This document provides information on enzymes. It begins by defining enzymes as soluble, colloidal organic catalysts formed by living cells that are specific, protein in nature, and inactive at 0°C. It notes that all enzymes are proteins and that activity is lost through denaturation or dissociation. The document discusses enzymes as proteins that often require cofactors. It provides details on different types of enzymes based on their structure, location, and method of secretion. The rest of the document covers enzyme nomenclature, classification, specificity, and mechanisms of action. It discusses factors that affect enzyme activity such as temperature, pH, product concentration, and more.

























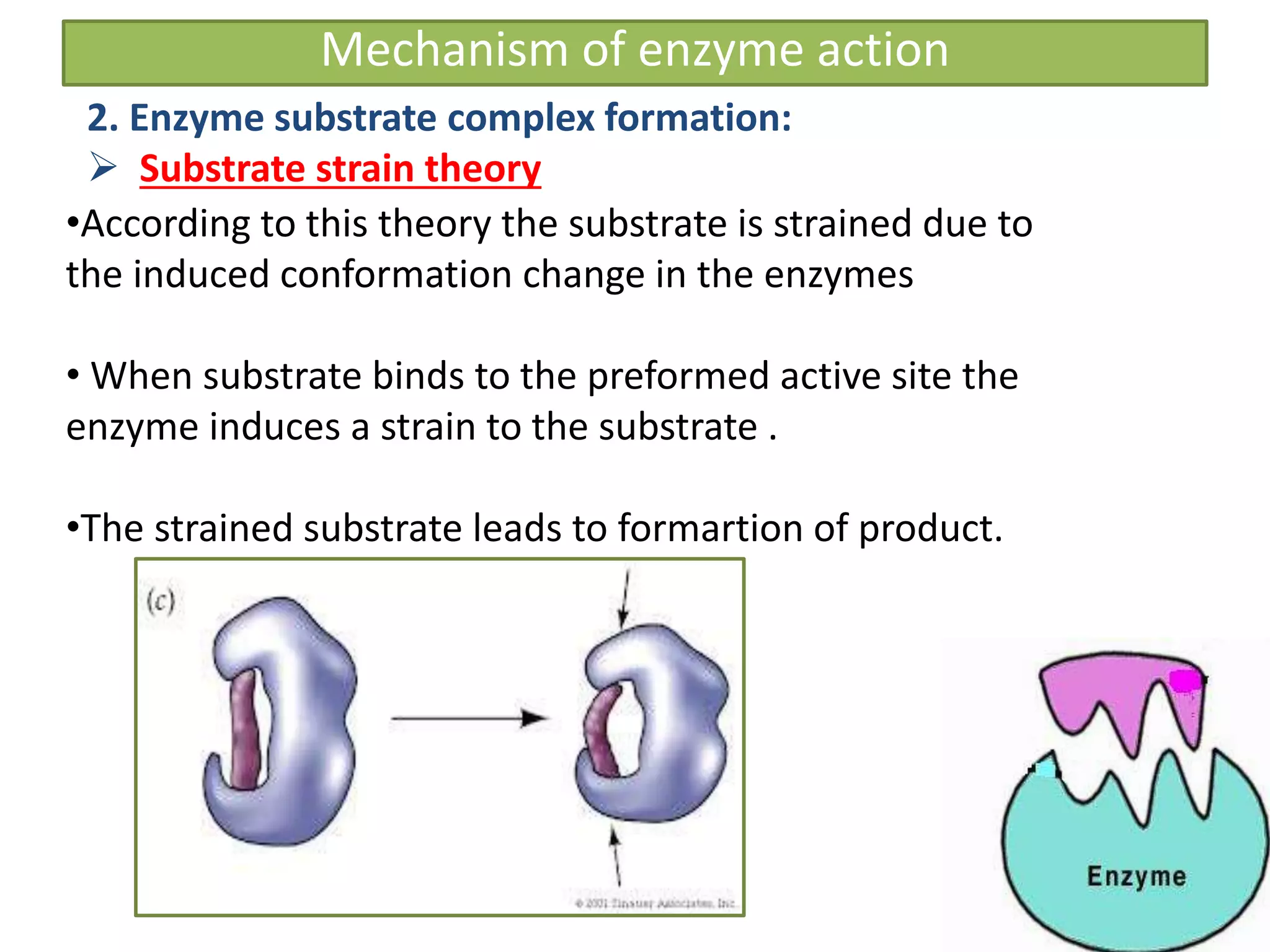

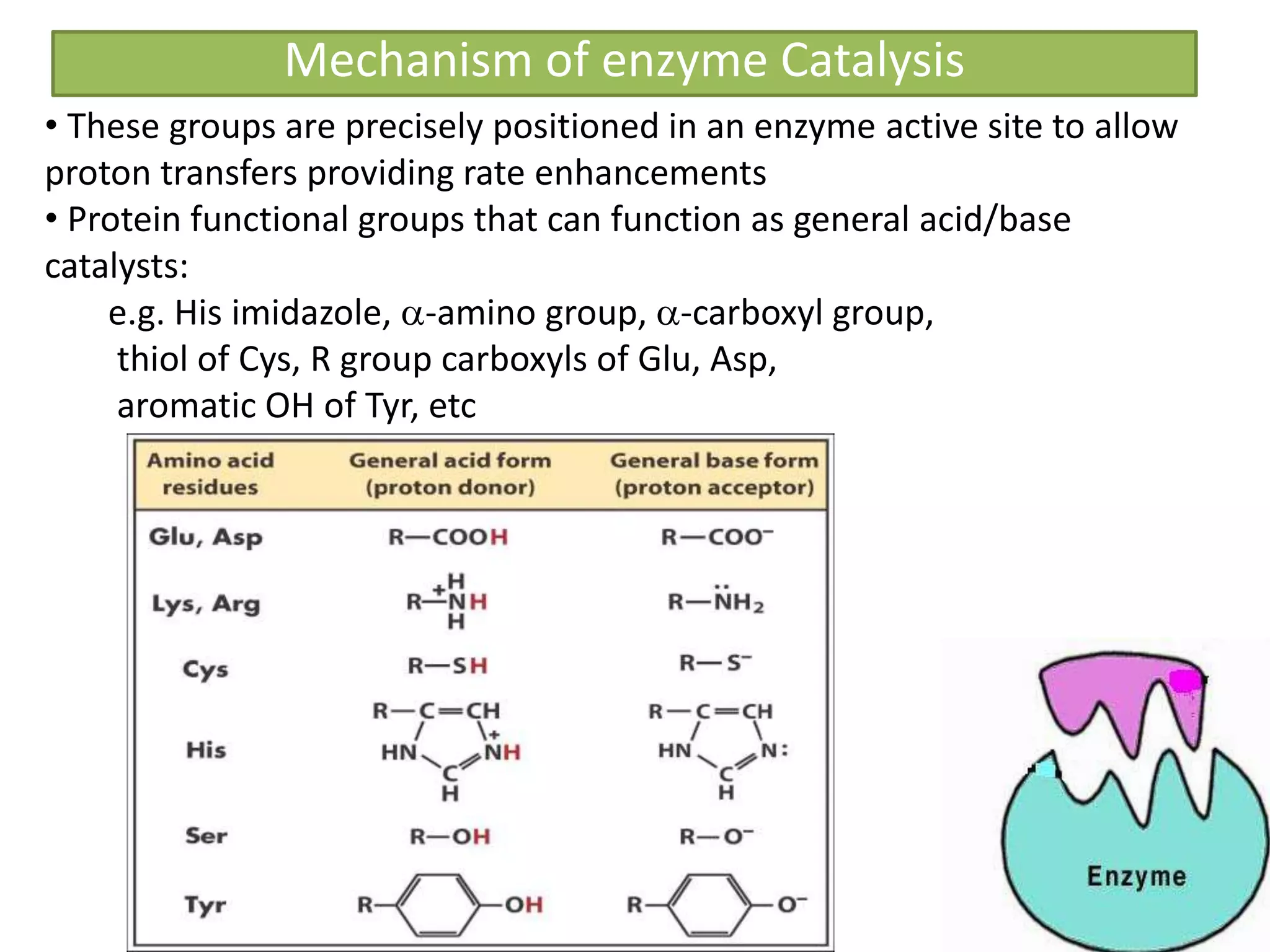















![Significance of KM
When V= ½ Vmax, what is [S]?](https://image.slidesharecdn.com/enzymesandenzymeinhibition-190509102317/75/Enzymes-and-enzyme-inhibition-48-2048.jpg)






















![Enzyme Inhibition (Mechanism)
I
I
S
S
S I
I
I II
S
Competitive Non-competitive Uncompetitive
E
E
Different site
Compete for
active siteInhibitor
Substrate
CartoonGuideEquationandDescription
[I] binds to free [E] only,
and competes with [S];
increasing [S] overcomes
Inhibition by [I].
[I] binds to free [E] or [ES]
complex; Increasing [S] can
not overcome [I] inhibition.
[I] binds to [ES] complex
only, increasing [S] favors
the inhibition by [I].
E + S → ES →E + P
+
I
↓
EI
←
↑
E + S → ES →E + P
+ +
I I
↓ ↓
EI+ S→EIS
←
↑ ↑
E + S → ES →E + P
+
I
↓
EIS
←
↑
E
I
S
Juang RH (2004) BCbasics](https://image.slidesharecdn.com/enzymesandenzymeinhibition-190509102317/75/Enzymes-and-enzyme-inhibition-71-2048.jpg)
![Km
Enzyme Inhibition (Plots)
I II Competitive Non-competitive Uncompetitive
DirectPlotsDoubleReciprocal
Vmax Vmax
Km Km’ [S], mM
vo
[S], mM
vo
I I
Km [S], mM
Vmax
I
Km’
Vmax’Vmax’
Vmax unchanged
Km increased
Vmax decreased
Km unchanged Both Vmax & Km decreased
I
1/[S]1/Km
1/vo
1/Vmax
I
Two parallel
lines
I
Intersect
at X axis
1/vo
1/Vmax
1/[S]1/Km 1/[S]1/Km
1/Vmax
1/vo
Intersect
at Y axis
= Km’](https://image.slidesharecdn.com/enzymesandenzymeinhibition-190509102317/75/Enzymes-and-enzyme-inhibition-72-2048.jpg)





