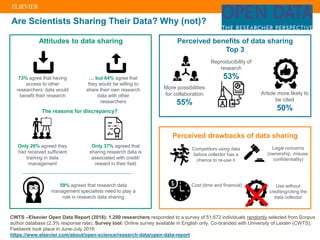
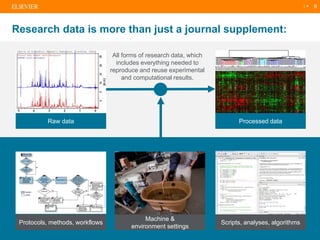
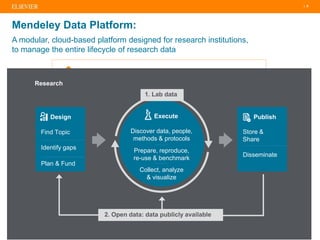
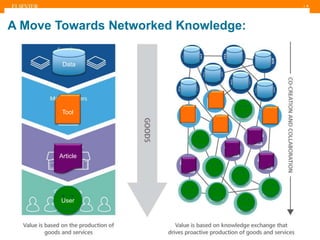
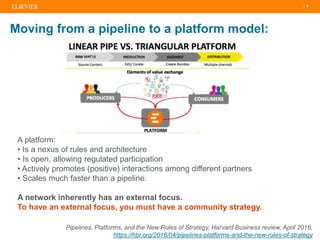
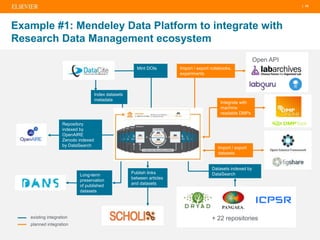
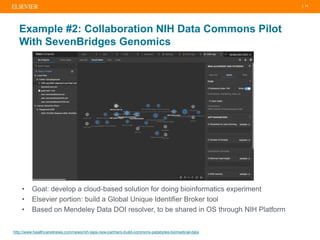
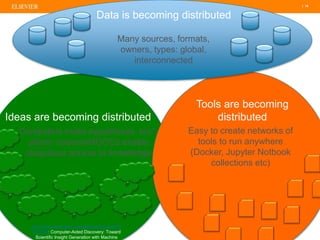
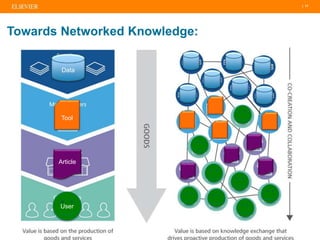
The document discusses publishers' roles in data sharing and challenges in open science. It notes that while most scientists agree access to others' data would benefit research, fewer are willing to share their own data due to lack of training and incentives. Publishers are working to establish data sharing guidelines and integrate platforms to store, share, and analyze research data and tools. However, many questions remain around publishing data science given distributed and interconnected data, tools, and knowledge networks. Publishers will need to transition from pipelines to platforms and enable these new network effects.