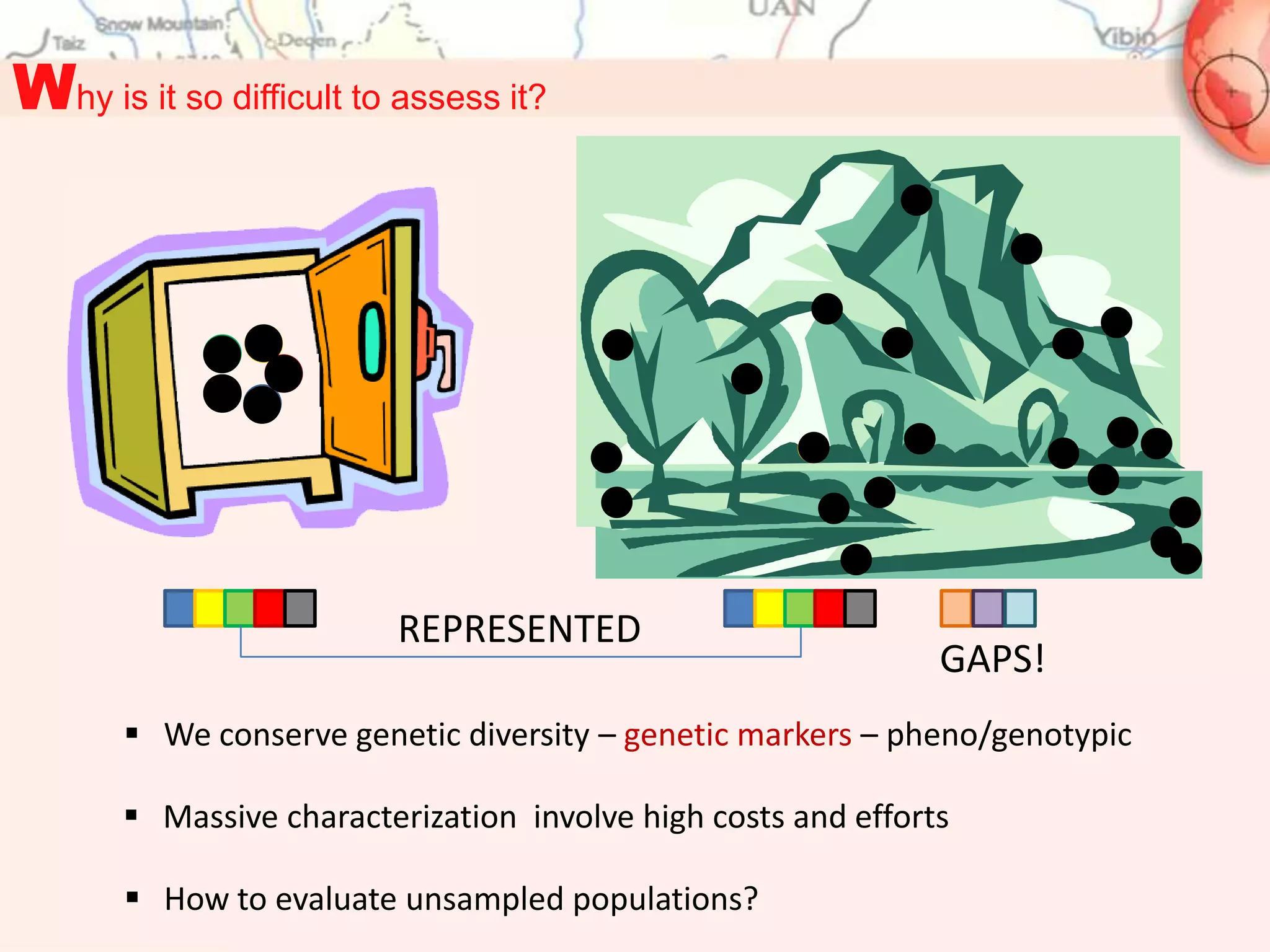
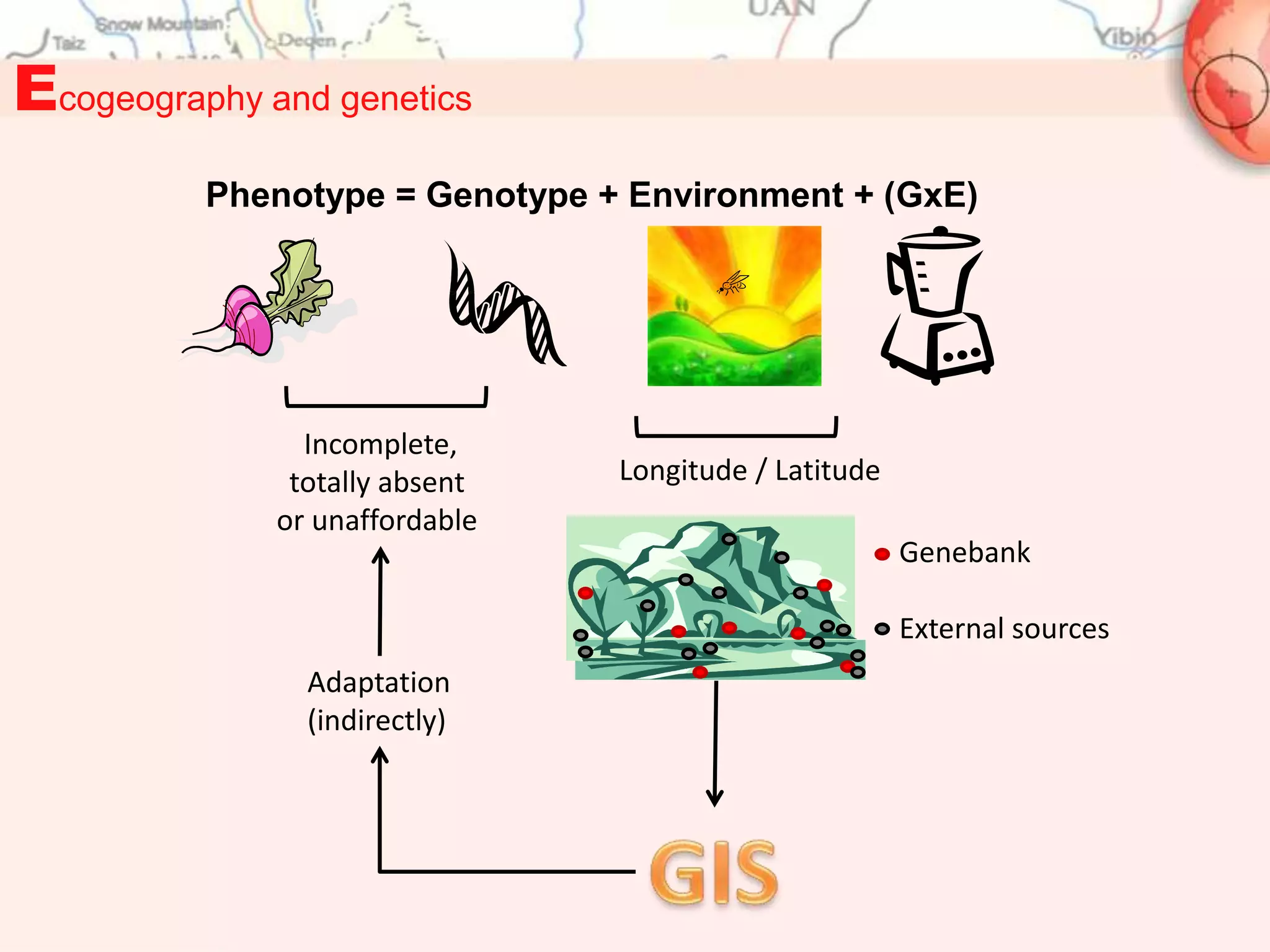
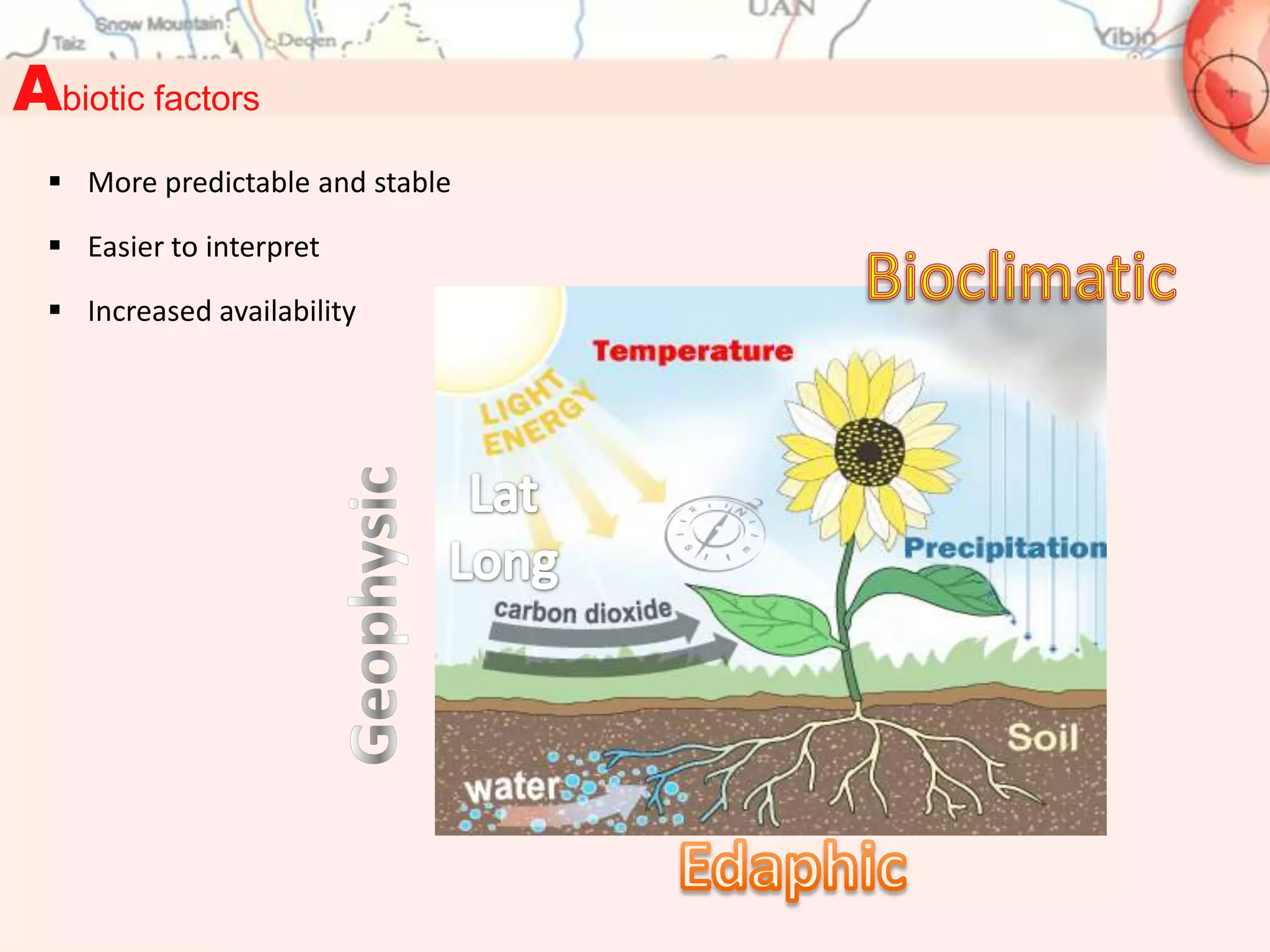
This document discusses representativeness in genebanks and methods to assess it. It is difficult to fully evaluate representativeness due to the large number of unsampled populations and costs of extensive characterization. Ecogeography, which studies adaptive scenarios based on biotic and abiotic factors, can help map diversity and identify core collections. The Focused Identification of Germplasm Strategy uses ecogeographical data to predict traits and select a reduced number of accessions for breeders to evaluate, helping address utilization challenges with large genebank collections. Ecogeographical analyses and FIGS can boost utilization of plant genetic resources.








![More information
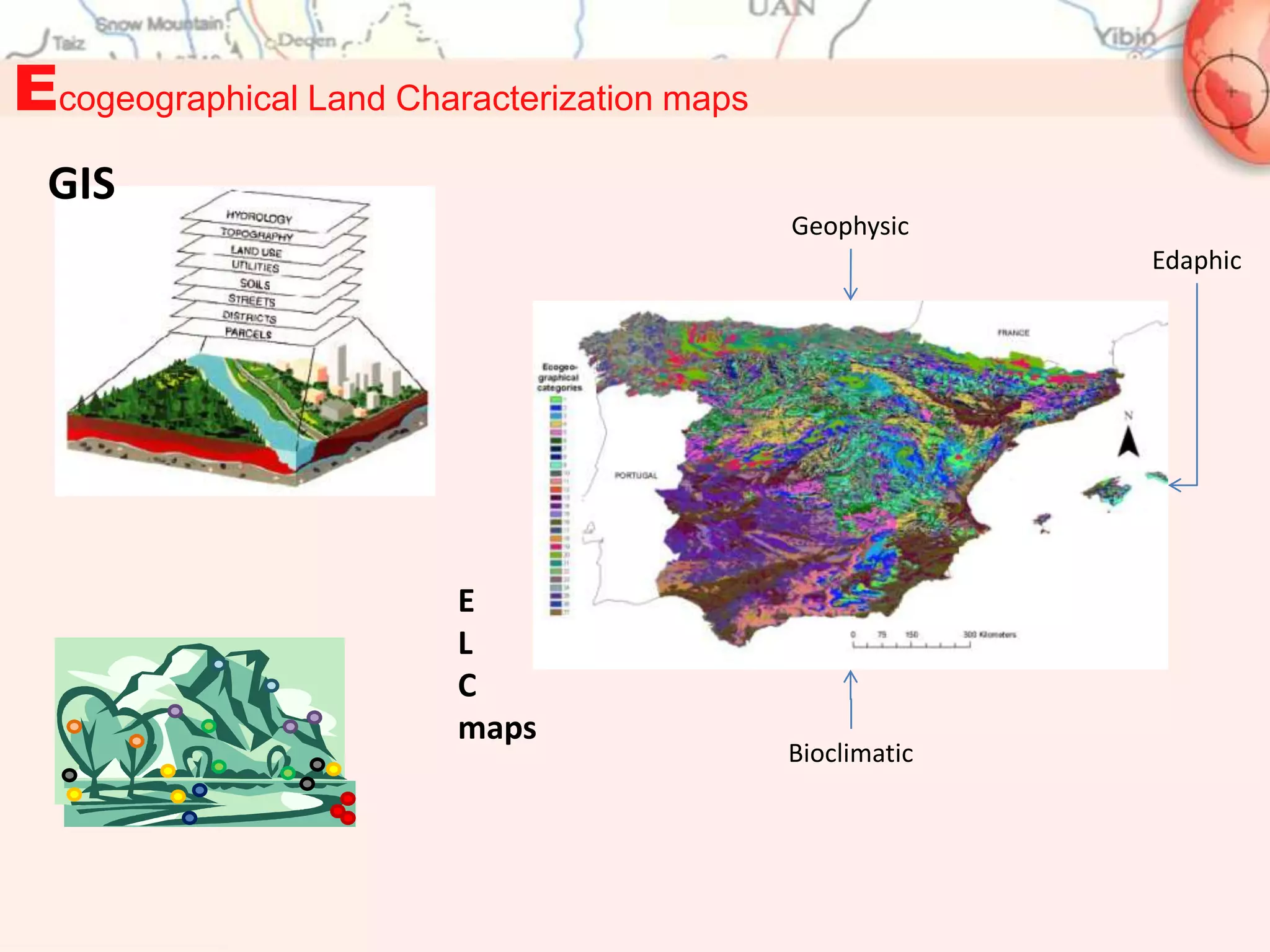
ELC maps:
1. M. Parra-Quijano, J.M. Iriondo , M.E. Torres. 2012. Ecogeographical land characterization maps as a tool for
assessing plant adaptation and their implications in agrobiodiversity studies. Genetic Resources and Crop
Evolution 59(2):205-217
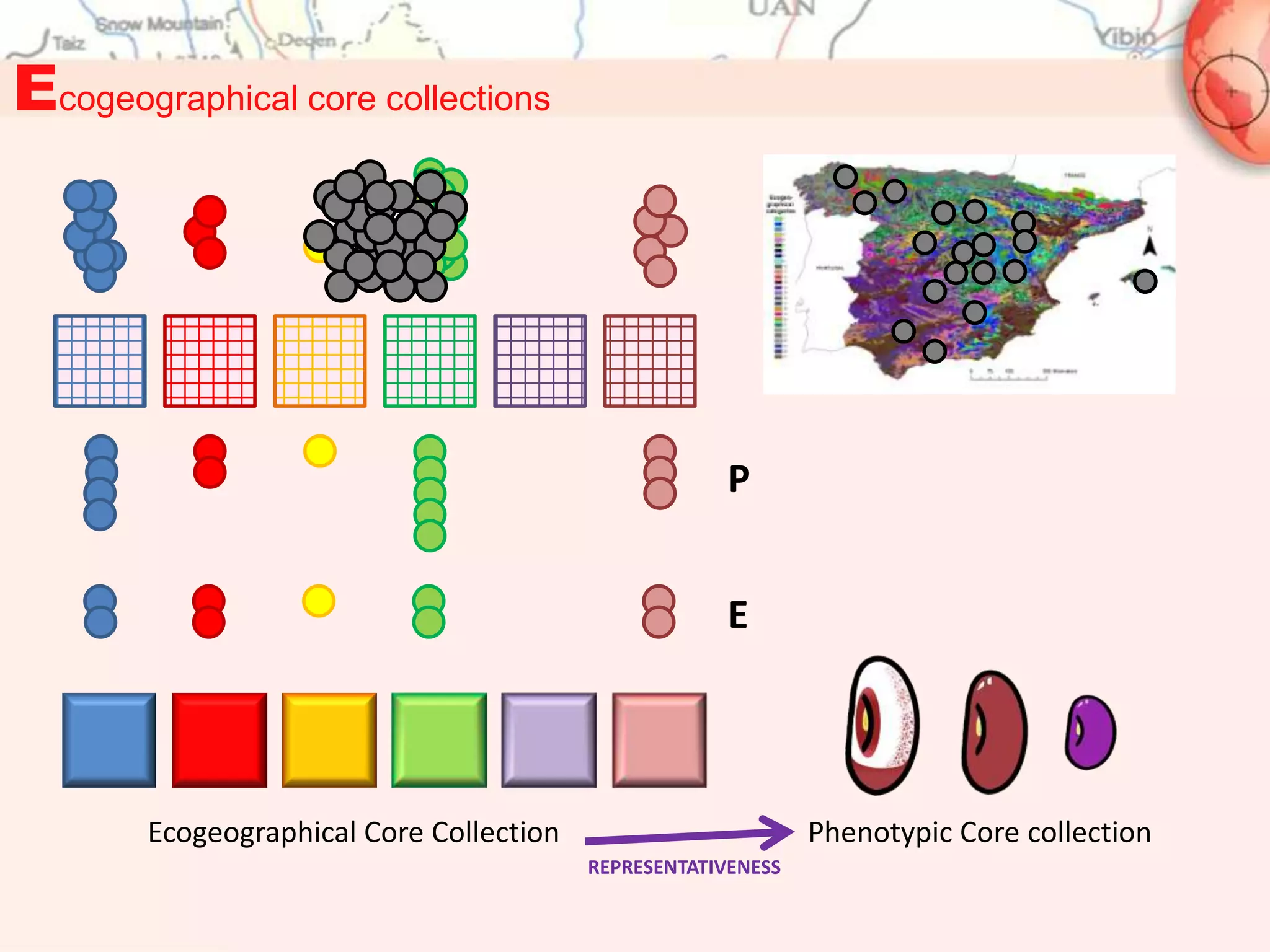
Ecogeographical Core Collections:
2. M. Parra-Quijano, J.M. Iriondo , M.E. Torres, L. De la Rosa. 2011. Evaluation and validation of ecogeographical core
collections using phenotypic data. Crop
Science 51:694-703
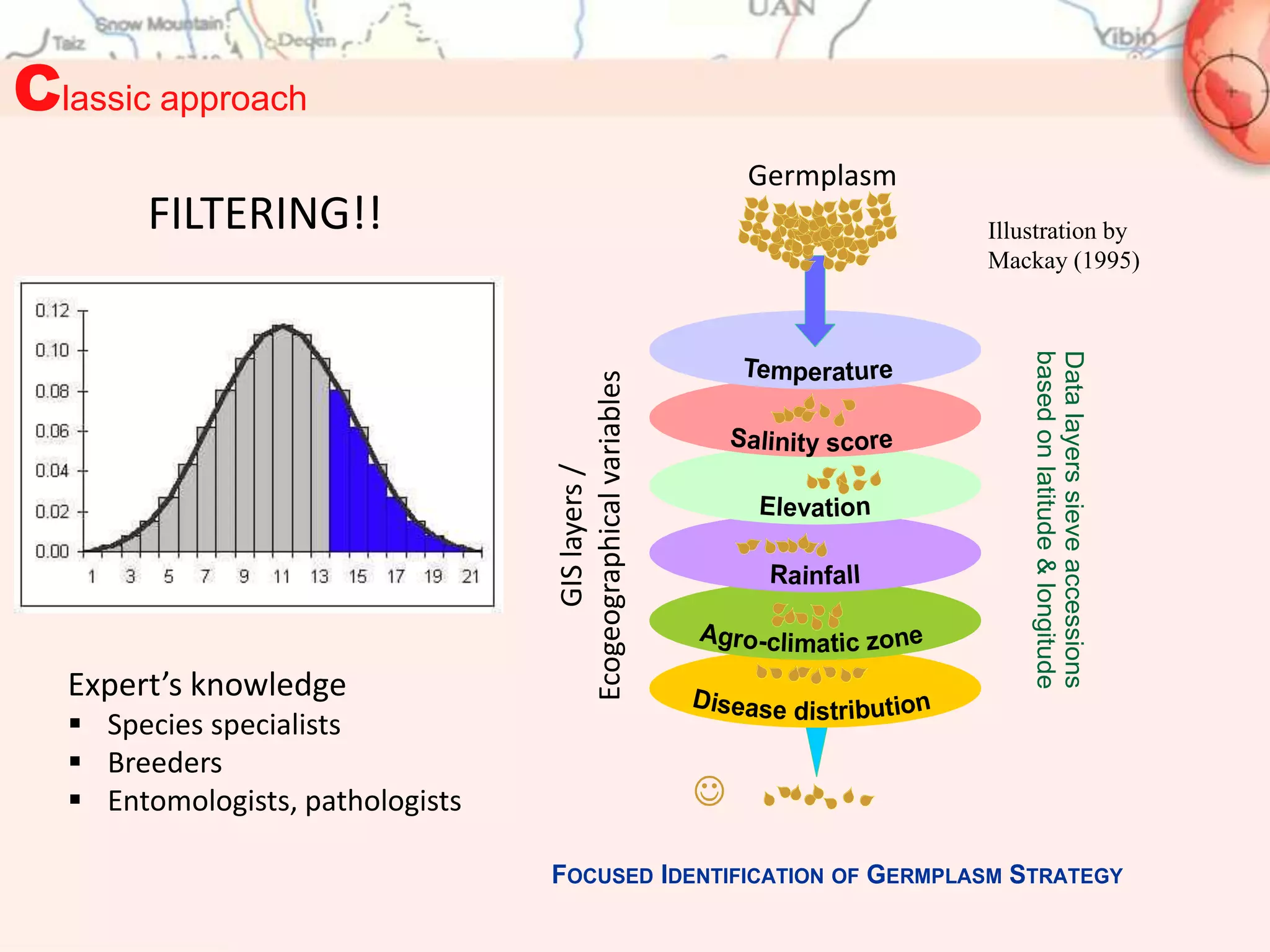
FIGS “classic” approach (study case):
3. El Bouhssini, M., et al.(2011). Sources of resistance in bread wheat to Russian wheat aphid (Diuvaphis noxia) in
Syria identified using the Focused Identification of Germplasm Strategy (FIGS). Plant Breeding 130: 97-97
FIGS modelling approach:
4. Bari, A., K. Street, , M. Mackay, D.T.F. Endresen, E. De Pauw, and A. Amri (2011). Focused Identification of
Germplasm Strategy (FIGS) detects wheat stem rust resistance linked to environment variables. Genetic
Resources and Crop Evolution [online first]. doi:10.1007/s10722-011-9775-5.
Websites
http://www.figstraitmine.org/ (FIGS subsets on wheat)
Review about ecogeographical and GIS tools
http://revistas.inia.es/index.php/sjar/article/view/1859/1673](https://image.slidesharecdn.com/ecogeographiccorecollectionsfigs-120711102144-phpapp01/75/Ecogeographic-core-collections-and-FIGS-17-2048.jpg)