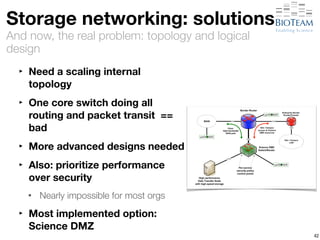
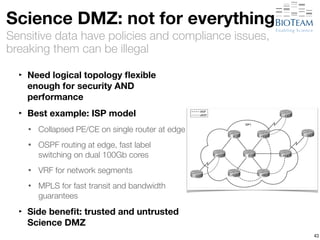
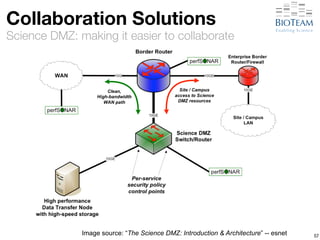
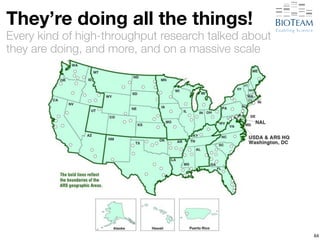
The document discusses the challenges and advancements in high-performance networking within life sciences, emphasizing the rapid increase of big data and its implications for research infrastructure. It highlights the urgency for improved IT solutions and networking capabilities to manage vast amounts of data generated by modern technologies like next-generation sequencing. Additionally, it explores the need for collaboration and sharing, as well as potential solutions such as the Science DMZ and cloud computing to facilitate data movement and accessibility.
























































![[Hyper-]convergence
It’s what we do
72
Laboratory Knowledge](https://image.slidesharecdn.com/20141028-berman-lifesciusecase-141027220919-conversion-gate02/85/High-Performance-Networking-Use-Cases-in-Life-Sciences-90-320.jpg)
![[Hyper-]convergence
It’s what we do
72
Laboratory Knowledge
Converged Solution](https://image.slidesharecdn.com/20141028-berman-lifesciusecase-141027220919-conversion-gate02/85/High-Performance-Networking-Use-Cases-in-Life-Sciences-91-320.jpg)
![[Hyper-]convergence
It’s what we do
72
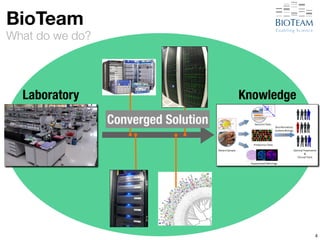
Laboratory Knowledge
Converged Solution](https://image.slidesharecdn.com/20141028-berman-lifesciusecase-141027220919-conversion-gate02/85/High-Performance-Networking-Use-Cases-in-Life-Sciences-92-320.jpg)