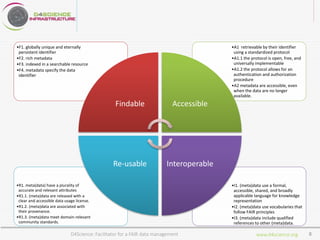
D4Science is a hybrid data infrastructure designed to facilitate fair data management, supporting over 2,500 scientists across 44 countries through virtual research environments (VREs). It emphasizes accessibility, interoperability, and reusability of data while ensuring low operational costs and compliance with FAIR principles. The platform integrates diverse data and services, enabling collaborative research and addressing complex research questions efficiently.