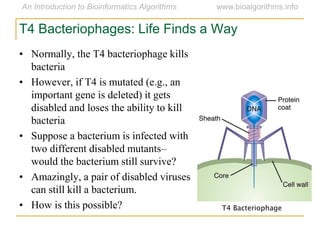
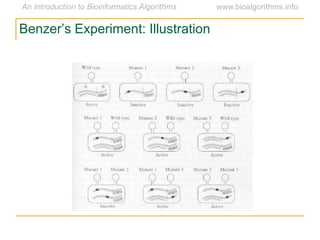
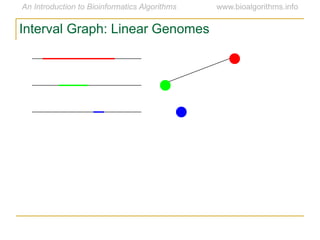
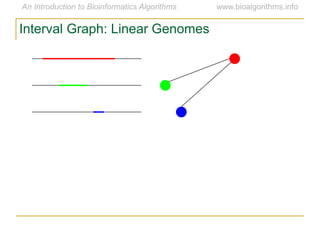
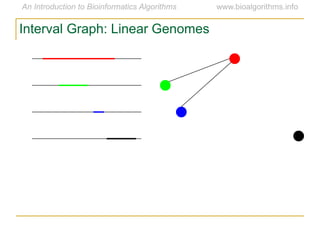
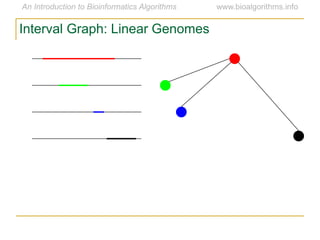
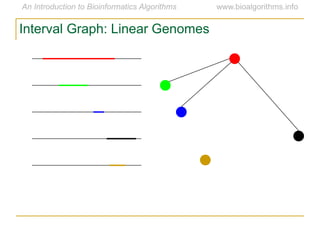
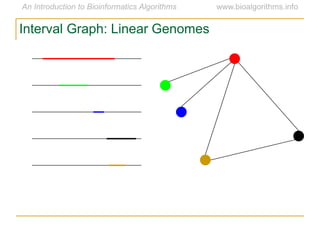
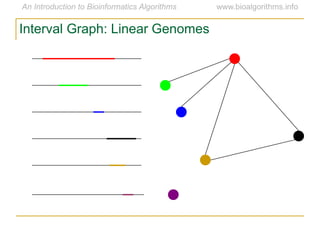
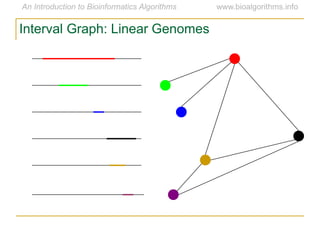
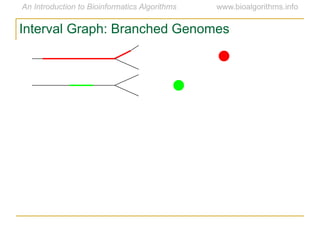
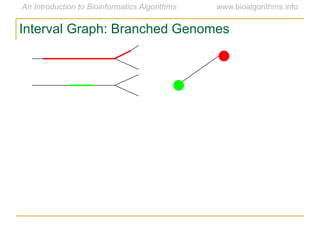
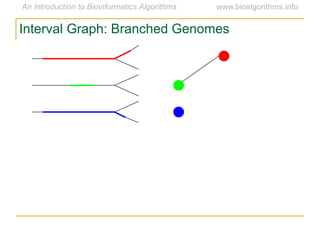
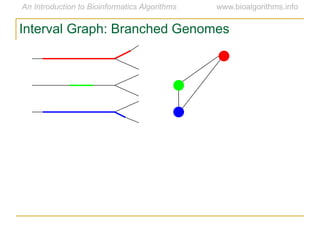
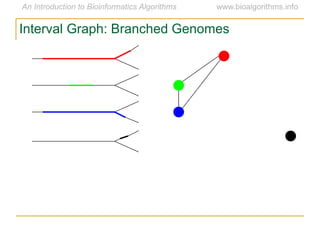
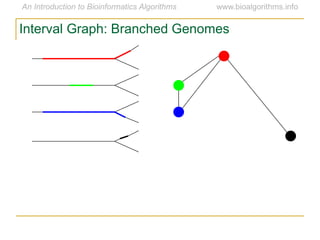
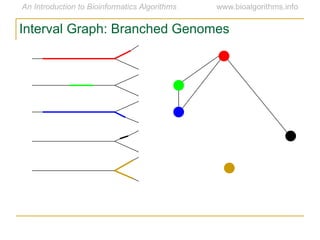
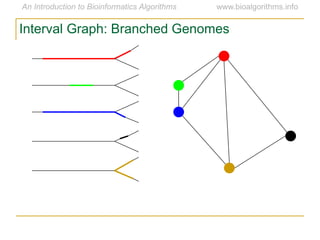
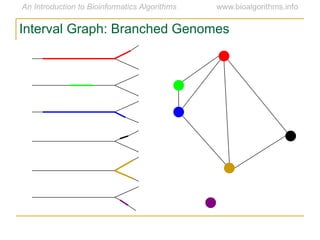
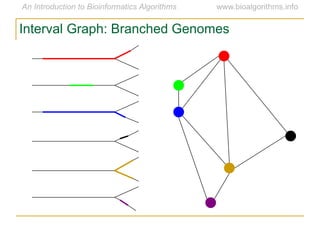
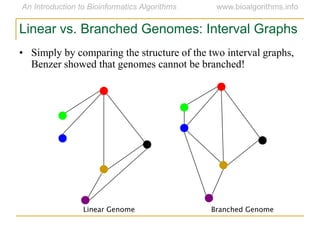
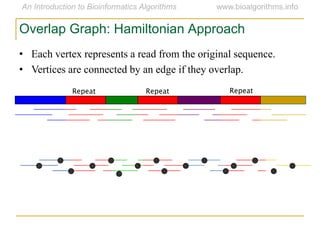
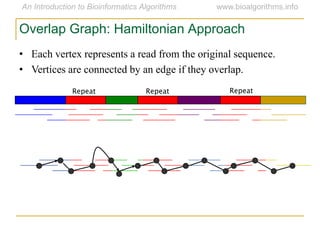
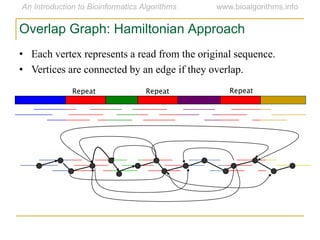
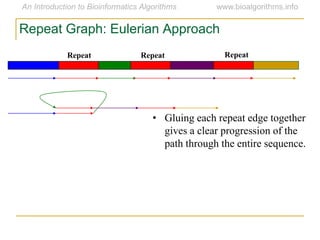
Graph algorithms have important applications in bioinformatics. Seymour Benzer used graph theory and interval graphs to study the structure of viral genomes. Through his experiment infecting bacteria with pairs of disabled bacteriophages, Benzer was able to show that the genome of these viruses was linear rather than branched based on whether the interval graph formed was interval or not. This provided insight into viral genome structure using mathematical concepts like graphs and intervals.