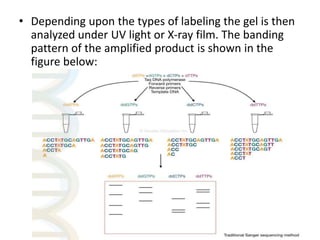
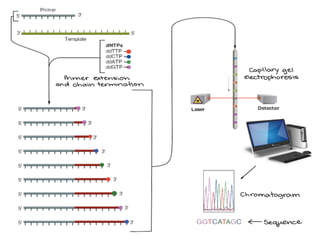
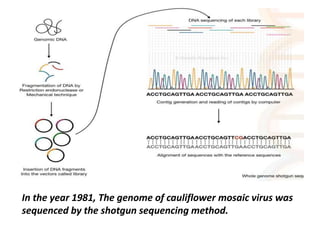
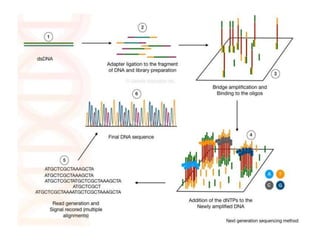
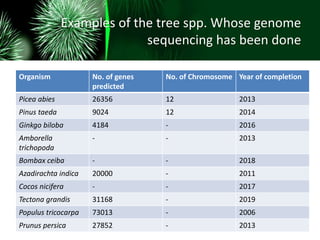
This document provides an overview of gene sequencing. It begins with an introduction to DNA structure and the goal of gene sequencing to determine the sequence of nucleotides. It then discusses the history of gene sequencing, from early RNA sequencing to the development of Sanger and next-generation sequencing methods. The document outlines the general steps in gene sequencing and describes several specific methods, including Maxam-Gilbert, Sanger, pyrosequencing, and shotgun sequencing. It concludes by reviewing applications of gene sequencing and providing examples of species whose genomes have been sequenced.


![Intoduction
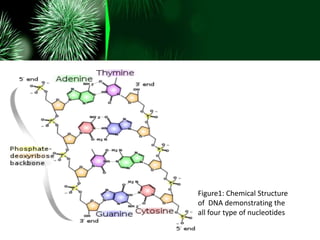
• DNA is composed of simpler monomeric units called
nucleotides.
• Each nucleotide is composed of one of four nitrogen-
containing nucleobases (cytosine [C], guanine [G],
adenine [A] or thymine [T]), a sugar called
deoxyribose, and a phosphate group.
• DNA sequencing is the process of determining the
sequence of these nucleotide bases (As, Ts, Cs, and
Gs) in a piece of DNA.](https://image.slidesharecdn.com/genesequencing-240112141034-b1156ea7/85/Gene-sequencing-steps-involved-methods-used-and-applications-pptx-3-320.jpg)