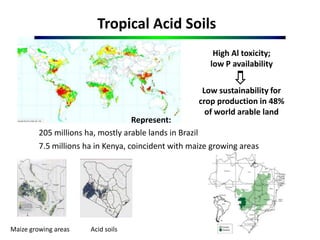
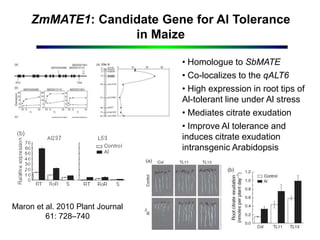
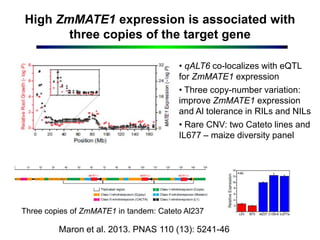
This document outlines objectives to validate genes associated with aluminum tolerance quantitative trait loci (QTLs), specifically ZmMATE genes, in maize crosses from Brazil and Kenya. It identifies ZmMATE1 as a candidate gene for aluminum tolerance in maize that encodes a transporter for citrate exudation from roots. Studies found higher ZmMATE1 expression and aluminum tolerance in lines with three copies of the gene. The objectives are to develop molecular tools like flanking markers for marker-assisted selection of the QTL, and to characterize and distribute genetic materials with improved aluminum tolerance. Key products include characterized germplasm, flanking markers, and capacity building activities.