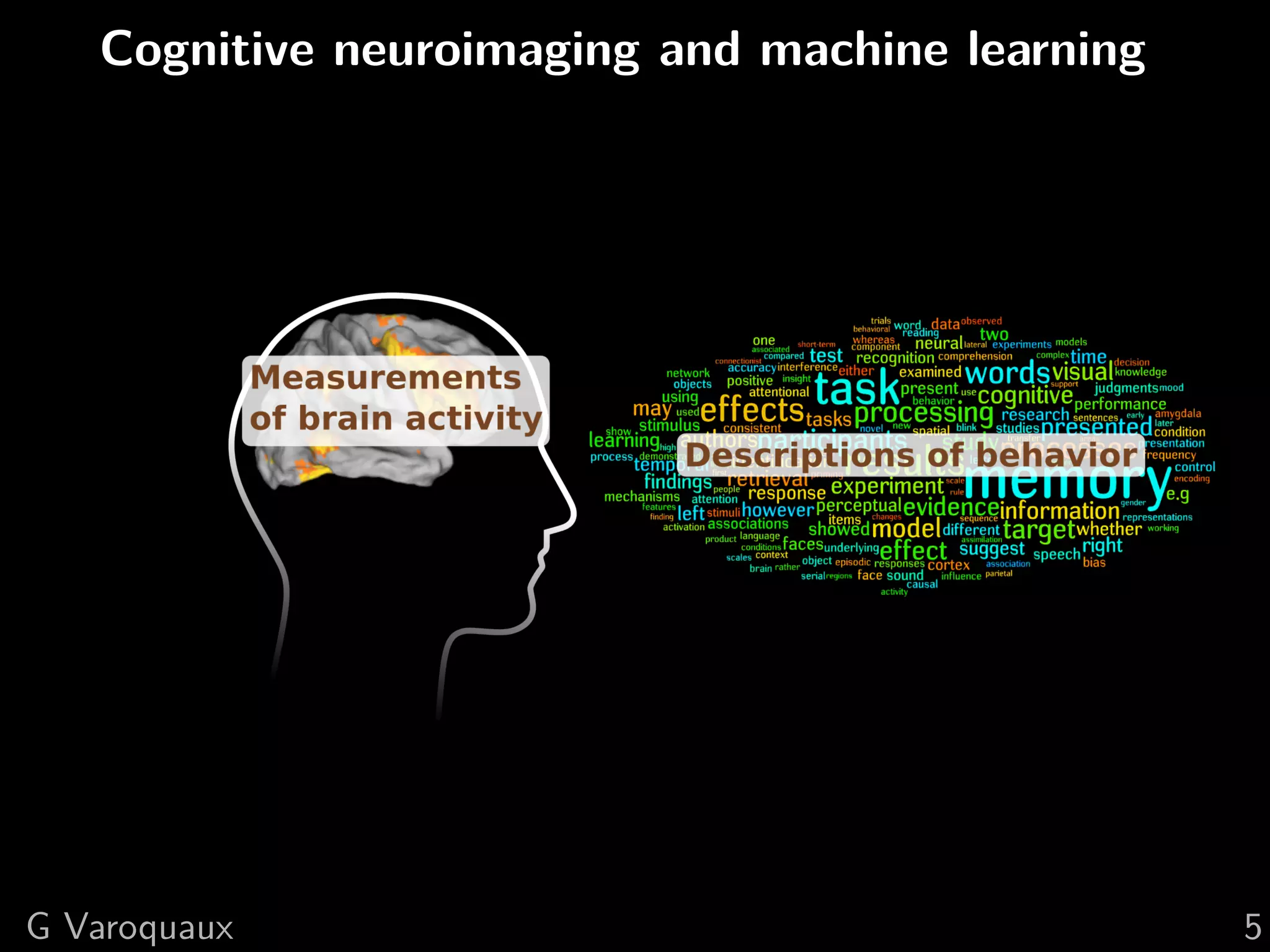
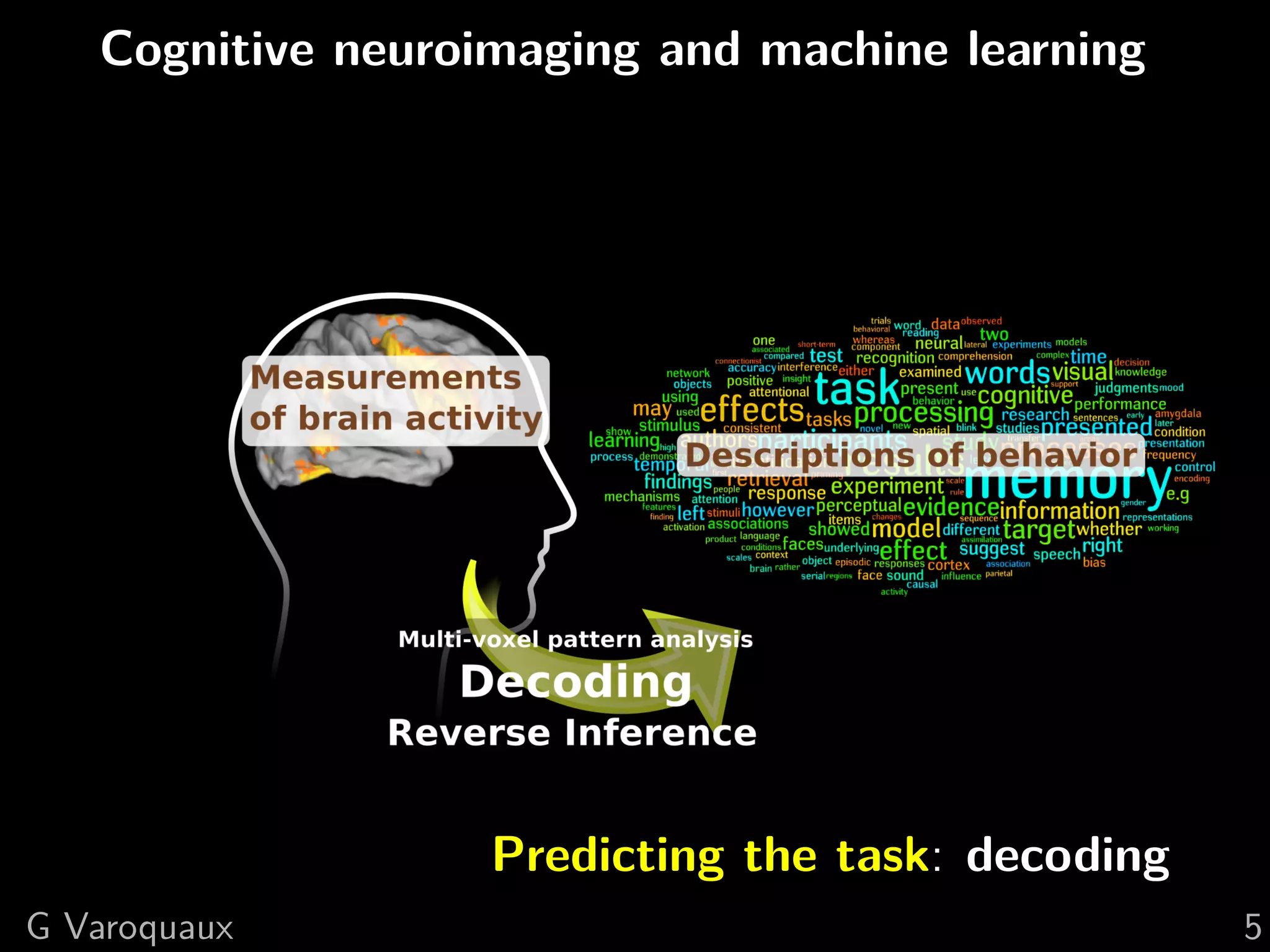
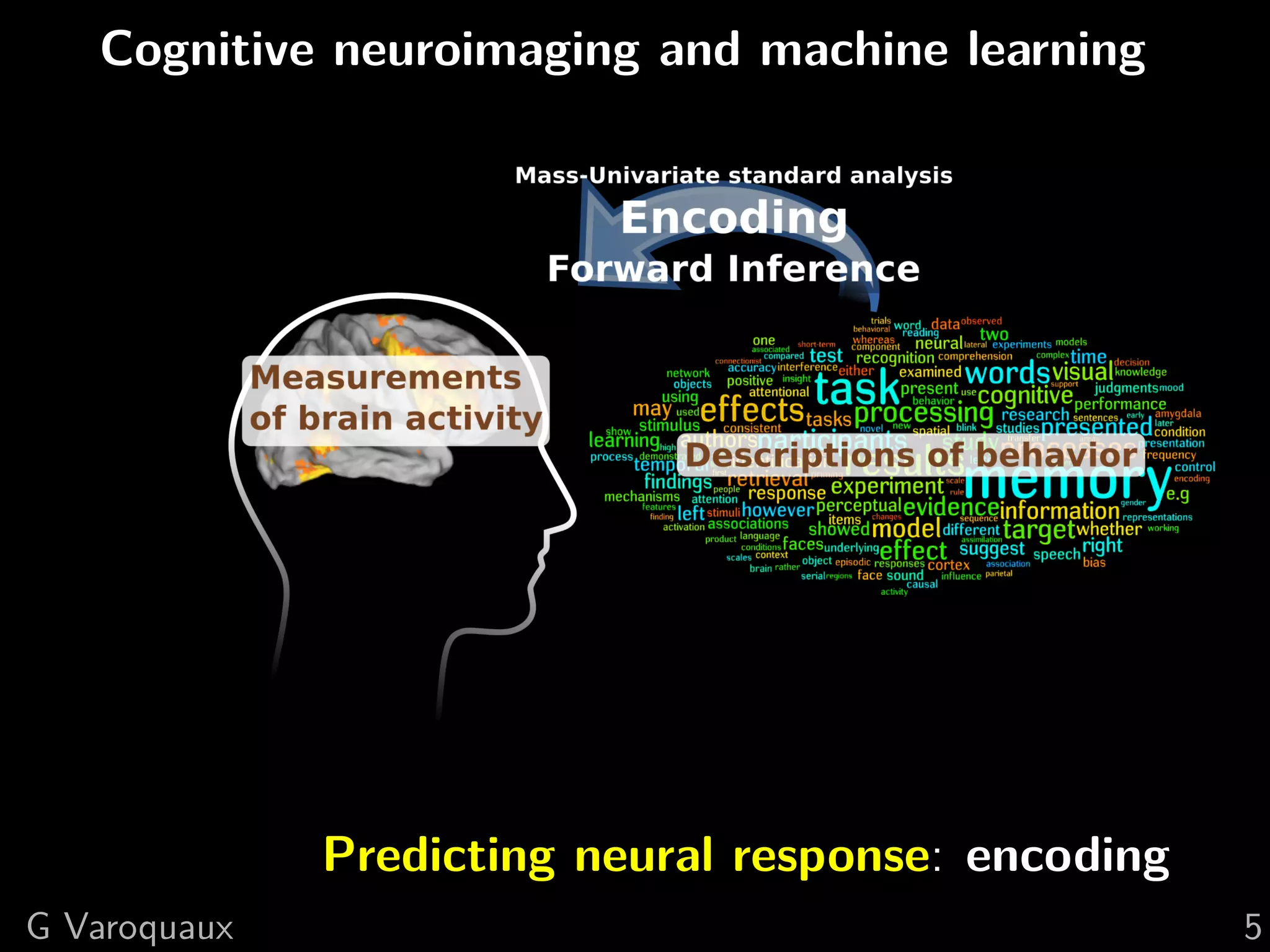
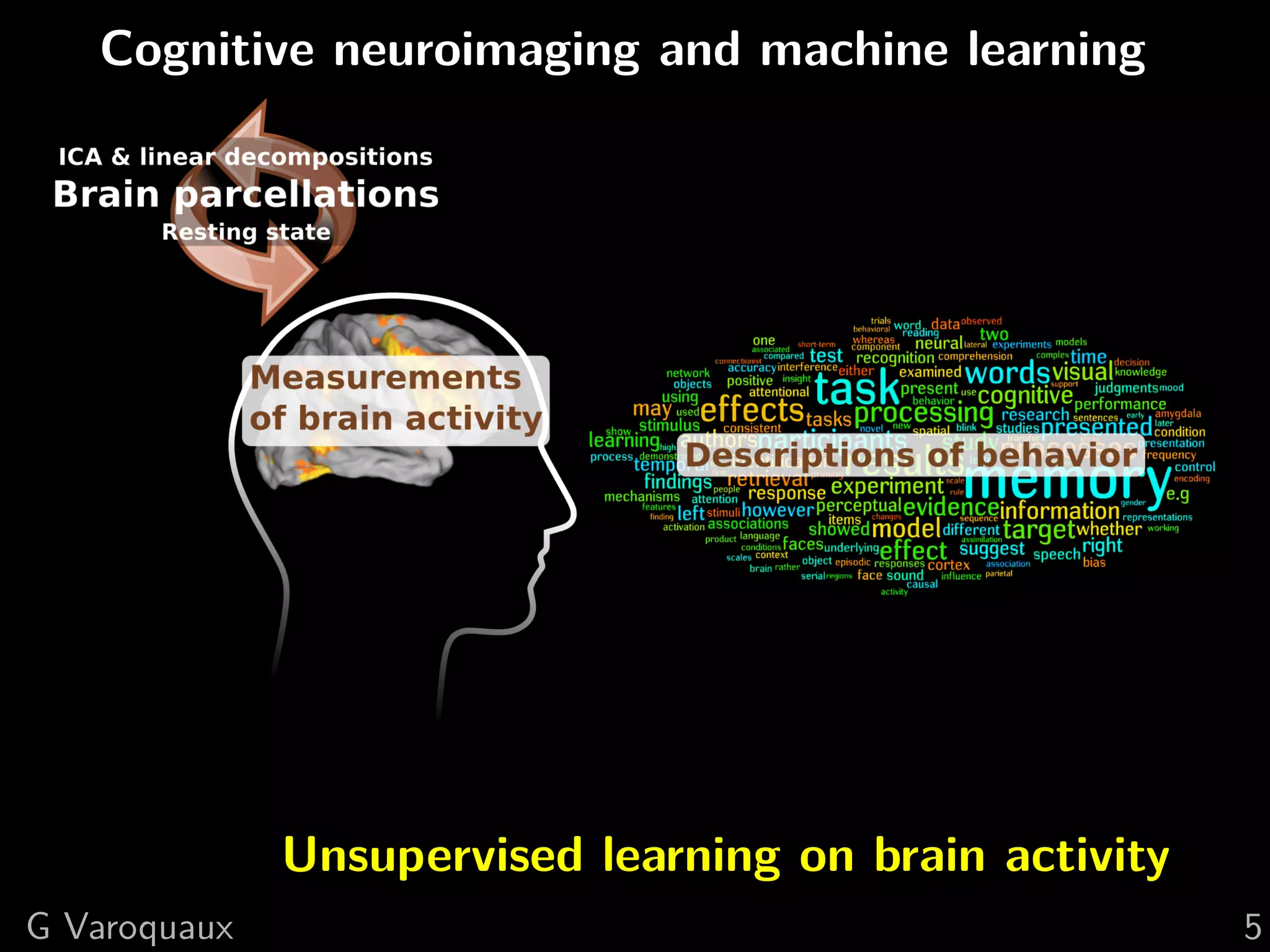

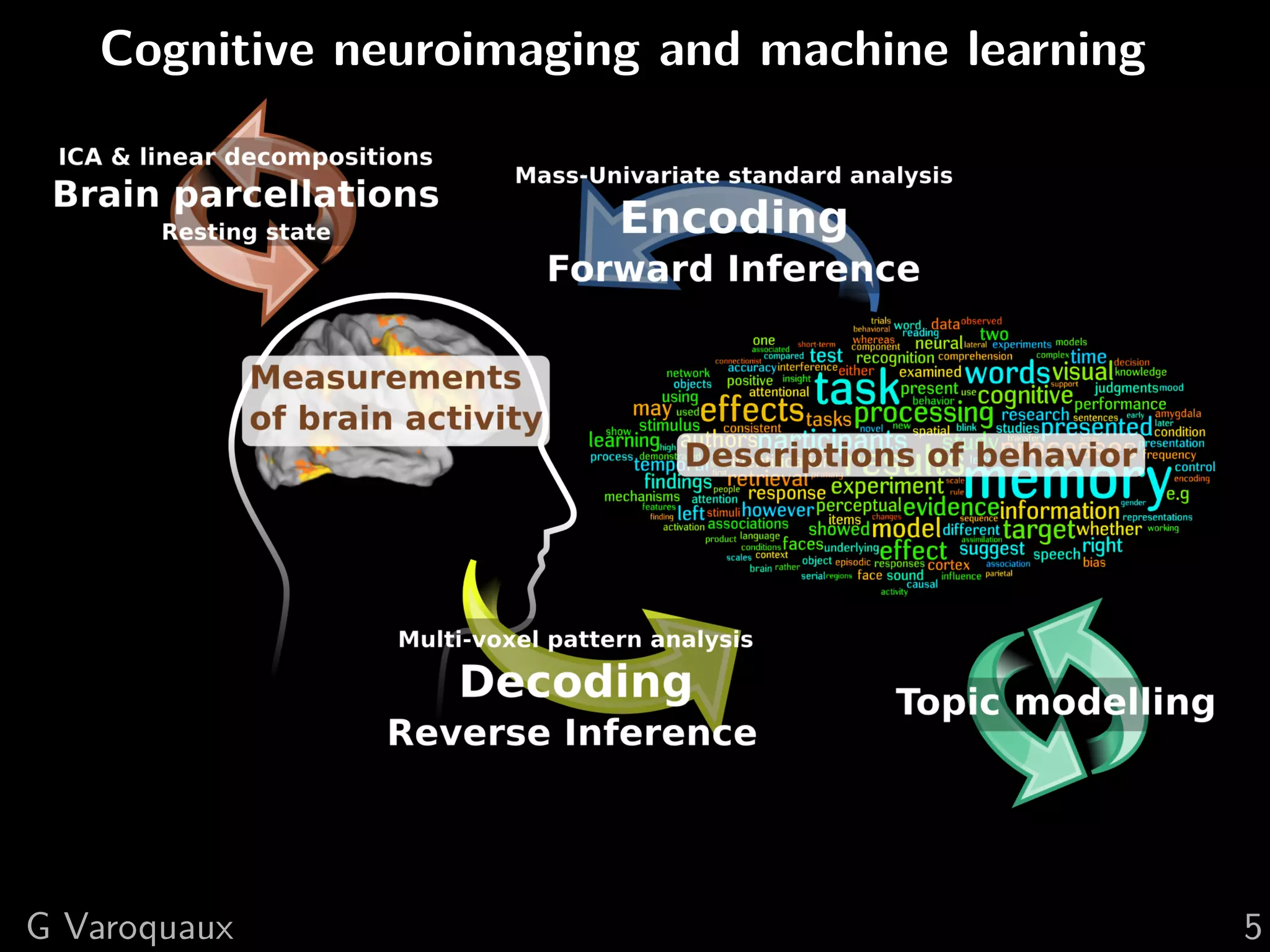
![1 Uncovering neural coding
Insights on breaking down cognitive functions into
atomic steps
[Hubel and Wiesel 1962]
Neurons receptive to
Gabors (edges)
G Varoquaux 8](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-19-2048.jpg)
![1 Uncovering neural coding
Insights on breaking down cognitive functions into
atomic steps
[Hubel and Wiesel 1962]
Neurons receptive to
Gabors (edges)
[Logothetis... 1995]
Shapes in inferior
temporal cortex
G Varoquaux 8](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-20-2048.jpg)
![1 Uncovering neural coding: richer models
Insights on breaking down cognitive functions into
atomic steps
[Hubel and Wiesel 1962]
Neurons receptive to
Gabors (edges)
[Logothetis... 1995]
Shapes in inferior
temporal cortex
Machine learning:
computer-vision models mapped to brain activity
[Yamins... 2014]
G Varoquaux 8](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-21-2048.jpg)
![1 Uncovering neural coding: in fMRI
Model-based fMRI [O’Doherty... 2007]
[Harvey... 2013]
High-level descriptions [Mitchell... 2008]
Natural stimuly [Kay... 2008]
G Varoquaux 9](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-22-2048.jpg)


![1 Decomposing visual stimuli
Low-level visual cortex is tuned
to natural image statistics
[Olshausen et al. 1996]
What drives high-level representations?
G Varoquaux 13](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-29-2048.jpg)
![1 Decomposing visual stimuli
Low-level visual cortex is tuned
to natural image statistics
[Olshausen et al. 1996]
What drives high-level representations?
Convolutional Net
G Varoquaux 13](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-30-2048.jpg)
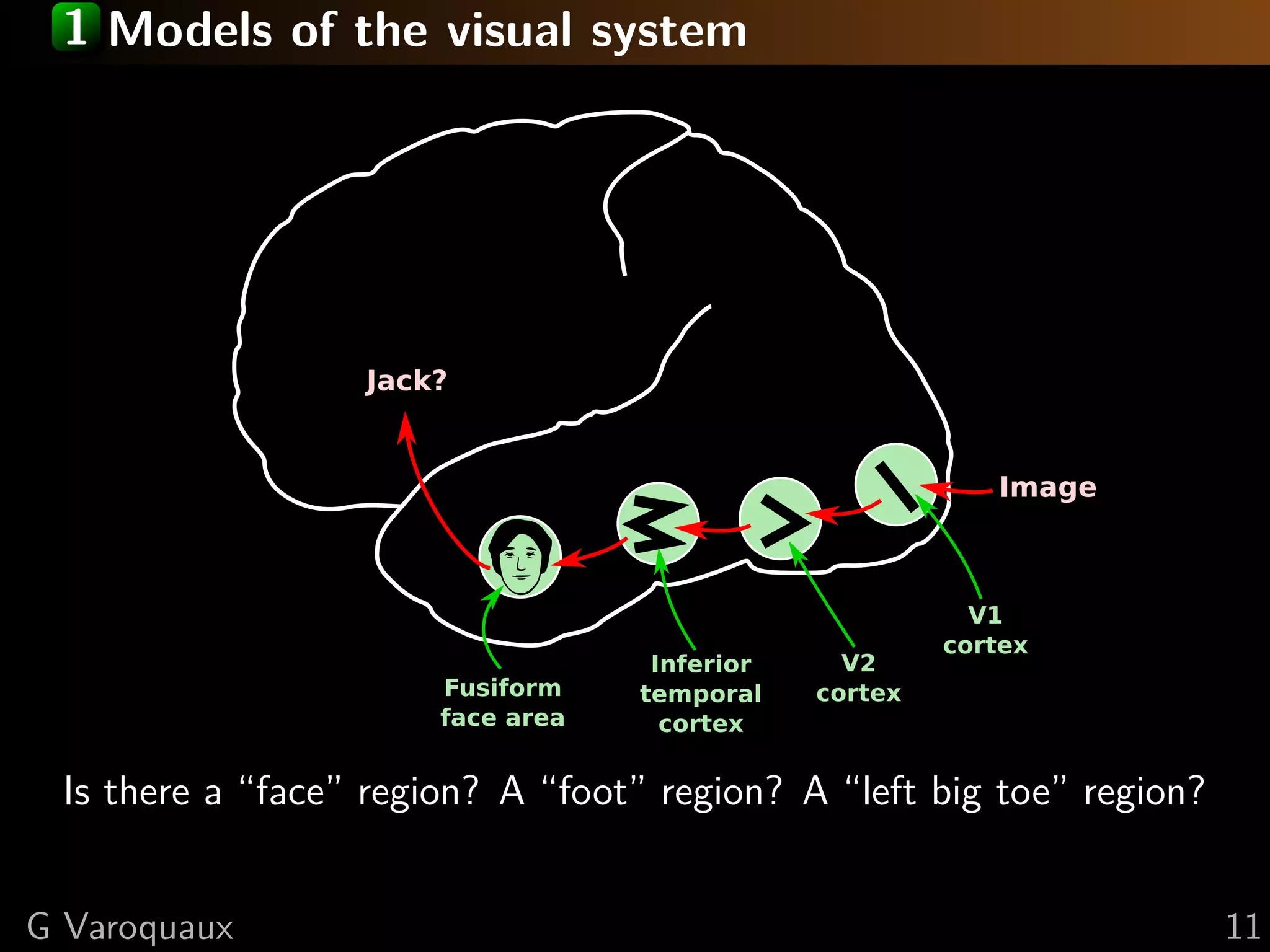
![Data-driven encoding models
Image
V1
cortex
V2
cortex
Inferior
temporal
cortex
Fusiform
face area
Jack?
[Khaligh-Razavi and Kriegeskorte 2014, Güçlü and van Gerven 2015]
FMRI beyond a handfull of contrasts
⇒ Sets us free from the paradigm
G Varoquaux 14](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-31-2048.jpg)
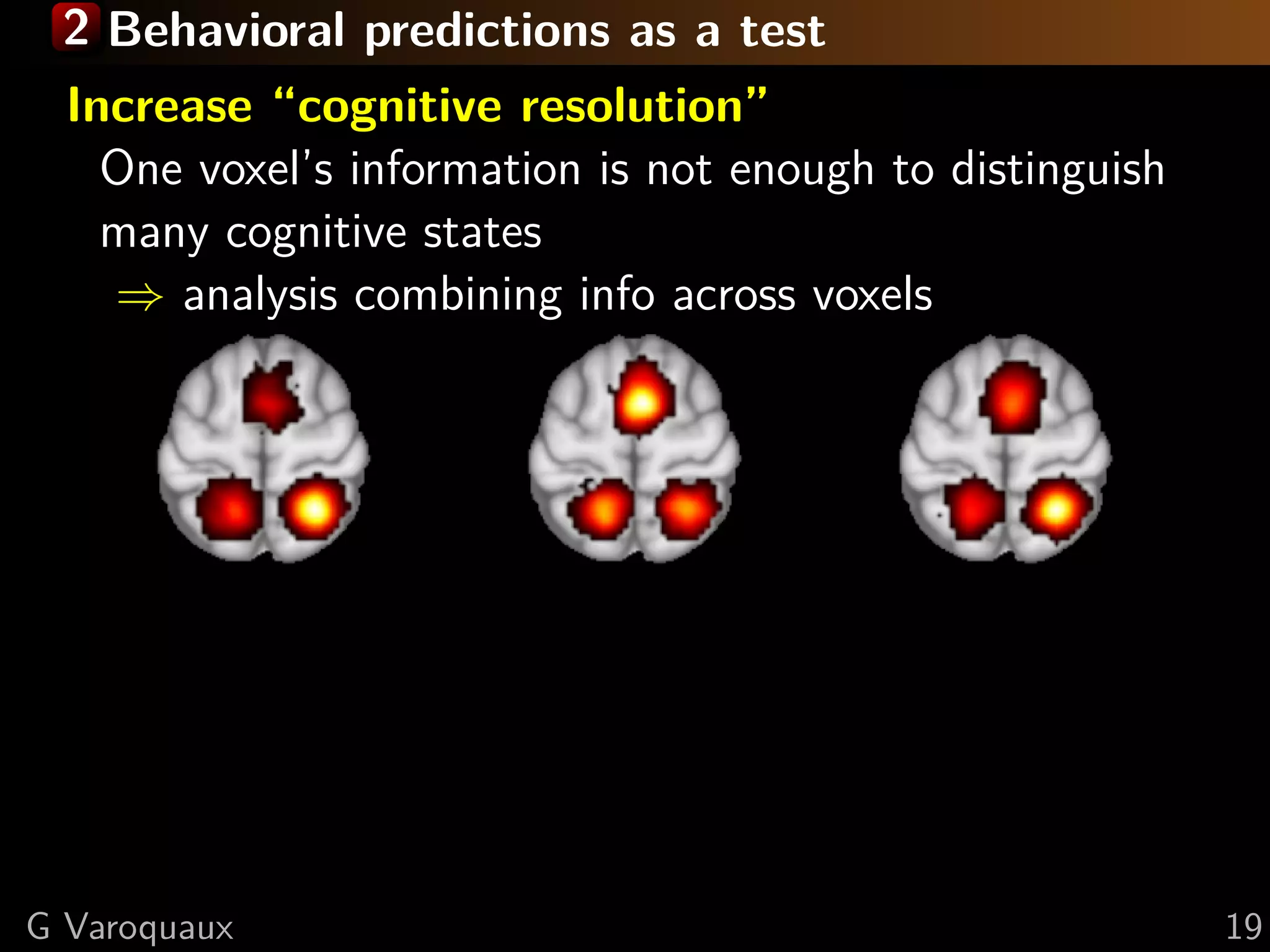
![2 Increased sensitivity
“Given the goal of detecting the presence of a particular
mental representation in the brain, the primary advantage
of MVPA methods over individual-voxel-based methods is
increased sensitivity.” — [Norman... 2006]
G Varoquaux 16](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-33-2048.jpg)
![2 Increased sensitivity
An omnibus test
“Given the goal of detecting the presence of a particular
mental representation in the brain, the primary advantage
of MVPA methods over individual-voxel-based methods is
increased sensitivity.” — [Norman... 2006]
Is there “information” about a
stimuli in a given region?
G Varoquaux 16](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-34-2048.jpg)
![2 Increased sensitivity
An omnibus test
“Given the goal of detecting the presence of a particular
mental representation in the brain, the primary advantage
of MVPA methods over individual-voxel-based methods is
increased sensitivity.” — [Norman... 2006]
“However, these maps are not guaranteed to include all
the voxels that are involved in representing the categories
of interest.” — [Norman... 2006]
G Varoquaux 16](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-35-2048.jpg)
![Non-linear
cognitive model
Linear
predictive models
Representations
Stimuli
2 Increased sensitivity
An omnibus test
Decoding used to test / compare encoding models
[Naselaris... 2011]
G Varoquaux 17](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-36-2048.jpg)
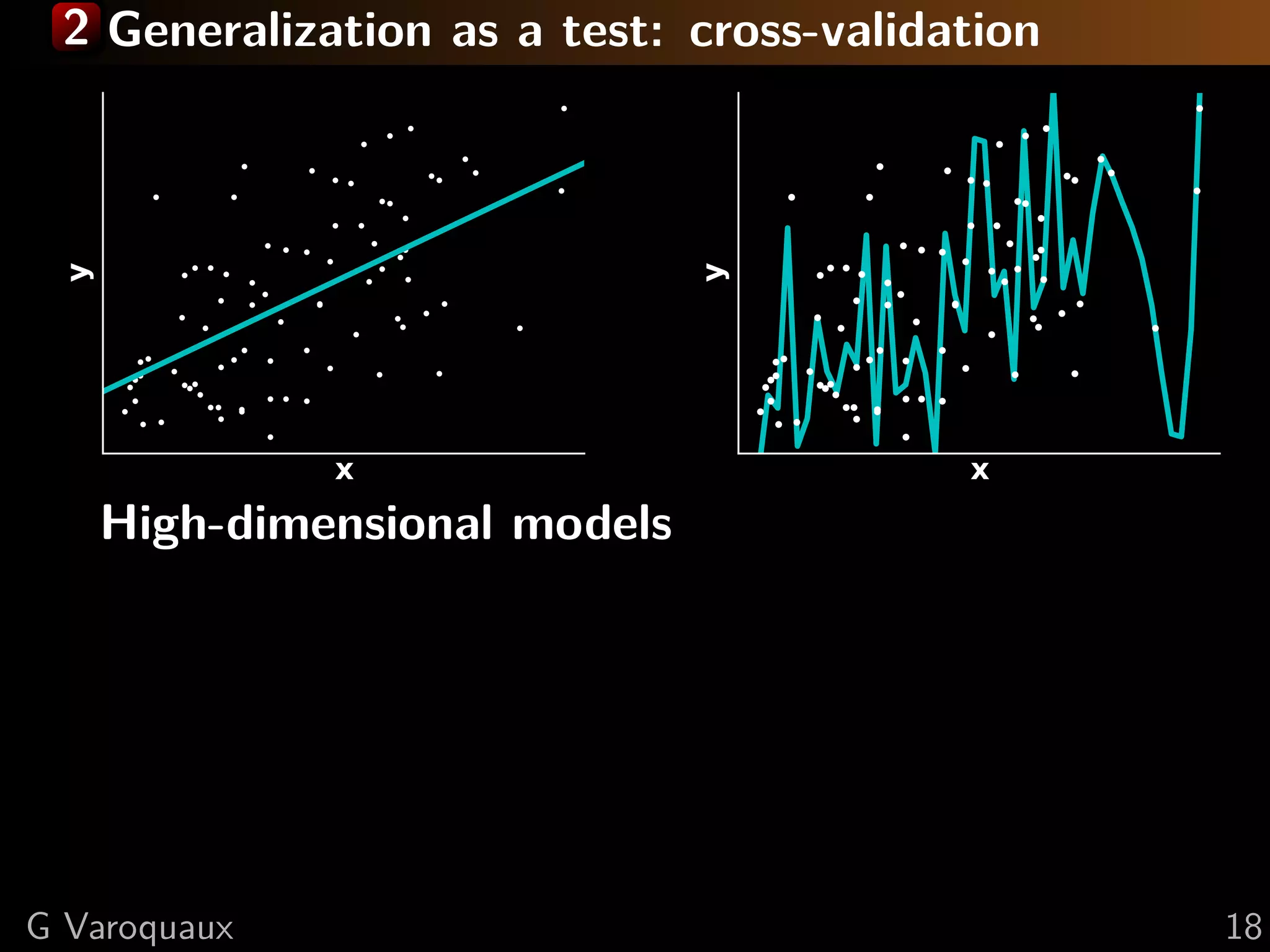
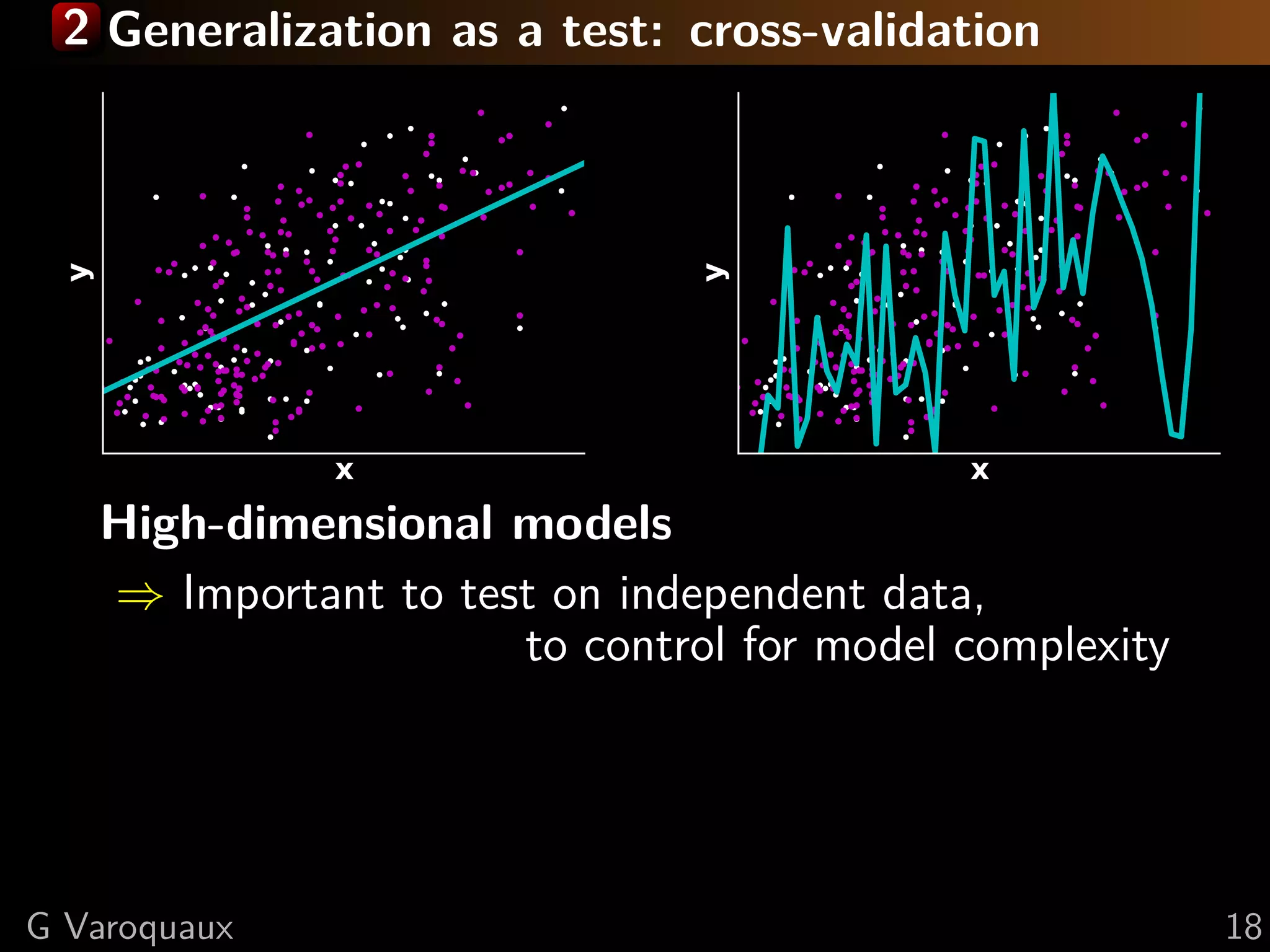
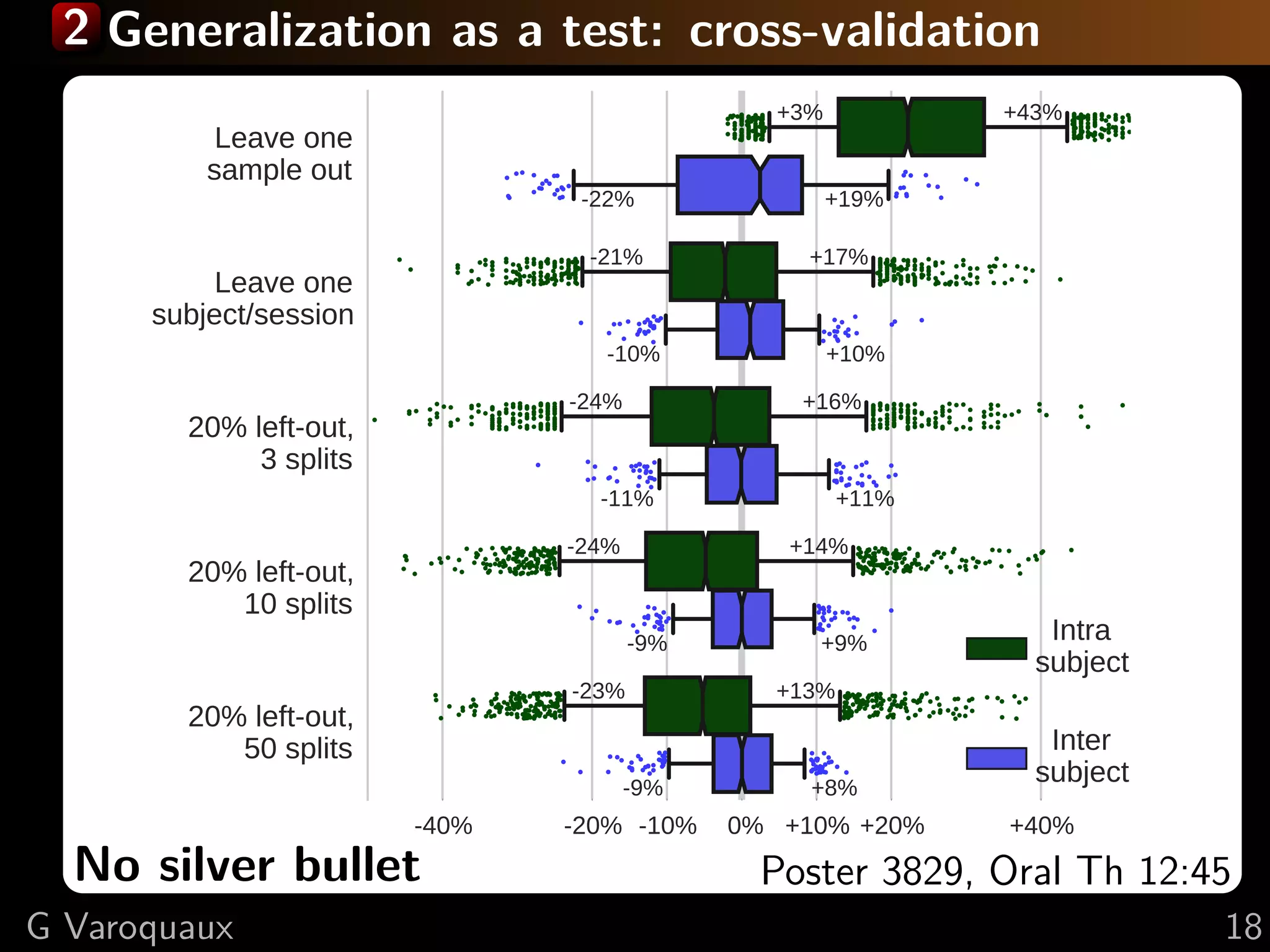

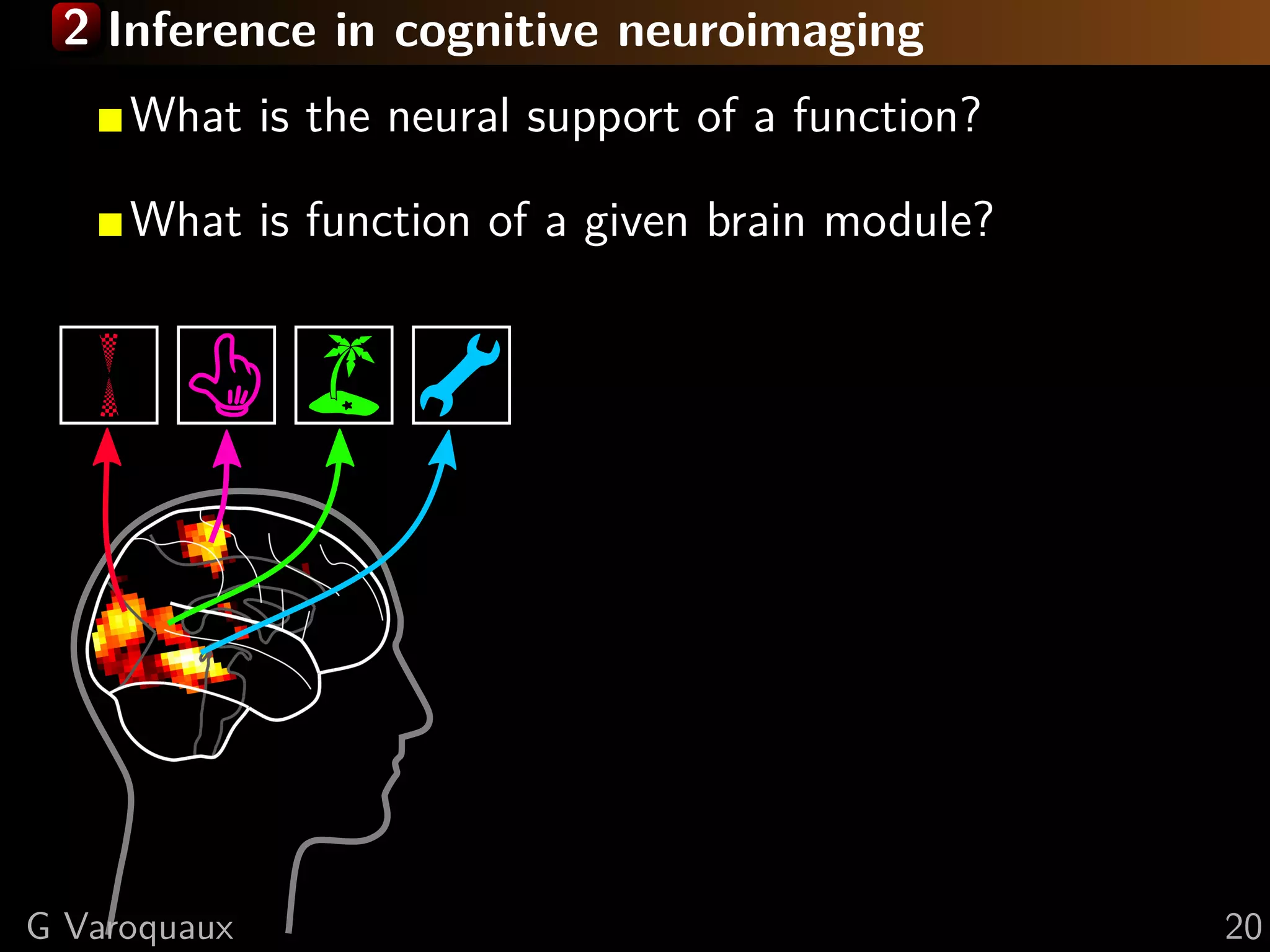
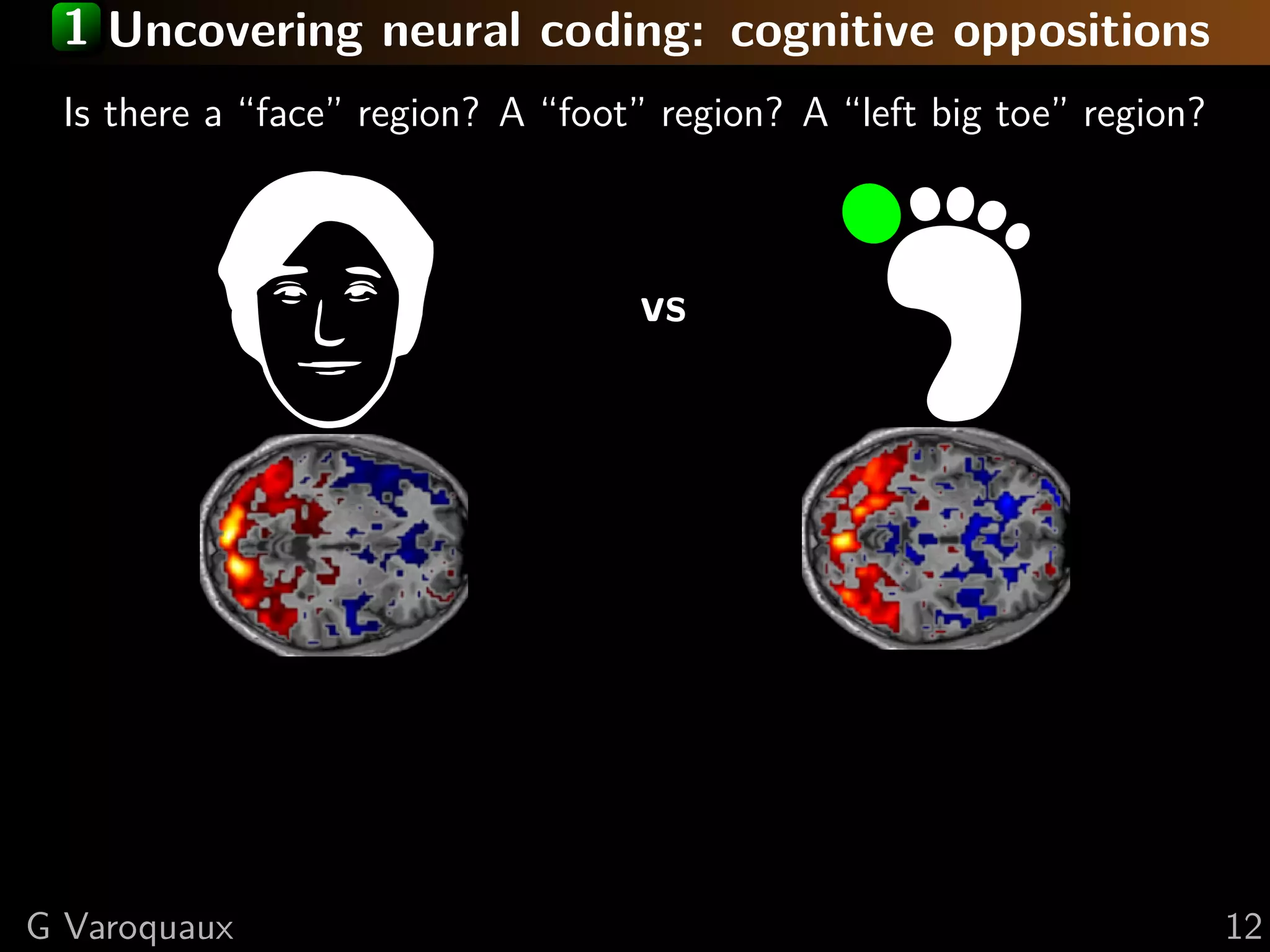
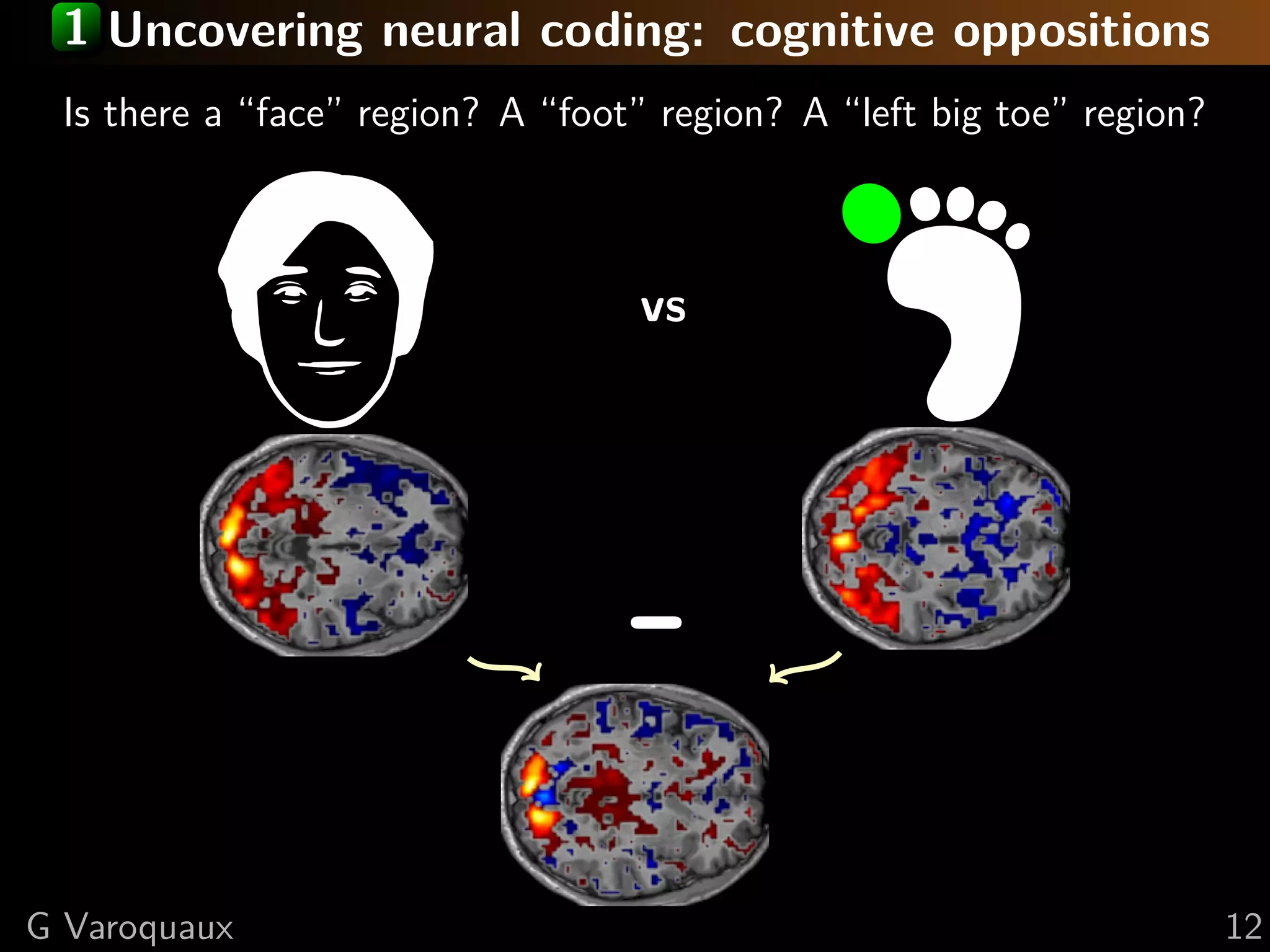
![2 Inference in cognitive neuroimaging
[Poldrack 2006, Henson 2006]
What is the neural support of a function?
What is function of a given brain module?
Reverse inference
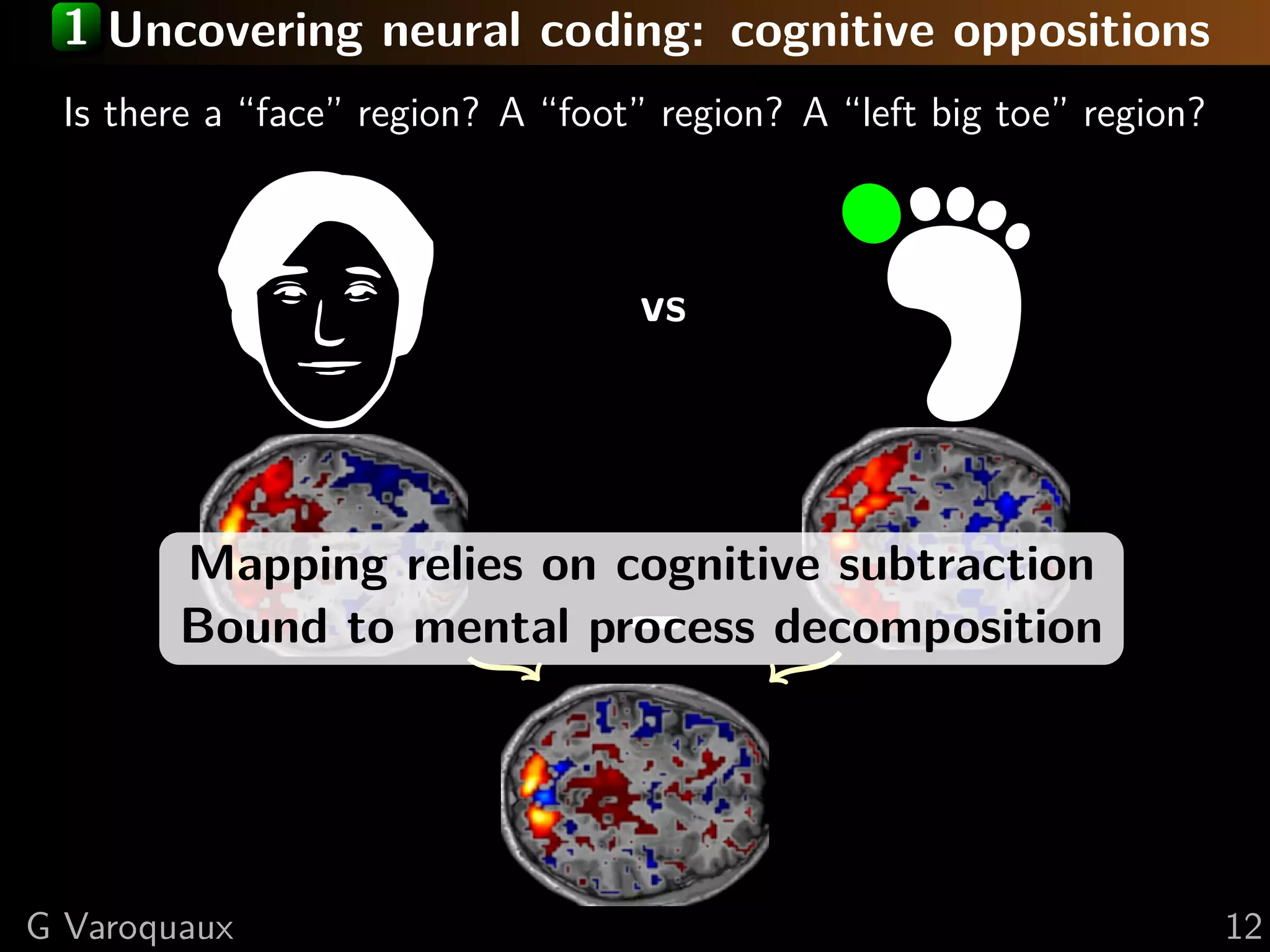
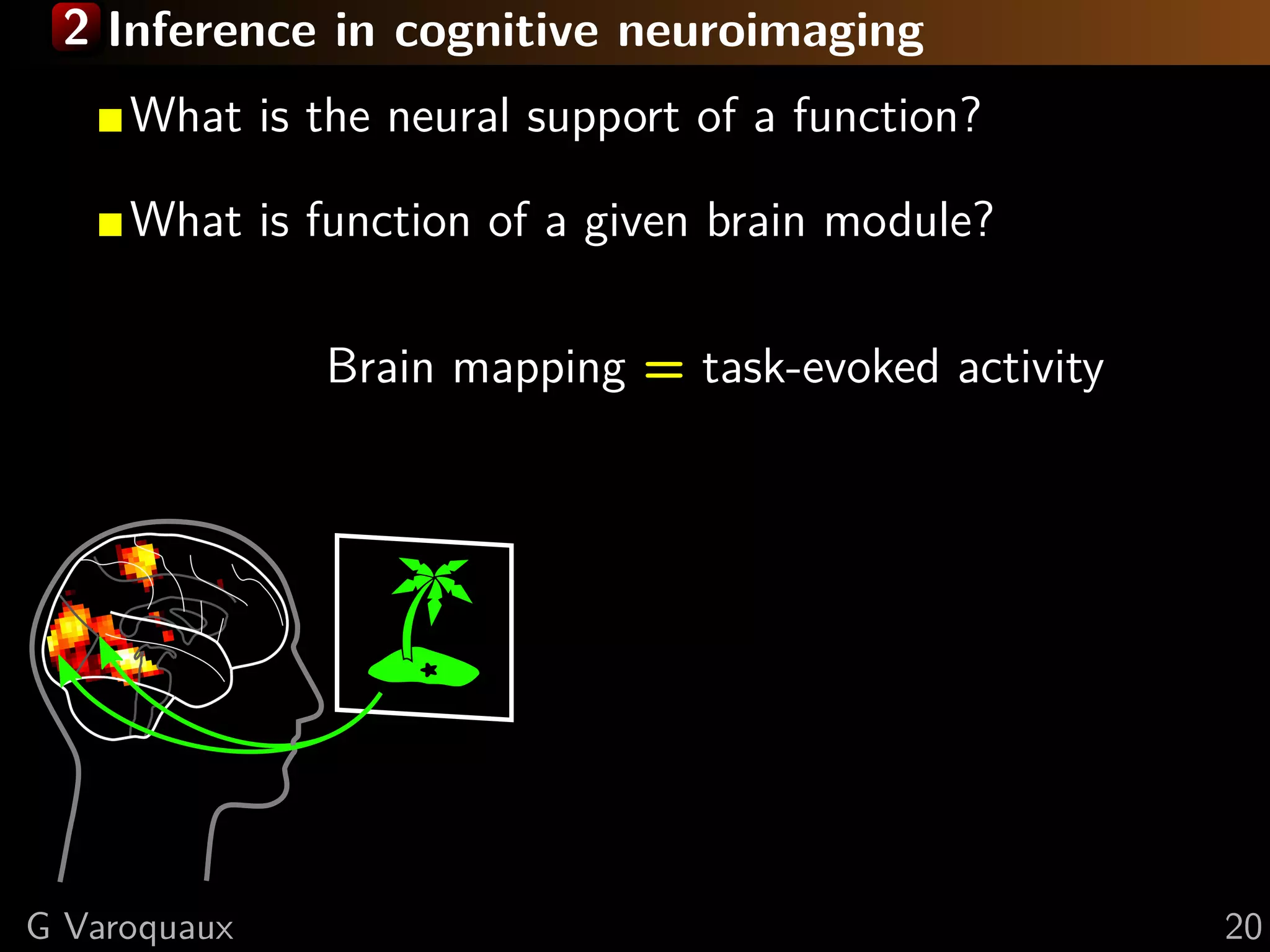
Brain mapping = task-evoked activity
+ crafting “contrasts” to isolate effects
G Varoquaux 20](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-44-2048.jpg)
![2 Inference in cognitive neuroimaging
[Kanwisher... 1997, Gauthier... 2000, Hanson and Halchenko 2008]
What is the neural support of a function?
What is function of a given brain module?
Reverse inference
Is there a face area?
G Varoquaux 20](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-45-2048.jpg)
![2 Inference in cognitive neuroimaging
[Poldrack... 2009, Schwartz... 2013]
What is the neural support of a function?
What is function of a given brain module?
Reverse inference
Decoding: Find regions that
predict observed cognition
G Varoquaux 20](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-46-2048.jpg)
![2 Decoding for reverse inference
[Poldrack... 2009, Schwartz... 2013]
Prediction = proxy for implication
Need large cognitive coverage
G Varoquaux 21](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-47-2048.jpg)
![2 Decoding for reverse inference
[Poldrack... 2009, Schwartz... 2013]
Prediction = proxy for implication
Need large cognitive coverage
Interpretation of the “grandmother neuron”
“more than a neuron re-
sponds to one concept and
[...] neurons do not neces-
sarily respond to only one
concept are given by the
data itself
[Quian Quiroga and Kreiman 2010]
G Varoquaux 21](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-48-2048.jpg)
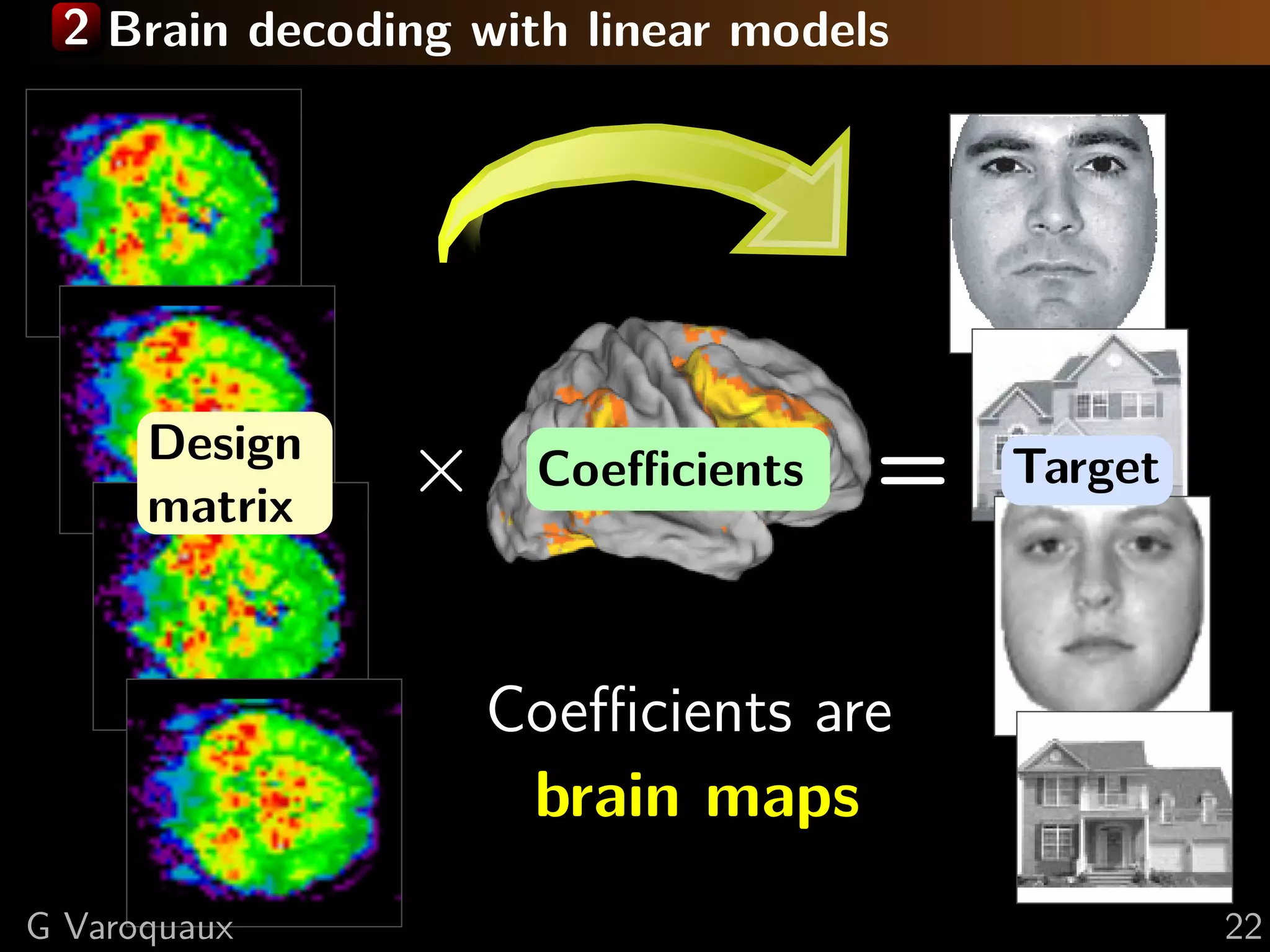
![2 Brain decoding to recover predictive regions?
Face vs house visual recognition [Haxby... 2001]
SVM
error: 26%
G Varoquaux 23](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-50-2048.jpg)
![2 Brain decoding to recover predictive regions?
Face vs house visual recognition [Haxby... 2001]
Sparse model
error: 19%
G Varoquaux 23](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-51-2048.jpg)
![2 Brain decoding to recover predictive regions?
Face vs house visual recognition [Haxby... 2001]
Ridge
error: 15%
Best predictor outlines the worst regions
Best maps predict worst
G Varoquaux 23](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-52-2048.jpg)
![2 Decoders as estimators [Gramfort... 2013]
Inverse problem
Minimize the error term:
ˆw = argmin
w
l(y − X w)
Ill-posed:
Many different w will give
the same prediction error
Choice driven by (implicit) priors of the decoder
SVM sparse ridge TV- 1
G Varoquaux 24](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-53-2048.jpg)
![2 Decoders as estimators [Gramfort... 2013]
Inverse problem
Minimize the error term:
ˆw = argmin
w
l(y − X w)
Ill-posed:
Many different w will give
the same prediction error
Choice driven by (implicit) priors of the decoder
SVM sparse ridge TV- 1
Inferences rely, explicitely or implicitely,
on the regions estimated by the decoder
G Varoquaux 24](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-54-2048.jpg)





![@GaelVaroquaux
Machine learning for cognitive neuroimaging
The description of cognition is hard ⇒ Encoding
Decoding as an omnibus test
Decoding for reverse inference
Estimation of predictive regions is difficult
Software: nilearn
In Python
http://nilearn.github.io
ni
[Varoquaux and Thirion 2014]
How machine learning is
shaping cognitive neuroimaging](https://image.slidesharecdn.com/slides-160626154057/75/Machine-learning-and-cognitive-neuroimaging-new-tools-can-answer-new-questions-60-2048.jpg)




