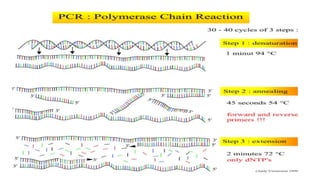
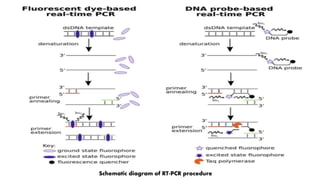
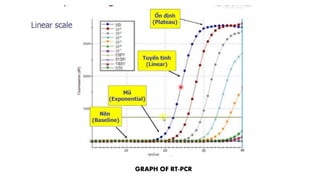
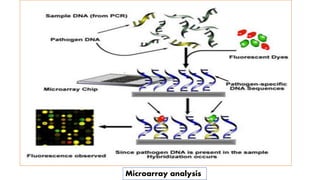
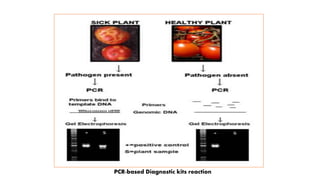
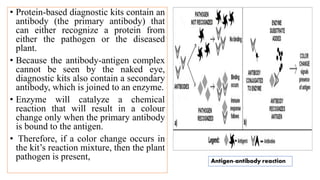
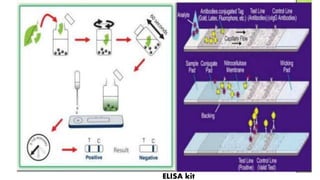
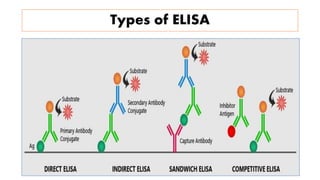
The document discusses the development of diagnostic kits for early detection of plant diseases, highlighting the significant annual losses caused by these diseases and the limitations of traditional diagnosis methods. It details various types of diagnostic kits, including nucleic acid-based and protein-based kits, their operational mechanisms, advantages, and specific pathogens they detect. The future of these diagnostic kits is emphasized as essential for reducing crop losses and improving agricultural practices, particularly in developing countries.