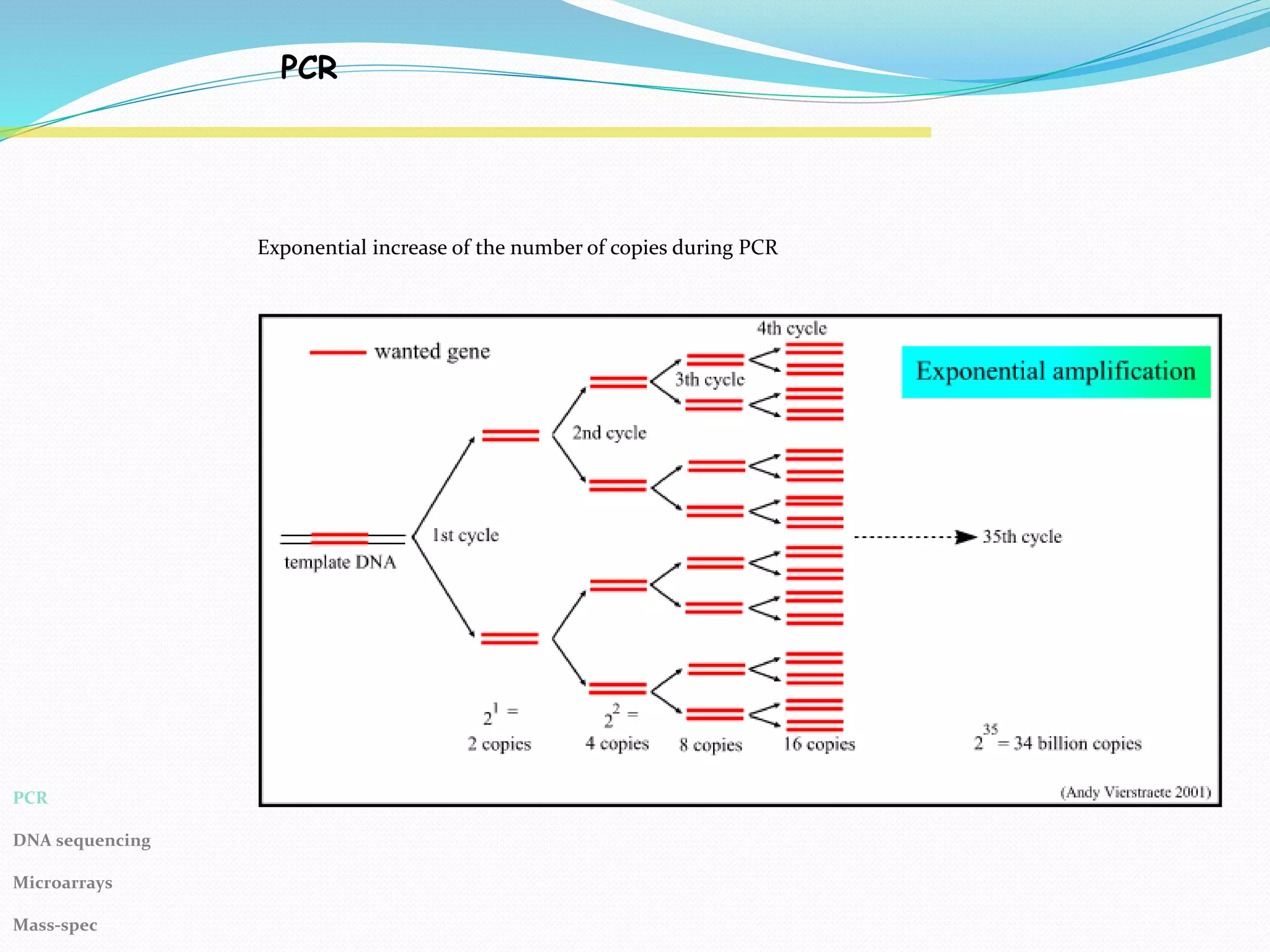
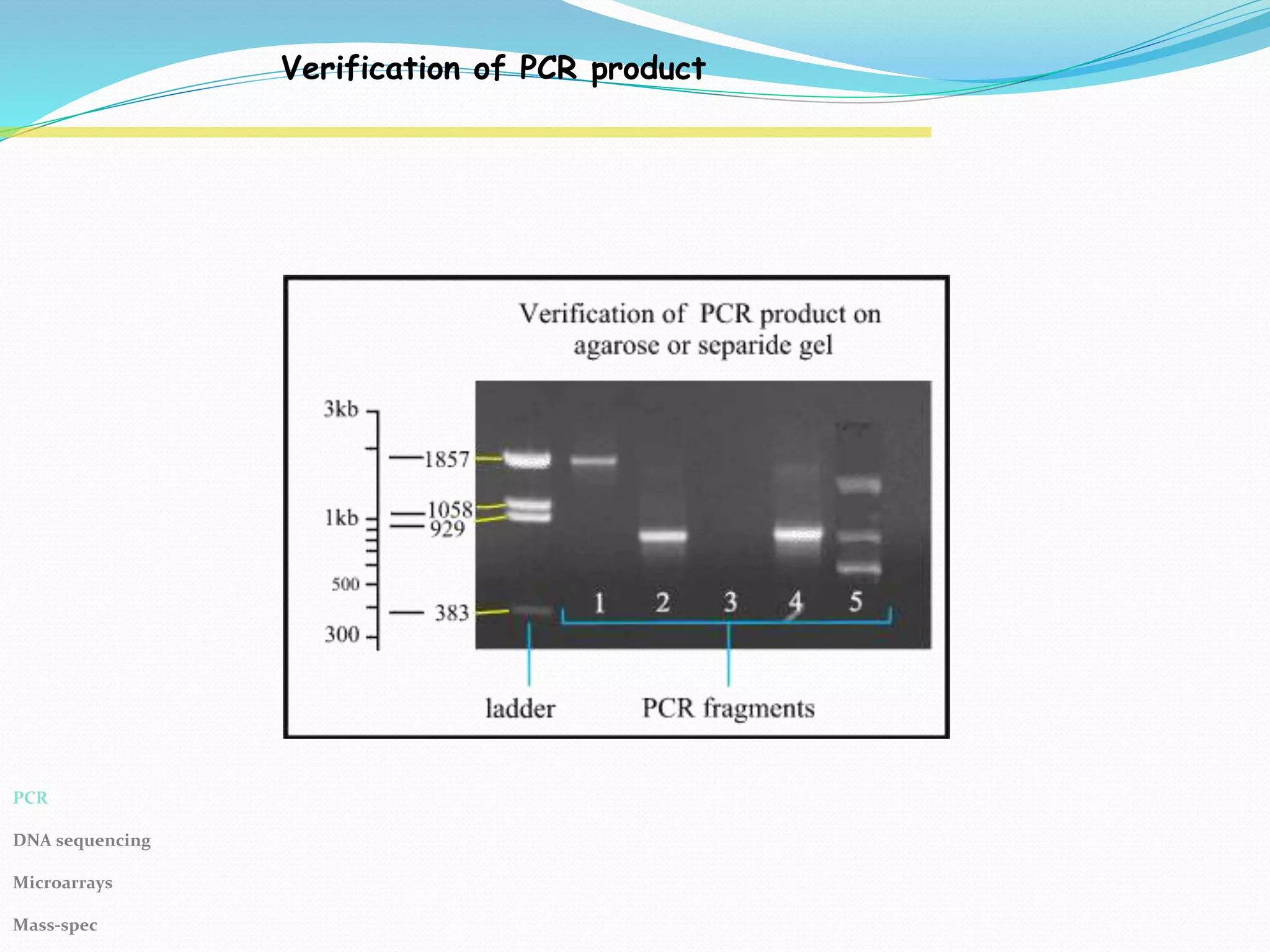
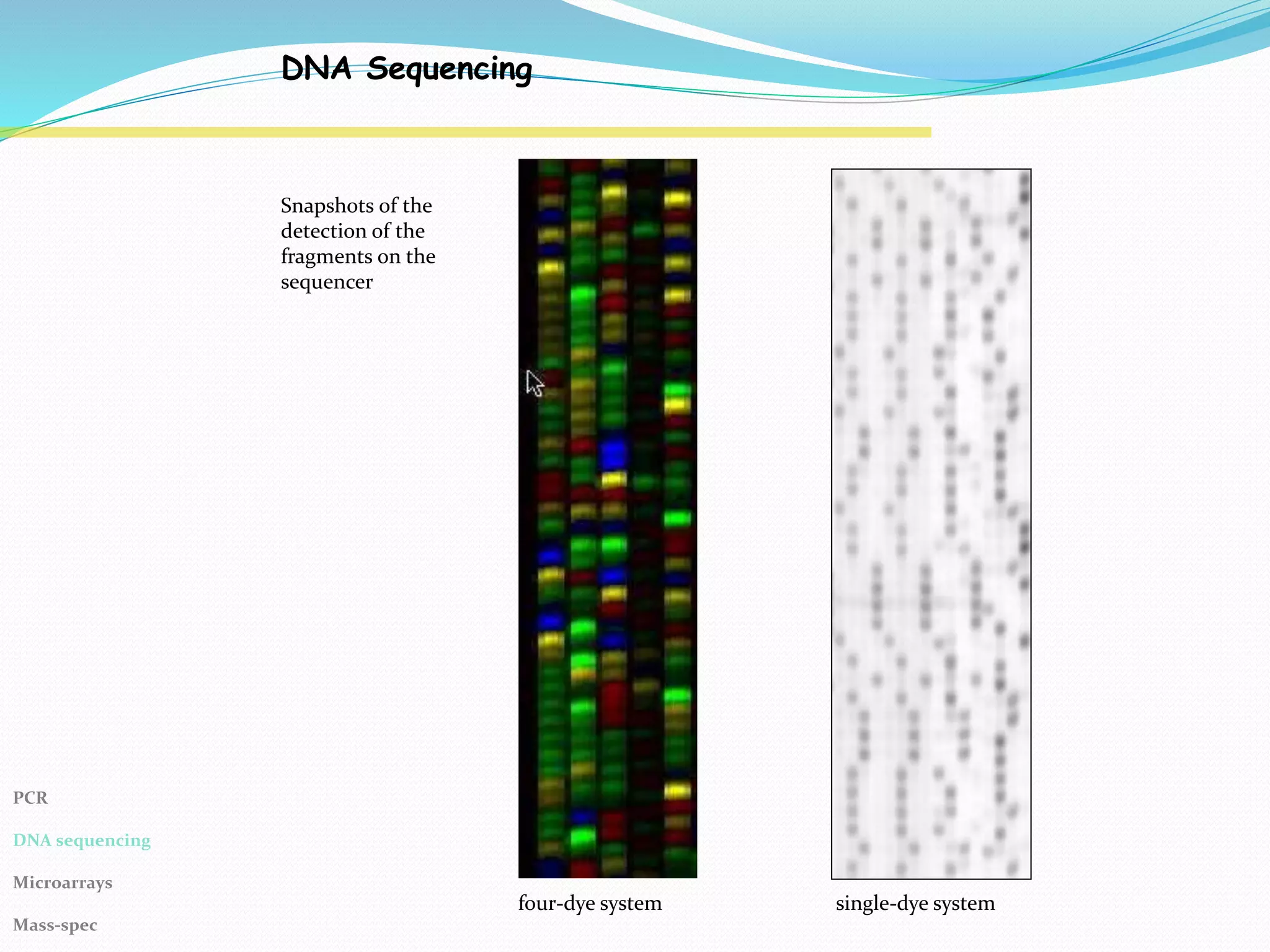
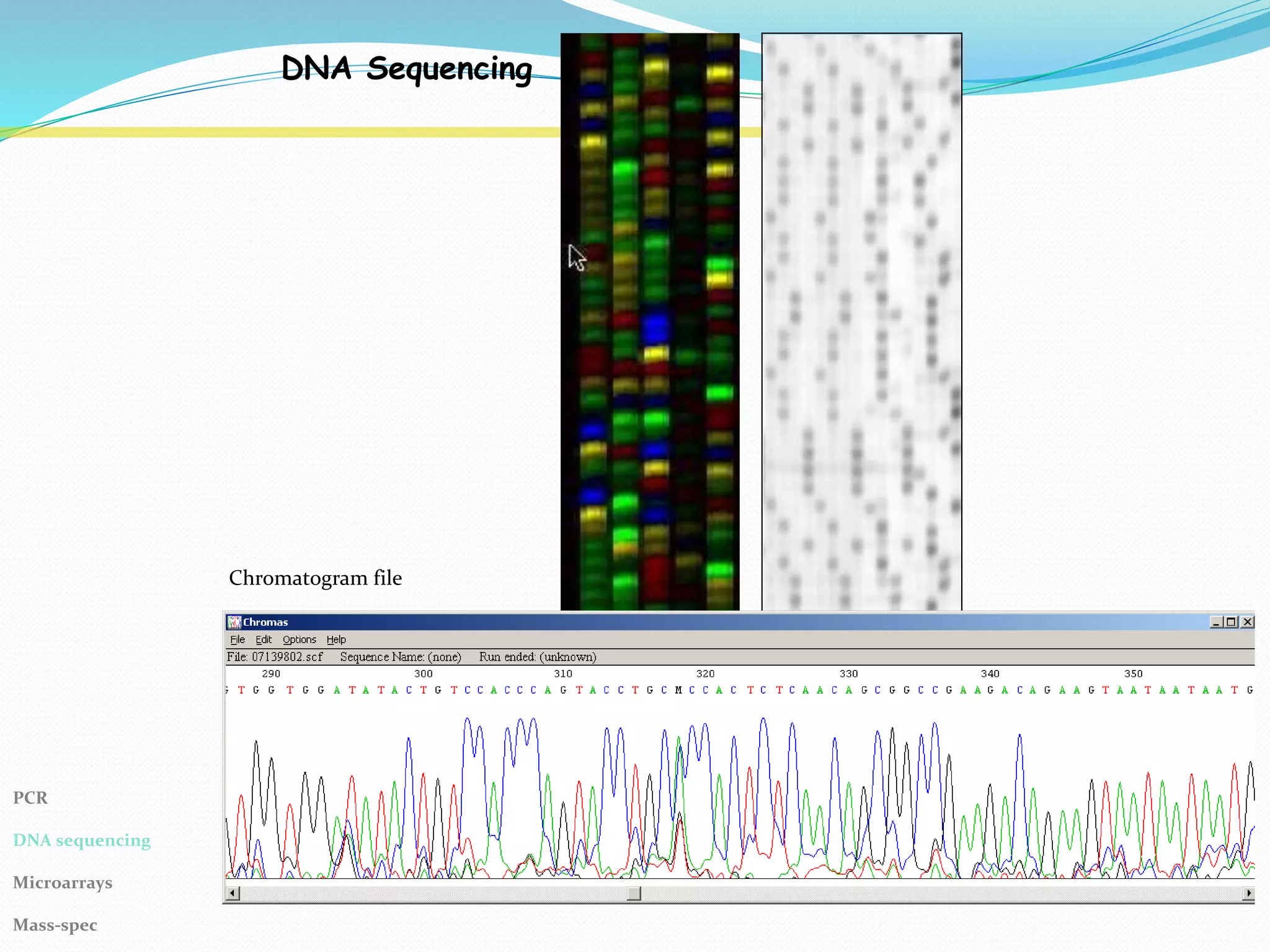
The Polymerase Chain Reaction (PCR) provides an extremely sensitive means of amplifying relatively large quantities of DNA. First described in 1985, it was made possible by the discovery of Taq polymerase. The primary reagents used in PCR are DNA nucleotides, template DNA, primers, and DNA polymerase.