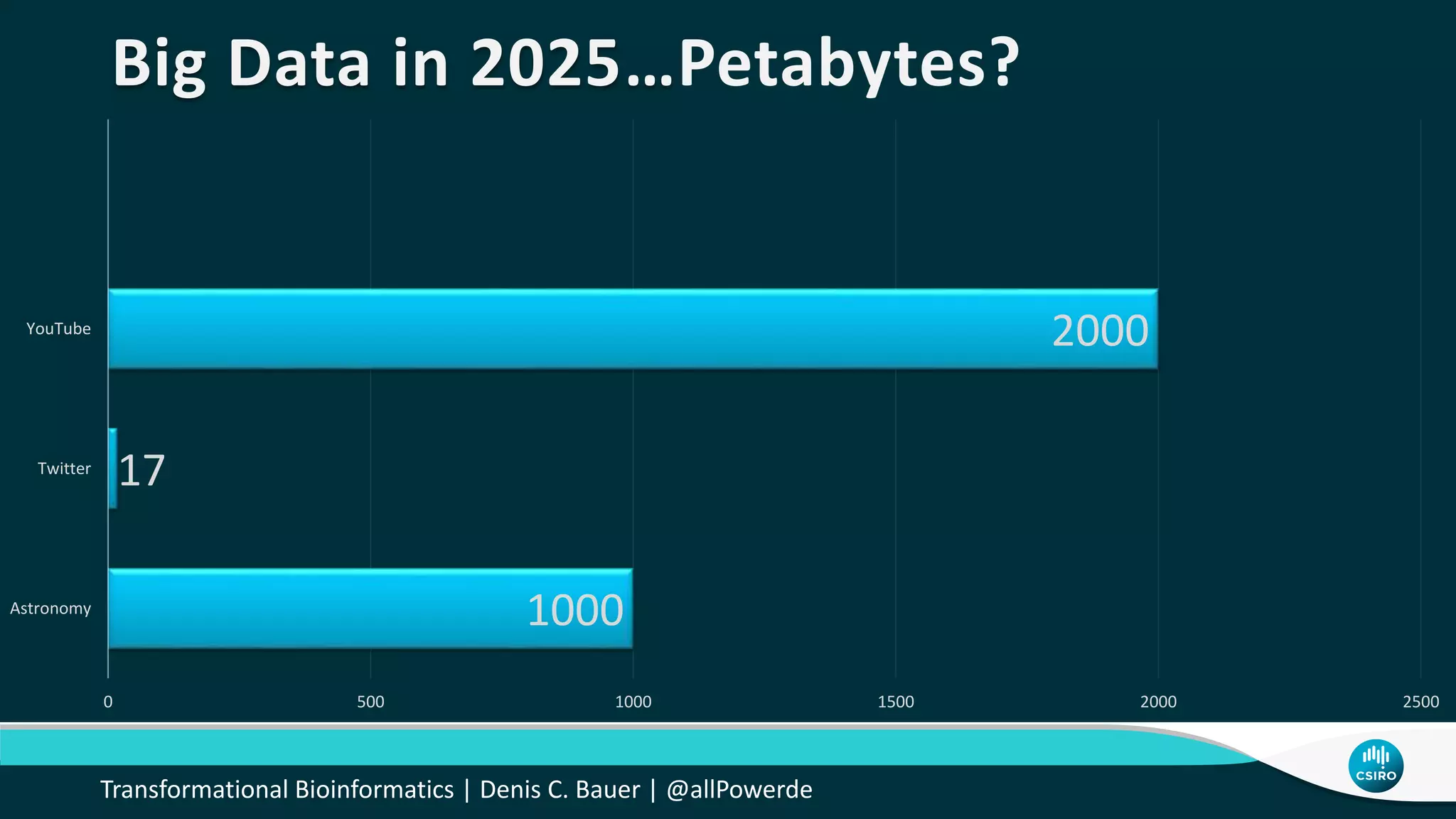
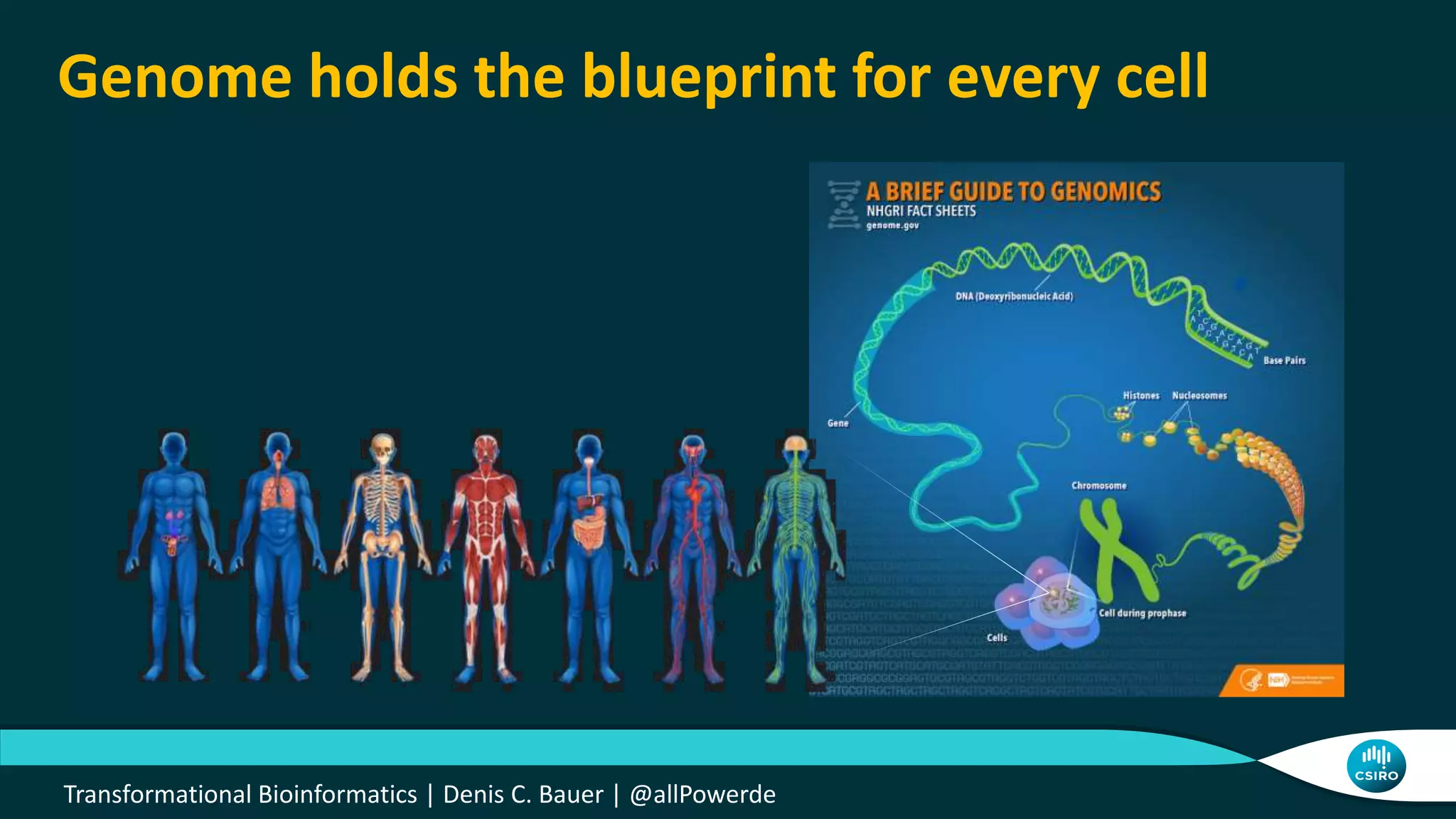
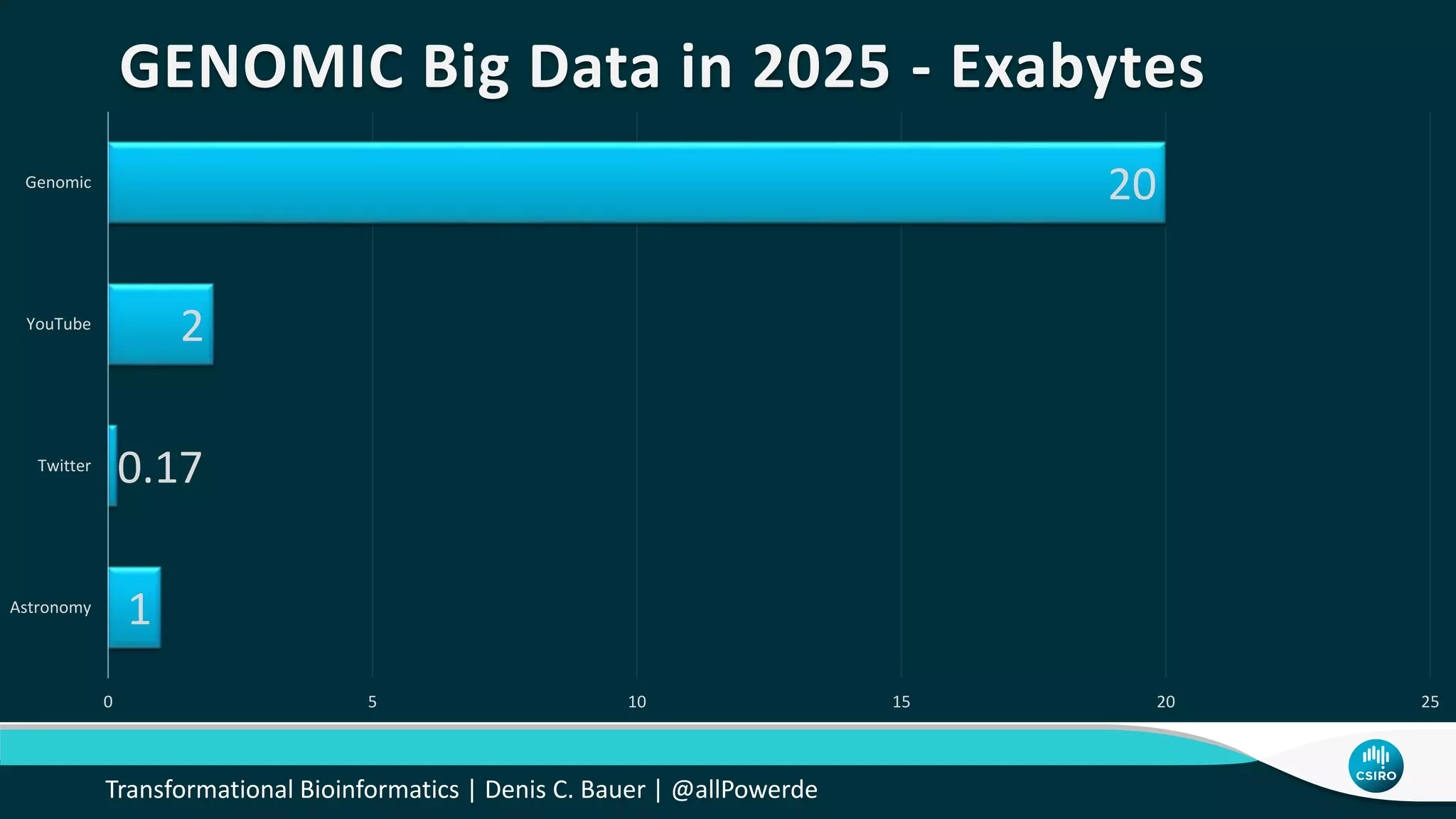
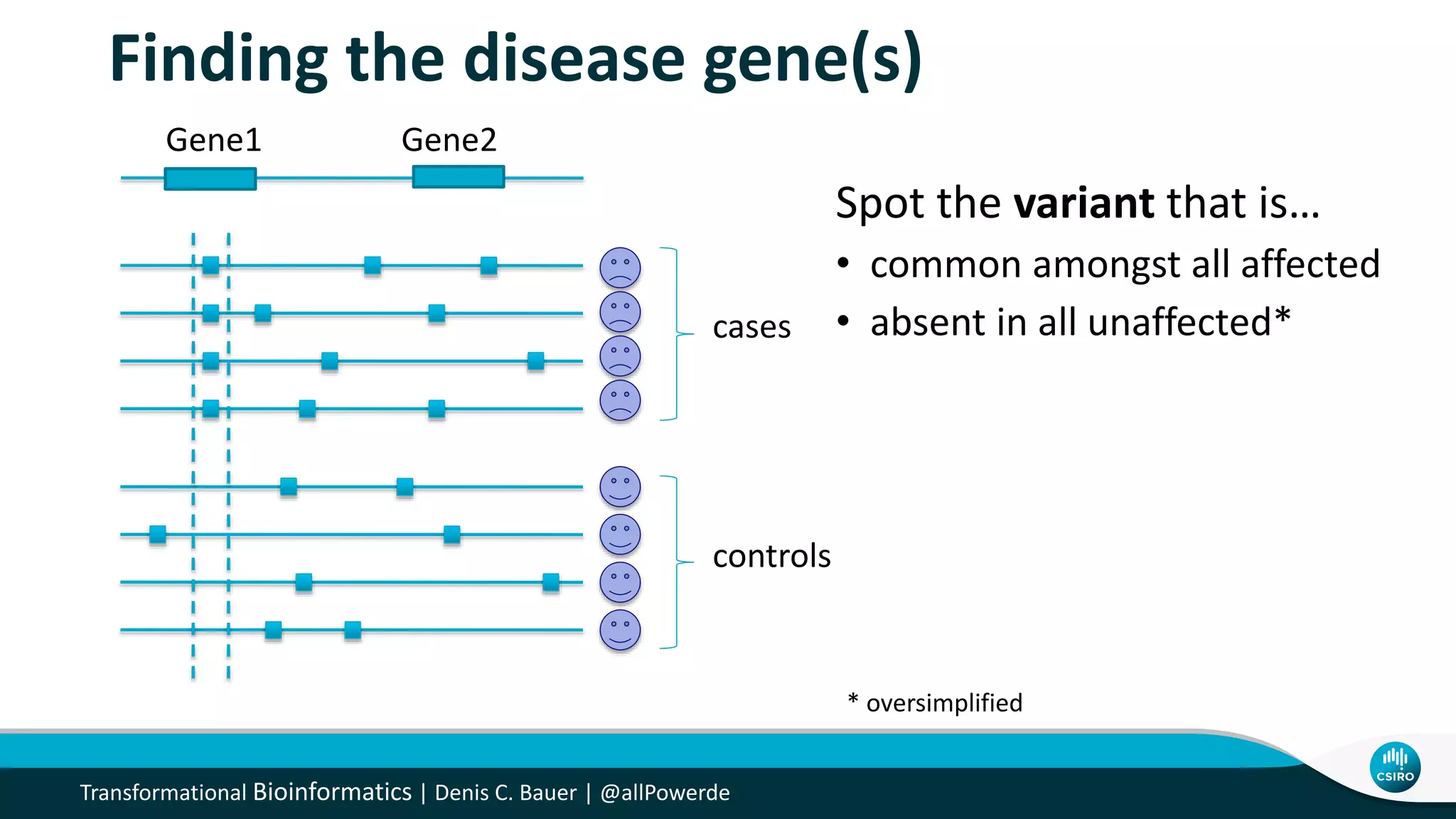
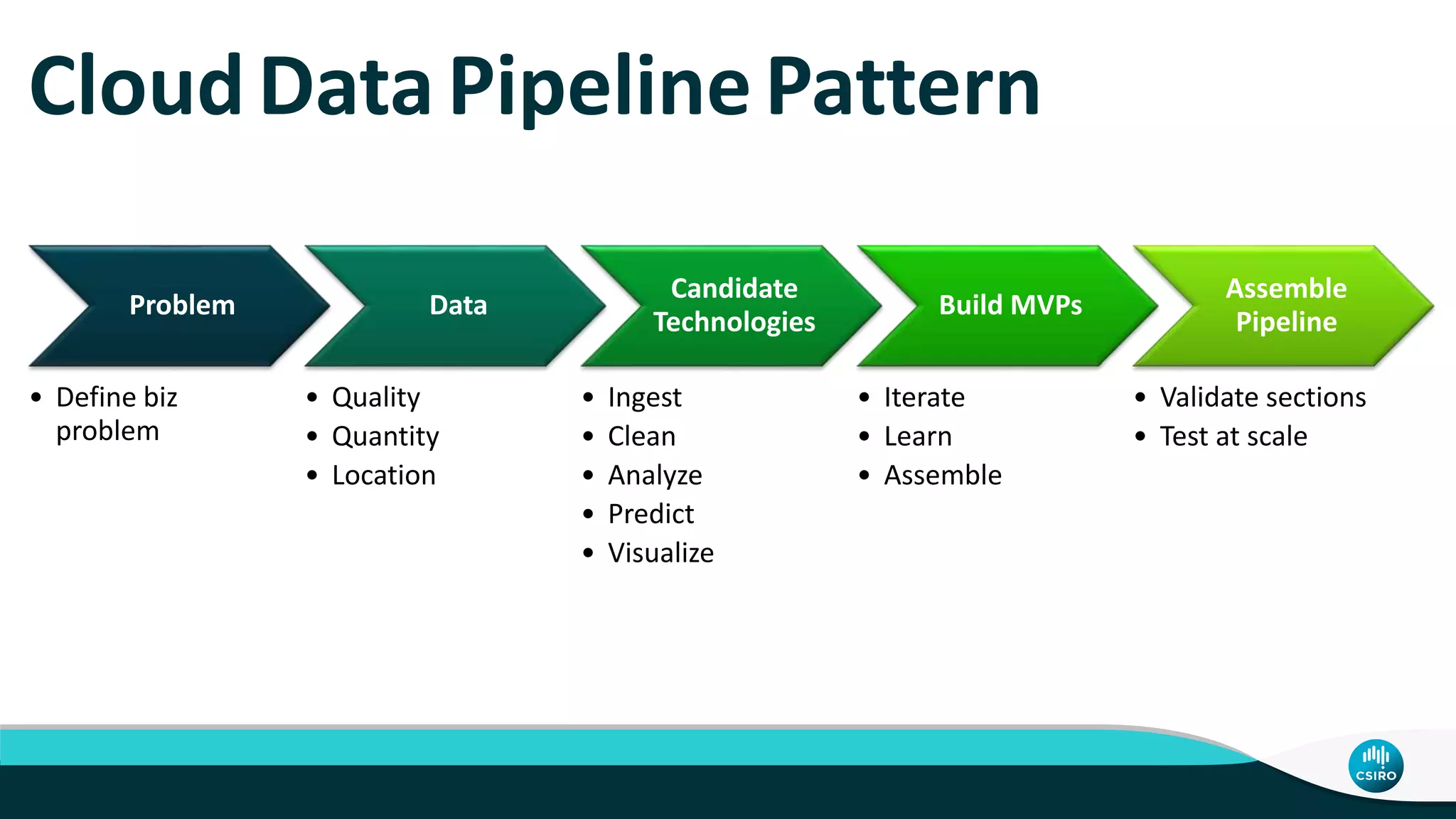
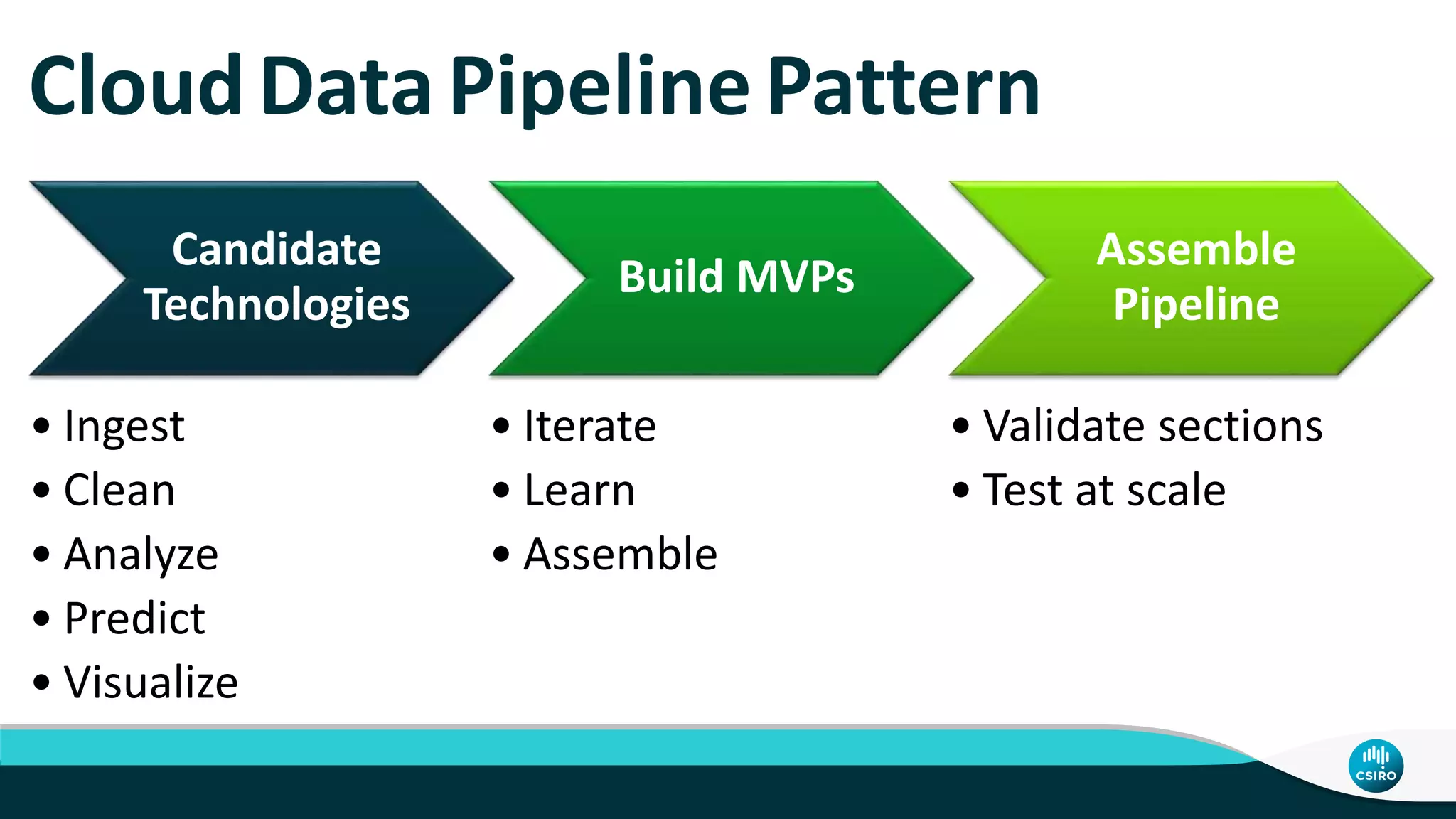
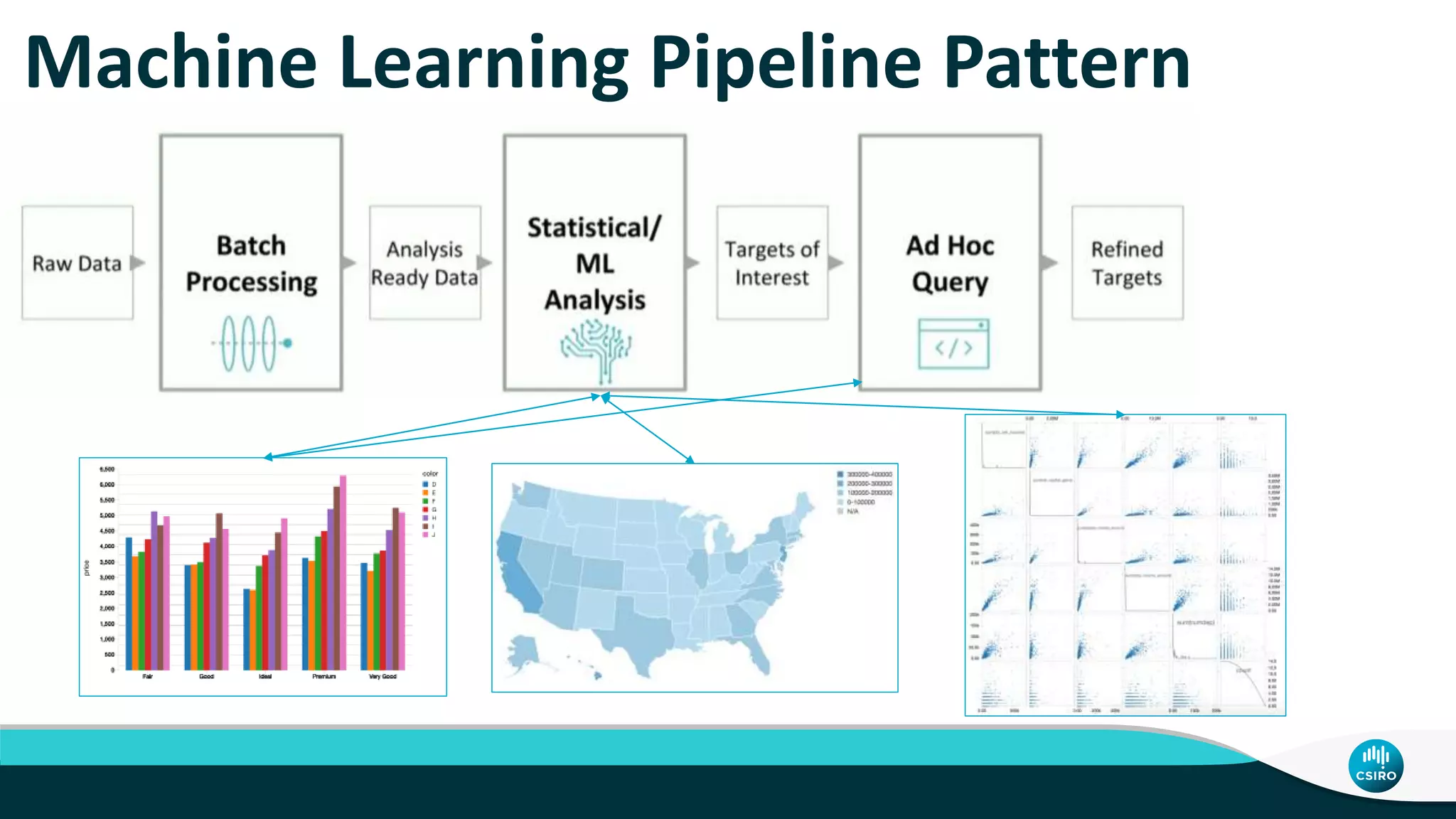
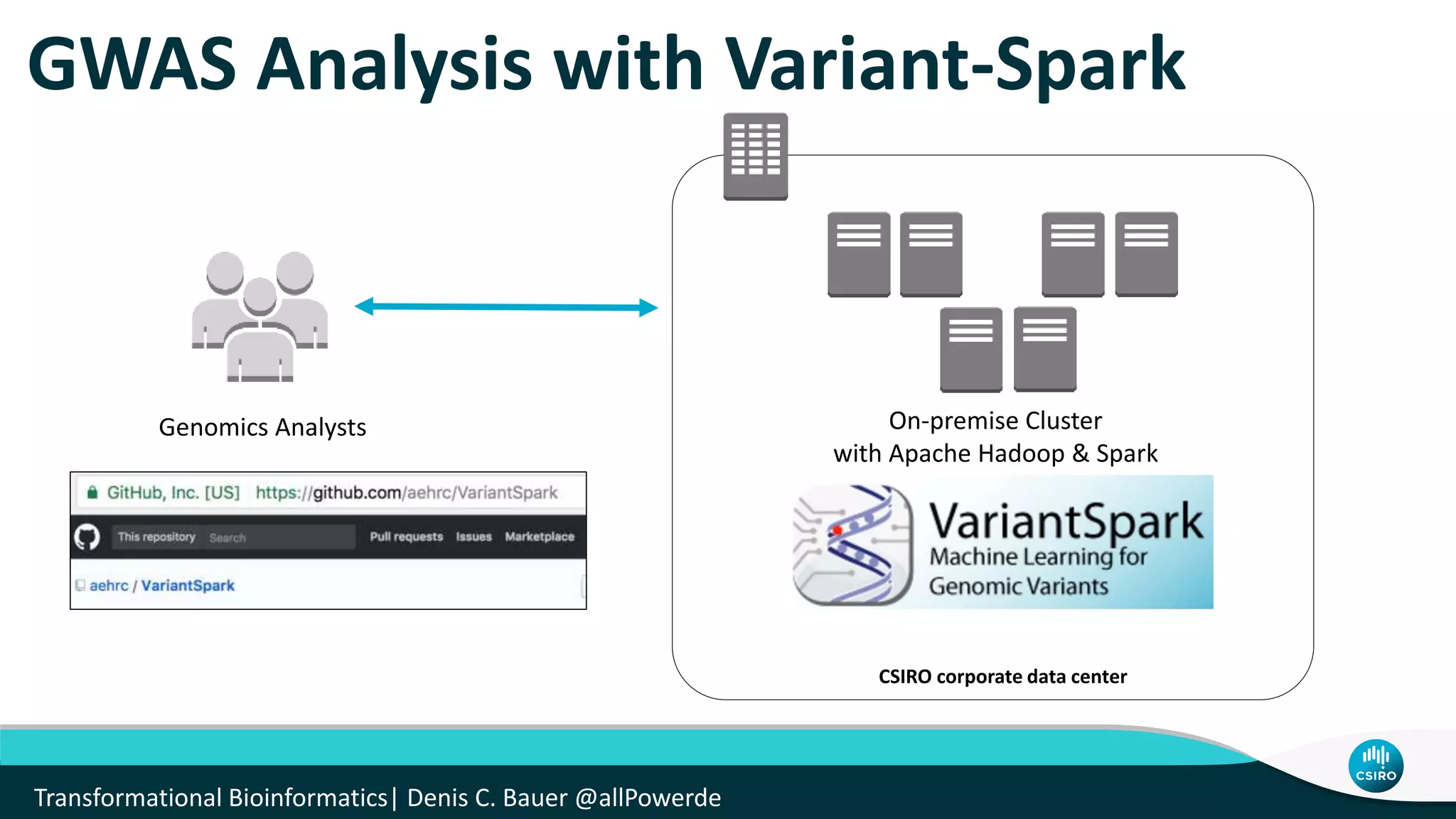
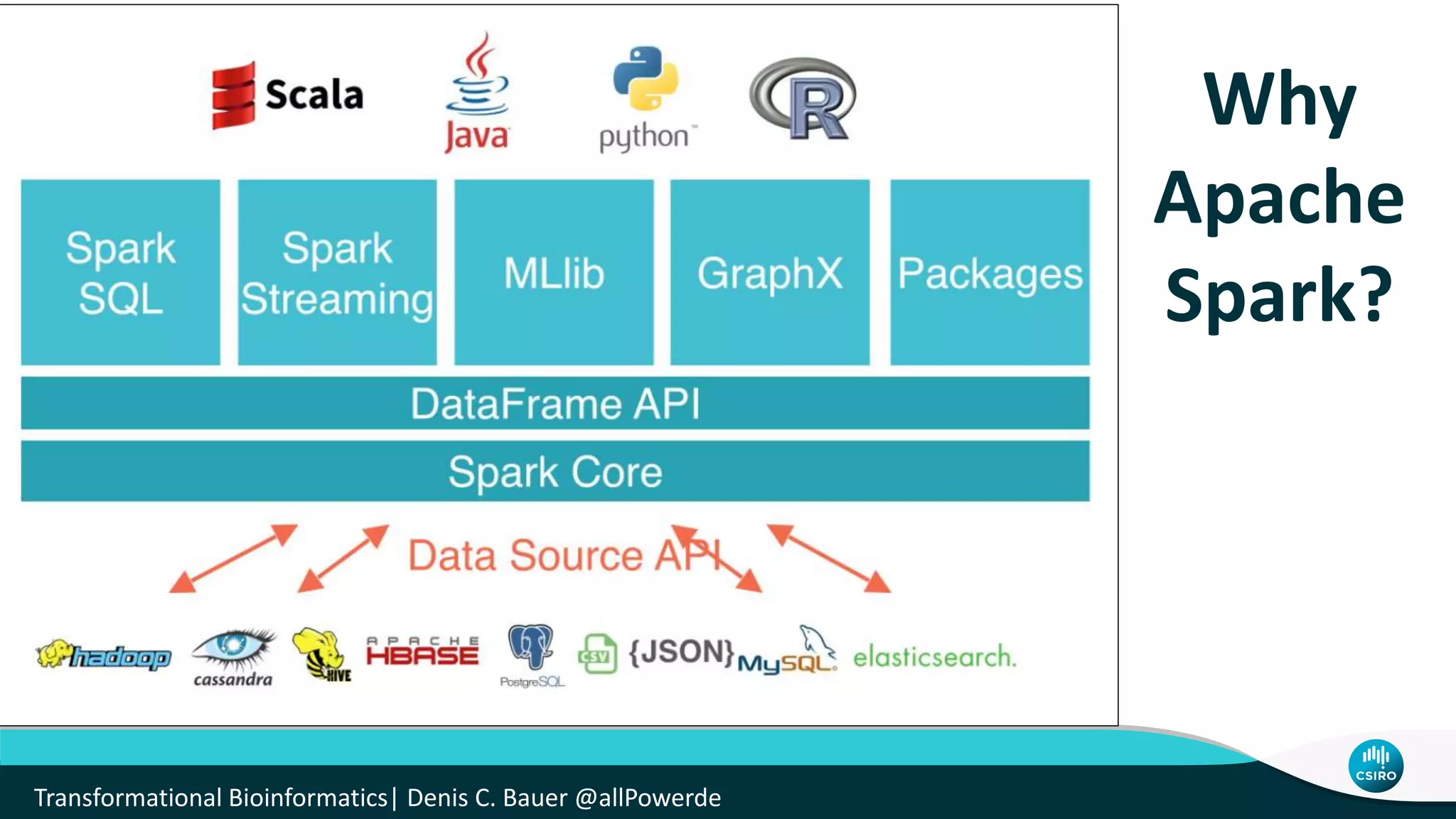
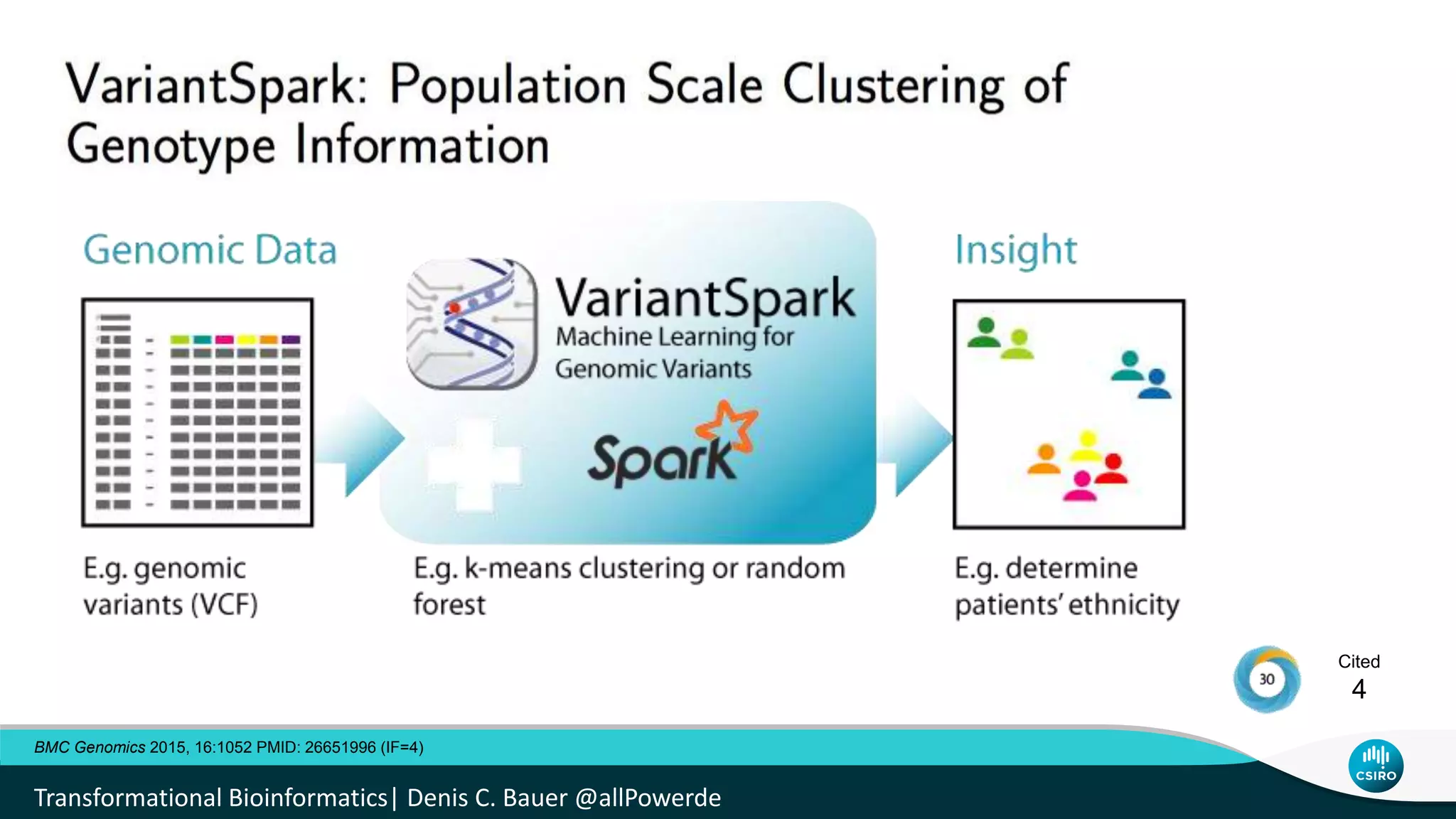
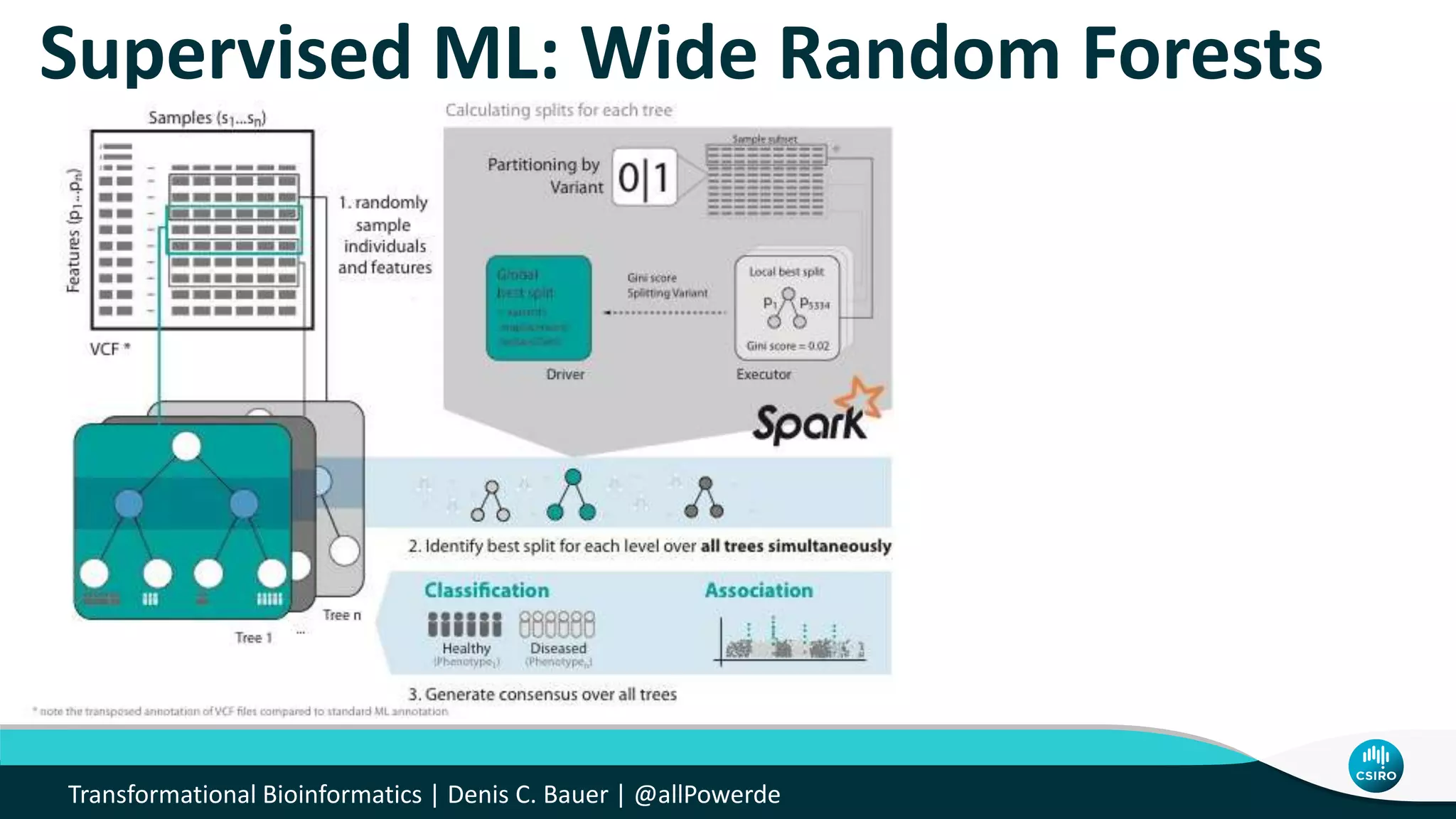
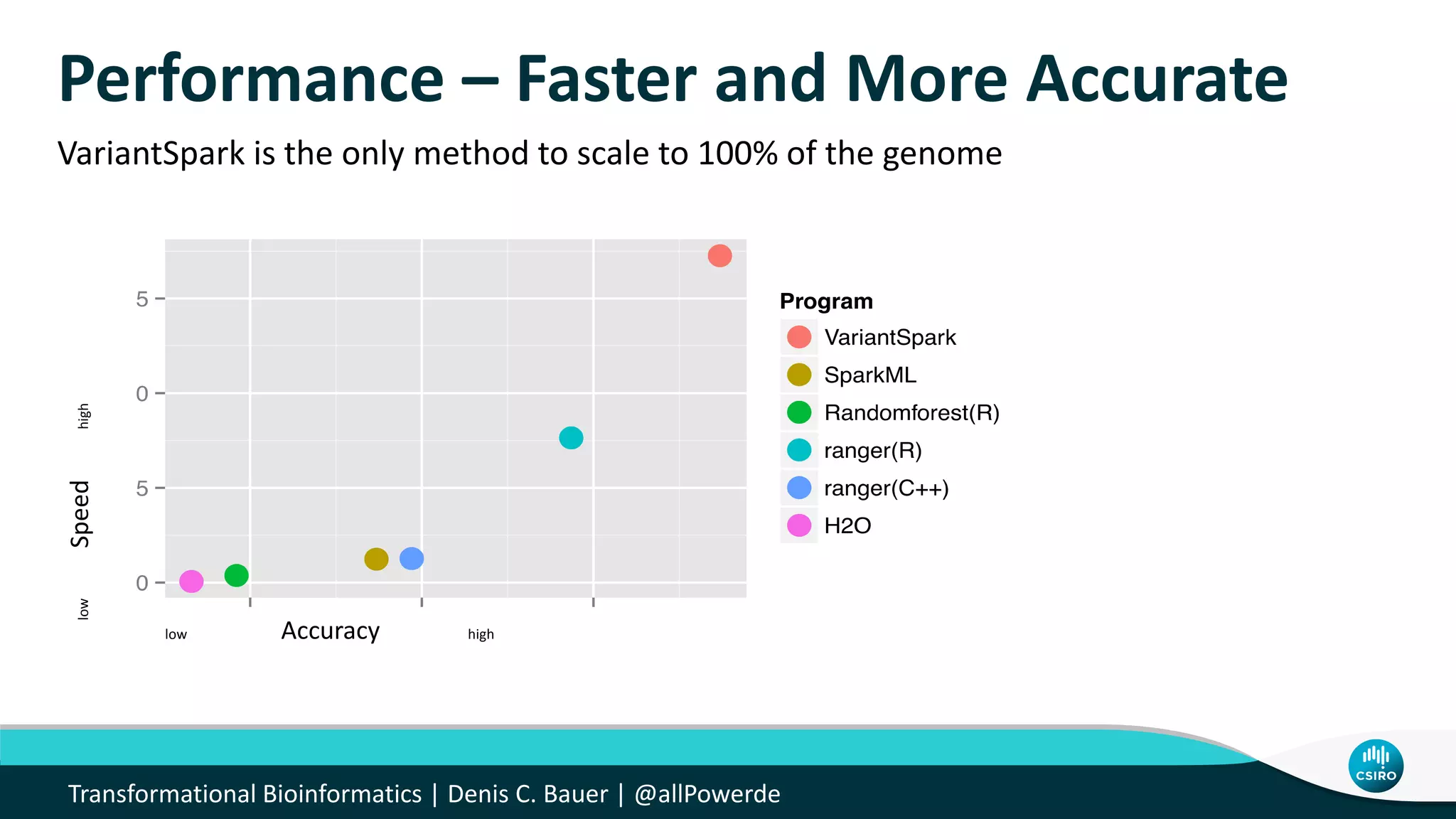
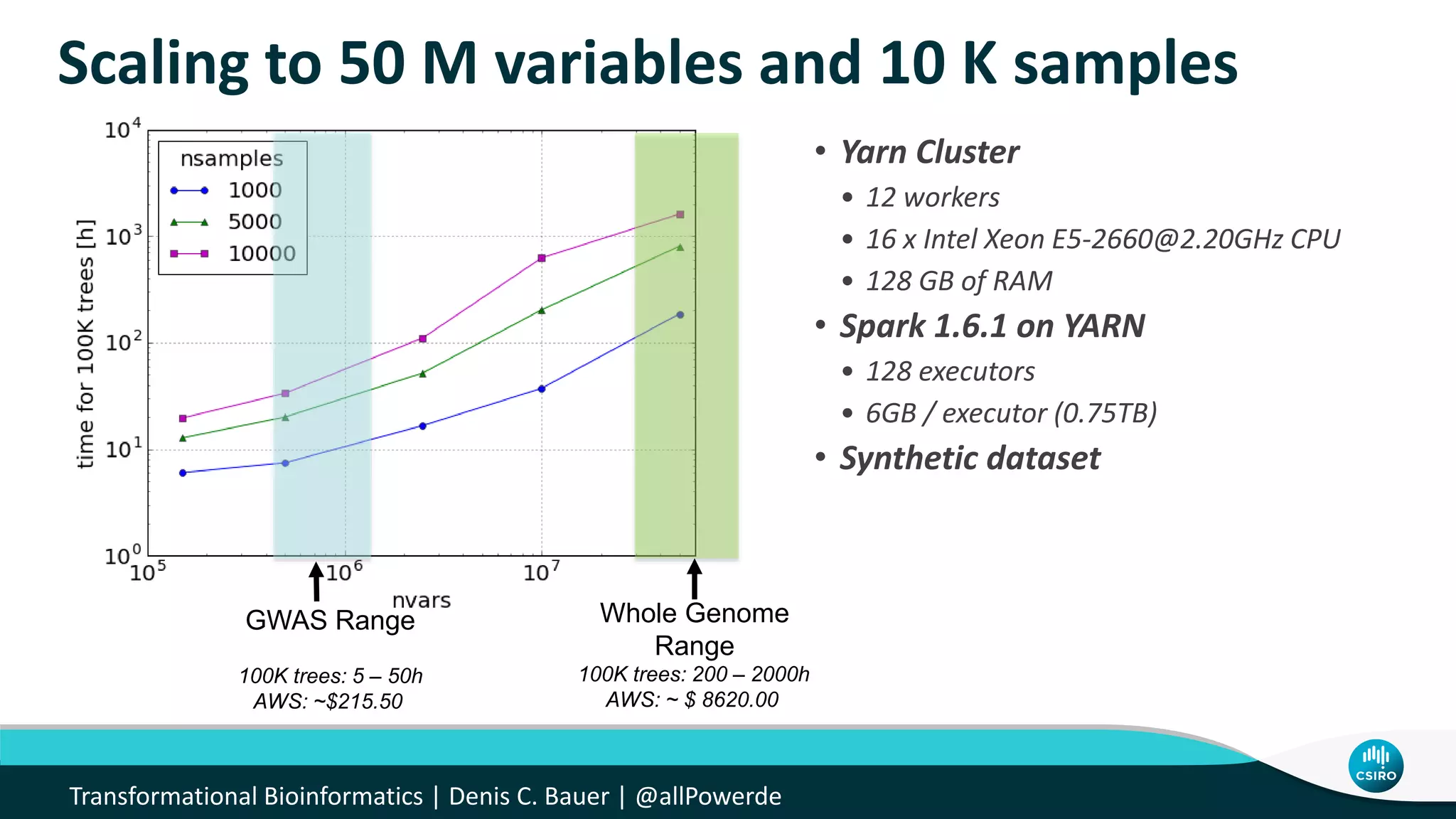
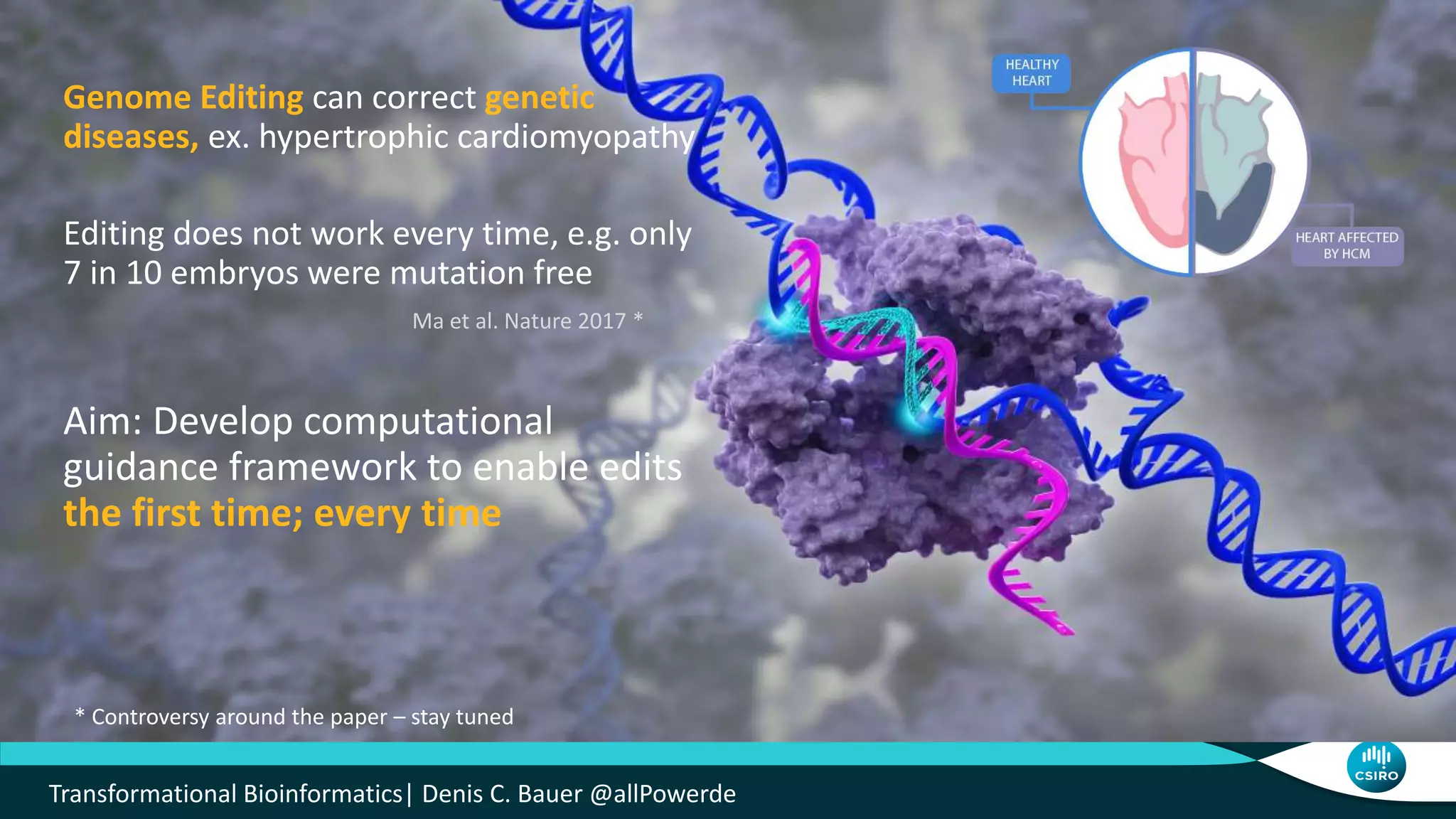
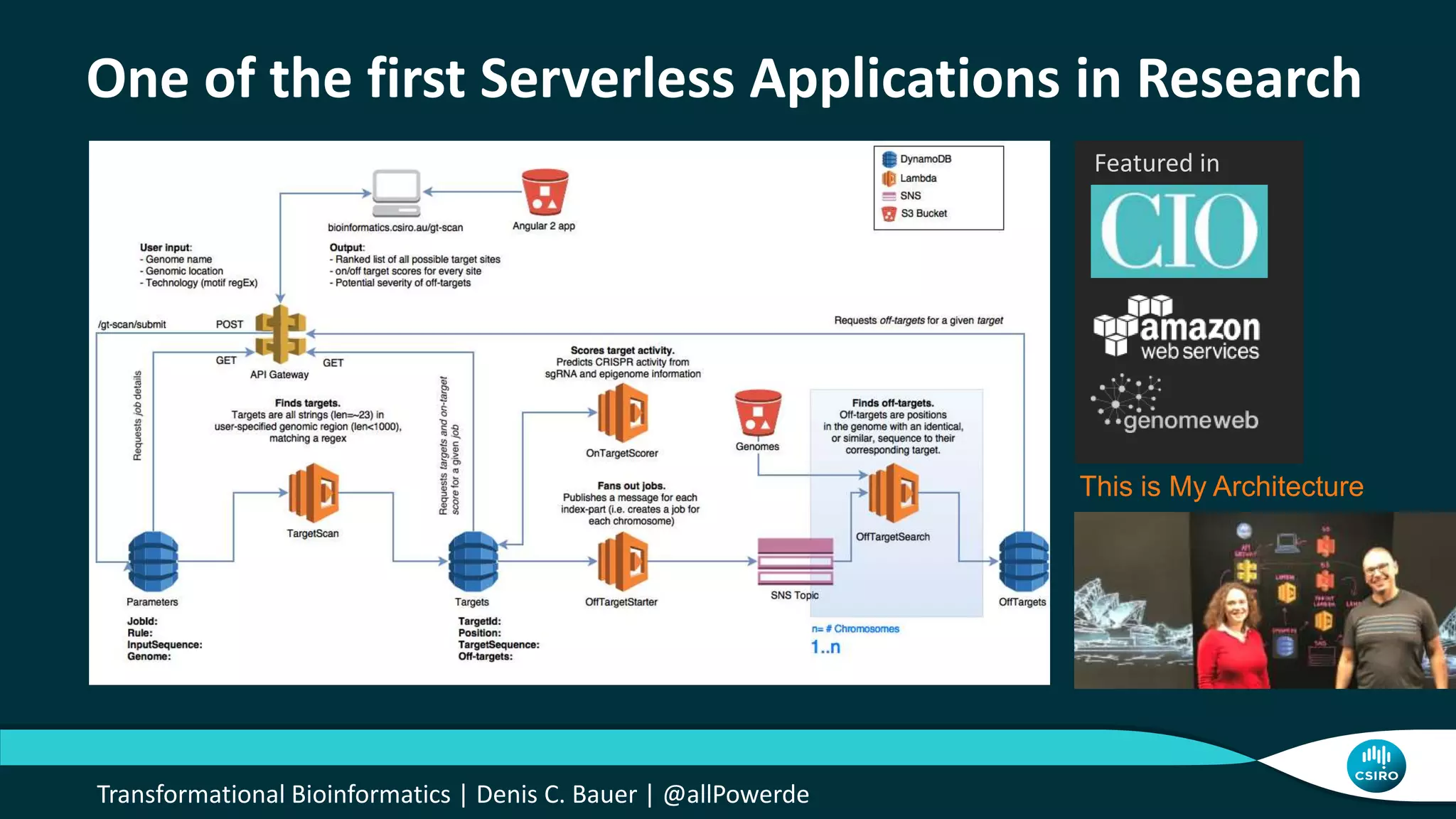
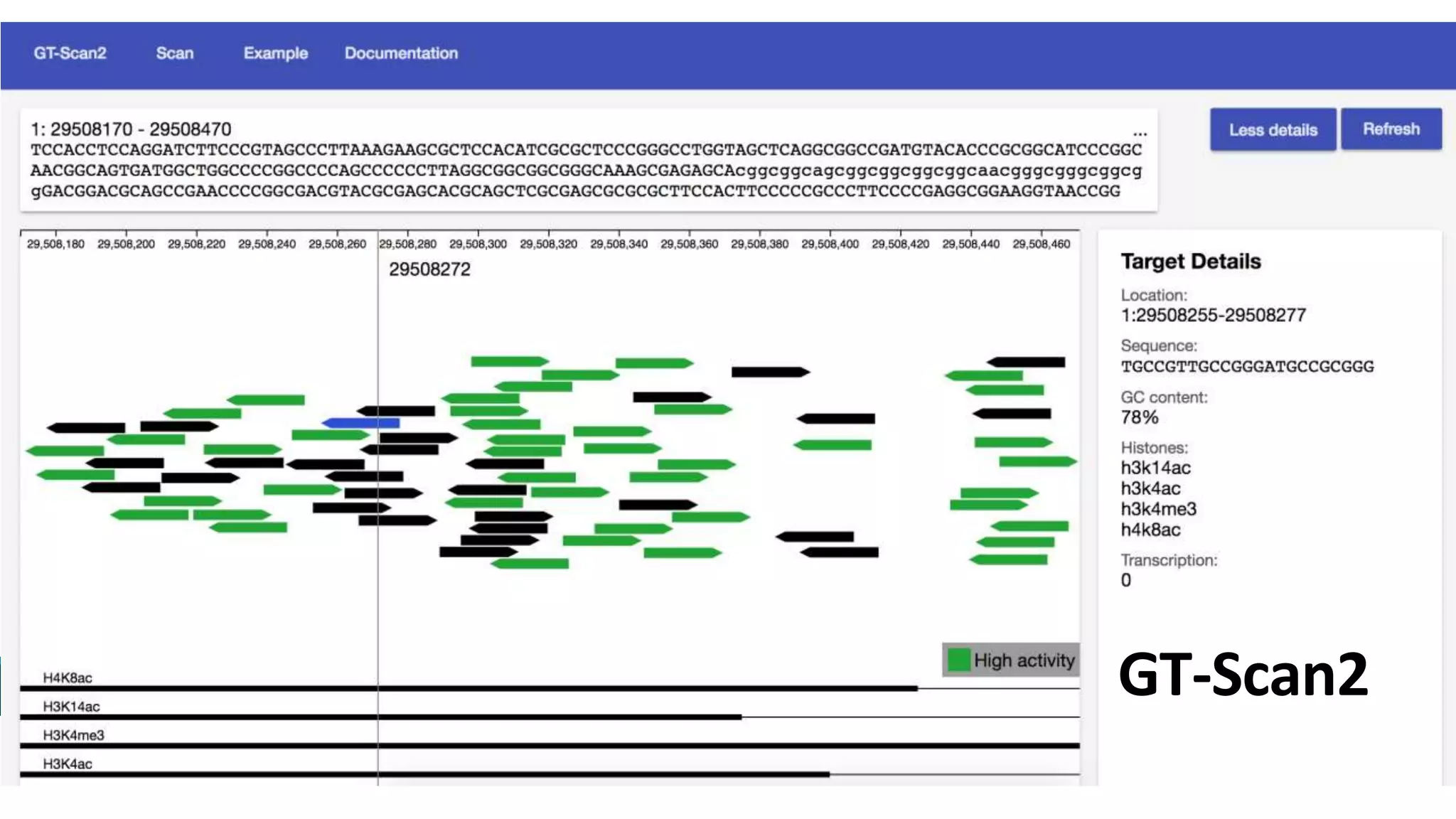
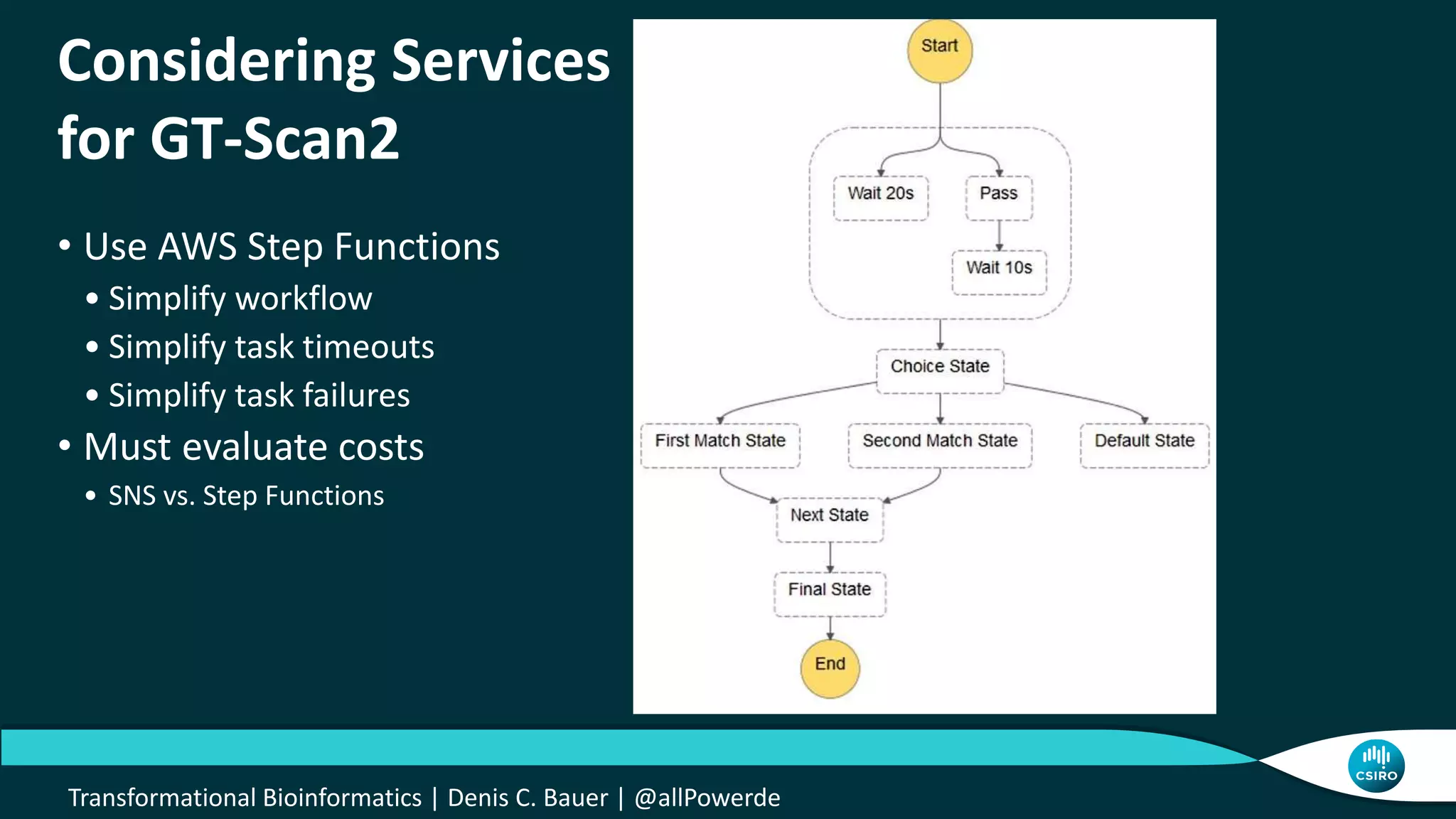
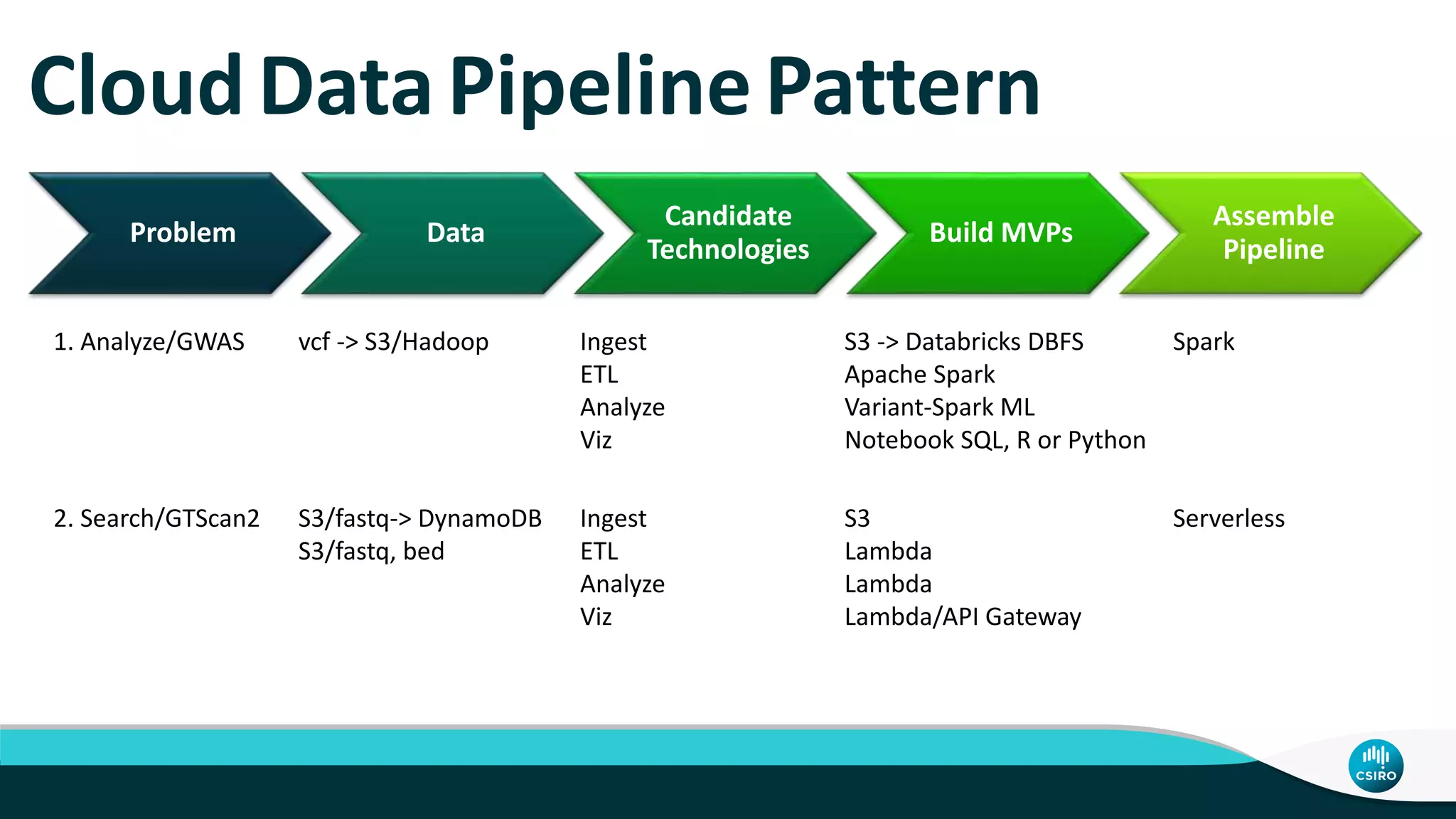
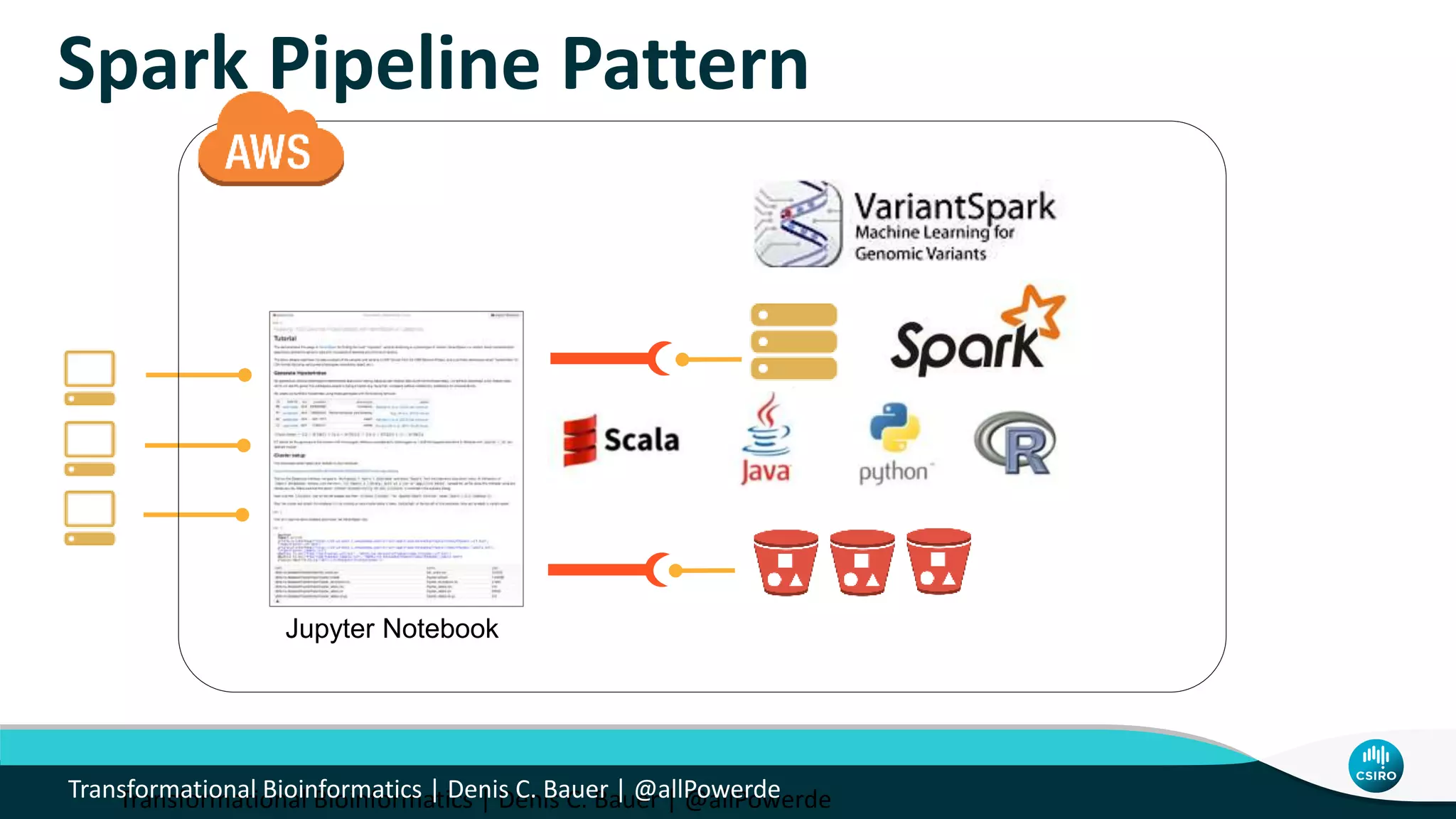
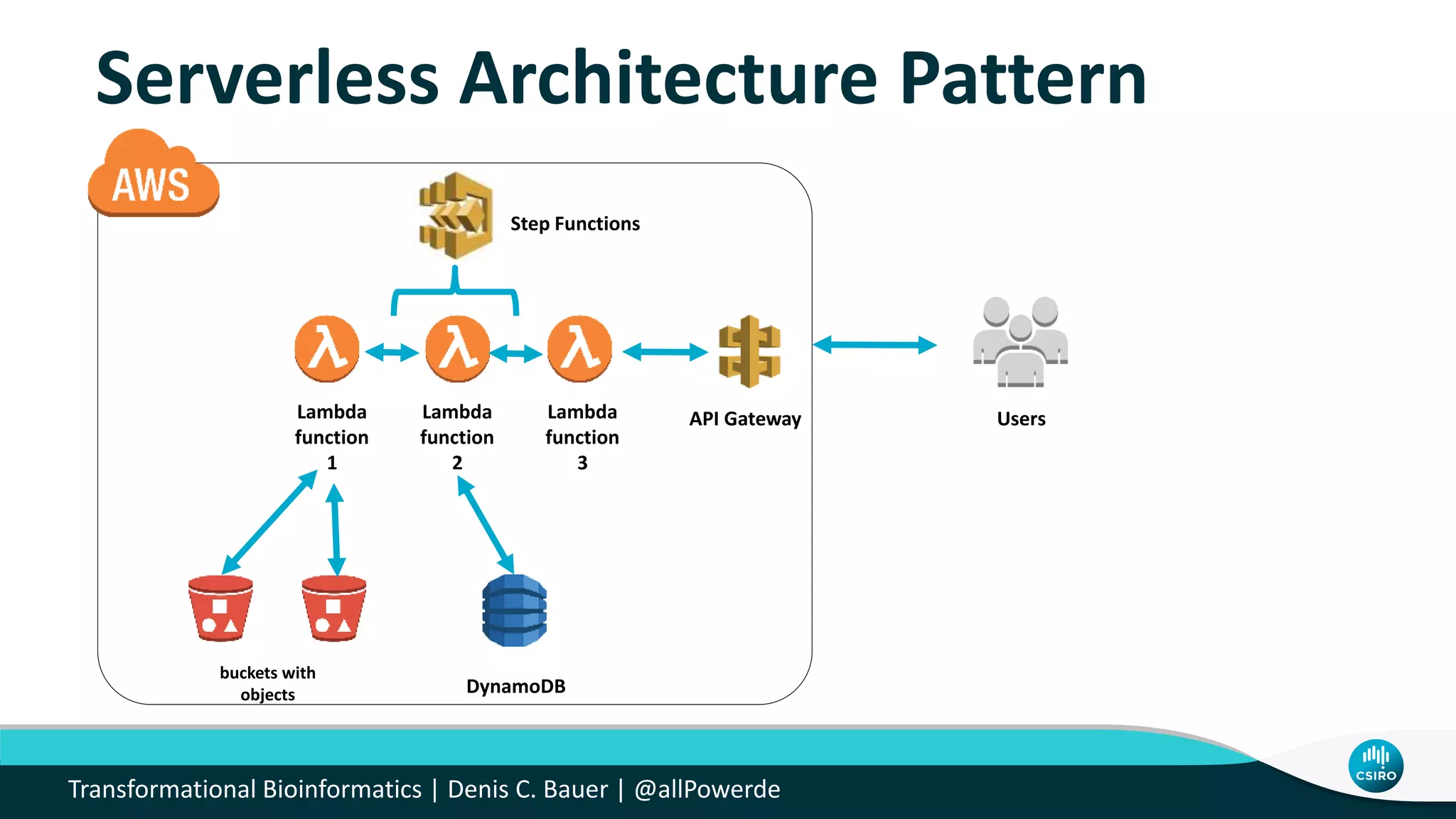
The document discusses the advancements in genomic-scale data pipelines and their implications for bioinformatics, focusing on cloud-based solutions like Apache Hadoop and Apache Spark for efficient data analysis and machine learning. It presents methodologies for genetic research, including identifying disease-related variants and genome editing technologies. Additionally, the document highlights the need for scalable and parallelized processes to handle increasing genomic data demands in research.