IB Biology Topic 1.3 - Cell Membrane Structure
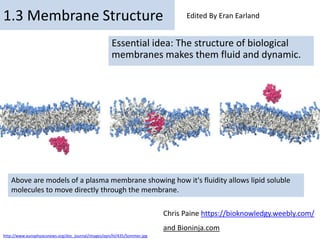
- 1. By Chris Paine https://bioknowledgy.weebly.com/ 1.3 Membrane Structure Essential idea: The structure of biological membranes makes them fluid and dynamic. Above are models of a plasma membrane showing how it's fluidity allows lipid soluble molecules to move directly through the membrane. Chris Paine https://bioknowledgy.weebly.com/ and Bioninja.com http://www.europhysicsnews.org/doc_journal/images/epn/hl/435/Sommer.jpg Edited By Eran Earland
- 2. Understandings, Applications and Skills Statement Guidance 1.3.U1 Phospholipids form bilayers in water due to the amphipathic properties of phospholipid molecules. Amphipathic phospholipids have hydrophilic and hydrophobic properties. 1.3.U2 Membrane proteins are diverse in terms of structure, position in the membrane and function. 1.3.U3 Cholesterol is a component of animal cell membranes. 1.3.A1 Cholesterol in mammalian membranes reduces membrane fluidity and permeability to some solutes. 1.3.S1 Drawing of the fluid mosaic model. Drawings of the fluid mosaic model of membrane structure can be two dimensional rather than three dimensional. Individual phospholipid molecules should be shown using the symbol of a circle with two parallel lines attached. A range of membrane proteins should be shown including glycoproteins. 1.3.S2 Analysis of evidence from electron microscopy that led to the proposal of the Davson-Danielli model. 1.3.S3 Analysis of the falsification of the Davson-Danielli model that led to the Singer-Nicolson model.
- 3. http://www.flickr.com/photos/zorin-denu/5385963280/ What happens when you put a drop of oil in water? 1.3.U1 Phospholipids form bilayers in water due to the amphipathic properties of phospholipid molecules.
- 4. http://www.flickr.com/photos/zorin-denu/5385963280/ The Oil droplet stays together and makes a perfect circular shape. The oil molecules are Hydrophobic Oil Molecules are non- polar and water molecules are polar. See 3.1.5 1.3.U1 Phospholipids form bilayers in water due to the amphipathic properties of phospholipid molecules.
- 5. Link to Topic 2.2 Water Understanding: 2.2 U1 - Water molecules are polar and hydrogen bonds form between them • Water is made up of two hydrogen atoms covalently bonded to an oxygen atom (molecular formula = H2O) • While this covalent bonding involves the sharing of electrons, they are not shared equally between the atoms • Oxygen (due to having a higher electronegativity) attracts the electrons more strongly • The shared electrons orbit closer to the oxygen atom than the hydrogen atoms resulting in polarity
- 6. Phospholipid molecules have a polar (charged) phosphate head and long non-polar lipid tails • The head is hydrophillic (attracted to water) • The tails are hydrophobic (repelled by water) 1.3.U1 Phospholipids form bilayers in water due to the amphipathic properties of phospholipid molecules. http://upload.wikimedia.org/wikipedia/commons/3/39/Phospholipid_TvanBrussel.jpg http://www.ib.bioninja.com.au/_Media/phospholipid_bilayer_med.jpeg When drawing a diagram of a phospholipid this is a good example which shows all the key features
- 7. When put into water, an emergent property is that phospholipids will self-organise to keep their heads ‘wet’ and their tails ‘dry’ micelle liposome 1.3.U1 Phospholipids form bilayers in water due to the amphipathic properties of phospholipid molecules. Arrangement in Membranes: Phospholipids spontaneously arrange into a bilayer The hydrophobic tail regions face inwards and are shielded from the surrounding polar fluids, while the two hydrophilic head regions associate with the cytosolic and extracellular fluids respectively
- 8. http://commons.wikimedia.org/wiki/File:Phospholipids_aqueous_solution_structures.svg In this 3D representation you can see that a phospholipid bilayer is one way that the tails can be removed from the water. Phospholipid molecules can flow past each other laterally but can’t move vertically 1.3.U1 Phospholipids form bilayers in water due to the amphipathic properties of phospholipid molecules.
- 9. Properties of the Phospholipid Bilayer: • The bilayer is held together by weak hydrophobic interactions between the tails • Hydrophilic / hydrophobic layers restrict the passage of many substances • Individual phospholipids can move within the bilayer, allowing for membrane fluidity and flexibility • This fluidity allows for the spontaneous breaking and reforming of membranes (endocytosis / exocytosis) 1.3.U1 Phospholipids form bilayers in water due to the amphipathic properties of phospholipid molecules.
- 10. But wait! there’s more! The plasma membrane is not just made of phospholipids http://commons.wikimedia.org/wiki/File:Cell_membrane_detailed_diagram_en.svg?uselang=en-gb 1.3.U1 Phospholipids form bilayers in water due to the amphipathic properties of phospholipid molecules.
- 11. Proteins: Phospholipid bilayers are embedded with proteins, which may be either permanently or temporarily attached to the membrane • Integral proteins are permanently attached to the membrane and are typically transmembrane (they span across the bilayer) • Peripheral proteins are temporarily attached by non-covalent interactions and associate with one surface of the membrane 1.3.U2 Membrane proteins are diverse in terms of structure, position in the membrane and function. http://commons.wikimedia.org/wiki/File:Cell_membrane_detailed_diagram_en.svg?uselang=en-gb
- 12. Connections to Topic 2.4 – Proteins http://ib.bioninja.com.au/standard-level/topic-2-molecular-biology/24-proteins/ Structure of Membrane Proteins The amino acids of a membrane protein are localized according to polarity: • Non-polar (hydrophobic) amino acids associate directly with the lipid bilayer • Polar (hydrophilic) amino acids are located internally and face aqueous solutions
- 13. Connections to Topic 2.4 – Proteins http://ib.bioninja.com.au/standard-level/topic-2-molecular-biology/24-proteins/ Structure of Membrane Proteins Transmembrane proteins typically adopt one of two tertiary structures: • Single helices / helical bundles • Beta barrels (common in channel proteins)
- 14. Functions of Membrane Proteins Membrane proteins can serve a variety of key functions: • Junctions – Serve to connect and join two cells together • Enzymes – Fixing to membranes localizes metabolic pathways • Transport – Responsible for facilitated diffusion and active transport • Recognition – May function as markers for cellular identification • Anchorage – Attachment points for cytoskeleton and extracellular matrix • Transduction – Function as receptors for peptide hormones Mnemonic: Jet Rat 1.3.U2 Membrane proteins are diverse in terms of structure, position in the membrane and function.
- 15. Cell Membrane Protein Functions
- 16. Glycoproteins: Are proteins with an oligosaccaride (oligo = few, saccharide = sugar) chain attached. They are important for cell recognition by the immune system and as hormone receptors http://commons.wikimedia.org/wiki/File:Cell_membrane_detailed_diagram_en.svg?uselang=en-gb 1.3.U2 Membrane proteins are diverse in terms of structure, position in the membrane and function.
- 17. 1.3.U2 Membrane proteins are diverse in terms of structure, position in the membrane and function. Transport: Protein channels (facilitated) and protein pumps (active) Receptors: Peptide-based hormones (insulin, glucagon, etc.) Anchorage: Cytoskeleton attachments and extracellular matrix Cell recognition: MHC proteins and antigens Intercellular joinings: Tight junctions and plasmodesmata Enzymatic activity: Metabolic pathways (e.g. electron transport chain) http://www.ib.bioninja.com.au/standard-level/topic-2-cells/24-membranes.html
- 18. Cholesterol: (It’s not all bad!) It makes the phospholipids pack more tightly and regulates the fluidity and flexibility of the membrane. Bad analogy: imagine a room full of people wearing fluffy jumpers (sweaters). It is crowded but they can slip past each other easily enough. Now sprinkle the crowd with people wearing Velcro™ suits… 1.3.U3 Cholesterol is a component of animal cell membranes.
- 19. Cholesterol is a component of animal cell membranes, where it functions to maintain integrity and mechanical stability It is absent in plant cells, as these plasma membranes are surrounded and supported by a rigid cell wall made of cellulose cholesterol molecule Cholesterol is an amphipathic molecule (like phospholipids), meaning it has both hydrophilic and hydrophobic regions • Cholesterol’s hydroxyl (-OH) group is hydrophilic and aligns towards the phosphate heads of phospholipids • The remainder of the molecule (steroid ring and hydrocarbon tail) is hydrophobic and associates with the phospholipid tails
- 20. Cholesterol 1.3.U3 Cholesterol is a component of animal cell membranes. http://www.cholesterol-and-health.com/images/Cholesterol_Structure.jpg http://www.uic.edu/classes/bios/bios100/lectf03am/cholesterol.jpg Hydroxyl group makes the head polar and hydrophilic - attracted to the phosphate heads on the periphery of the membrane. Carbon rings – it’s not classed as a fat or an oil, cholesterol is a steroid Non-polar (hydrophobic) tail –attracted to the hydrophobic tails of phospholipids in the centre of the membrane
- 21. Membrane fluidity 1.3.A1 Cholesterol in mammalian membranes reduces membrane fluidity and permeability to some solutes. It is important to regulate the degree of fluidity: • Membranes need to be fluid enough that the cell can move • Membranes need to be fluid enough that the required substances can move across the membrane • If too fluid however the membrane could not effectively restrict the movement of substances across itself http://www.nature.com/scitable/content/ne0000/ne0000/ne0000/ne0000/14668965/U2.cp5.3_membrane_f2.jpg The hydrophobic hydrocarbon tails usually behave as a liquid. Hydrophilic phosphate heads act more like a solid. Though it is difficult to determine whether the membrane is truly either a solid or liquid it can definitely be said to be fluid.
- 22. Cholesterol’s role in membrane fluidity 1.3.A1 Cholesterol in mammalian membranes reduces membrane fluidity and permeability to some solutes. The presence of cholesterol disrupts the regular packing of the of the hydrocarbon tails of phospholipid molecules - this is increases the flexibility as it prevents the tails from crystallising and hence behaving like a solid. Cholesterol also reduces the permeability to hydrophilic/water soluble molecules and ions such as sodium and hydrogen. 2. 3. http://www.nature.com/scitable/content/ne0000/ne0000/ne0000/ne0000/14668965/U2.cp5.3_membrane_f2.jpg The presence of cholesterol in the membrane restricts the movement of phospholipids and other molecules – this reduces membrane fluidity. 1.
- 23. 1.3.S1 Drawing of the fluid mosaic model. https://www.wisc-online.com//LearningContent/ap1101/index.html http://www.phschool.com/science/biology_place/biocoach/biomembrane1/regi ons.html http://www.bio.davidson.edu/people/macampbell/111/memb- swf/membranes.swf Use the tutorials to learn and review membrane structure
- 24. 1.3.S1 Drawing of the fluid mosaic model. http://www.youtube.com/watch?v=w9VBHGNoFrY
- 25. 1.3.S1 Drawing of the fluid mosaic model. http://www.ib.bioninja.com.au/_Media/phospholipid_bilayer_med.jpeg • Good use of space • Clear strong lines • Label lines are straight • Labels clearly written • (Scale bar if appropriate) • Lines touch the labeled structure • No unnecessary shading or colouring Reminder of features that make good diagrams:
- 26. • The fluid-mosaic model was not the first scientifically accepted paradigm to describe membrane structure • The first model that attempted to describe the position of proteins within the bilayer was proposed by Hugh Davson and James Danielli in 1935 • When viewed under a transmission electron microscope, membranes exhibit a characteristic 'trilaminar’ appearance • Trilaminar = 3 layers (two dark outer layers and a lighter inner region) 1.3.S2 Analysis of evidence from electron microscopy that led to the proposal of the Davson-Danielli model.
- 27. • Danielli and Davson proposed a model whereby two layers of protein flanked a central phospholipid bilayer • The model was described as a 'lipo-protein sandwich’, as the lipid layer was sandwiched between two protein layers • The dark segments seen under electron microscope were identified (wrongly) as representing the two protein layers 1.3.S2 Analysis of evidence from electron microscopy that led to the proposal of the Davson-Danielli model.
- 28. There were a number of problems with the lipo-protein sandwich model proposed by Davson and Danielli: • It assumed all membranes were of a uniform thickness and would have a constant lipid-protein ratio • It assumed all membranes would have symmetrical internal and external surfaces (i.e. not bifacial) • It did not account for the permeability of certain substances (did not recognise the need for hydrophilic pores) • The temperatures at which membranes solidified did not correlate with those expected under the proposed model 1.3.S3 Analysis of the falsification of the Davson-Danielli model that led to the Singer- Nicolson model.
- 29. Falsification Evidence: Membrane proteins were discovered to be insoluble in water indicating hydrophobic surfaces) and varied in size • Such proteins would not be able to form a uniform and continuous layer around the outer surface of a membrane Fluorescent antibody tagging of membrane proteins showed they were mobile and not fixed in place • Membrane proteins from two different cells were tagged with red and green fluorescent markers respectively • When the two cells were fused, the markers became mixed throughout the membrane of the fused cell • This demonstrated that the membrane proteins could move and did not form a static layer (as per Davson-Danielli) Freeze fracturing was used to split open the membrane and revealed irregular rough surfaces within the membrane • These rough surfaces were interpreted as being transmembrane proteins, demonstrating that proteins were not solely localised to the outside of the membrane structure 1.3.S3 Analysis of the falsification of the Davson-Danielli model that led to the Singer- Nicolson model.
- 30. New Model: • In light of these limitations, a new model was proposed by Seymour Singer and Garth Nicolson in 1972 • According to this model, proteins were embedded within the lipid bilayer rather than existing as separate layers • This model, known as the fluid-mosaic model, remains the model preferred by scientists today (with refinements) 1.3.S3 Analysis of the falsification of the Davson-Danielli model that led to the Singer- Nicolson model.
- 31. Our current model of the cell membrane is called the Singer-Nicholson fluid mosaic model http://commons.wikimedia.org/wiki/File:Cell_membrane_detailed_diagram_en.svg?uselang=en-gb 1.3.S3 Analysis of the falsification of the Davson-Danielli model that led to the Singer- Nicolson model. Key features: • Phospholipid molecules form a bilayer - phospholipids are fluid and move laterally • Peripheral proteins are bound to either the inner or outer surface of the membrane • Integral proteins - permeate the surface of the membrane • The membrane is a fluid mosaic of phospholipids and proteins • Proteins can move laterally along membrane
- 32. Our current model of the cell membrane is called the Singer-Nicholson fluid mosaic model There is strong evidence for this model: http://commons.wikimedia.org/wiki/File:Cell_membrane_detailed_diagram_en.svg?uselang=en-gb 1.3.S3 Analysis of the falsification of the Davson-Danielli model that led to the Singer- Nicolson model. Biochemical techniques • Membrane proteins were found to be very varied in size and globular in shape • Such proteins would be unable to form continuous layers on the periphery of the membrane. • The membrane proteins had hydrophobic regions and therefore would embed in the membrane not layer the outside
- 33. Our current model of the cell membrane is called the Singer-Nicholson fluid mosaic model There is strong evidence for this model: http://commons.wikimedia.org/wiki/File:Cell_membrane_detailed_diagram_en.svg?uselang=en-gb 1.3.S3 Analysis of the falsification of the Davson-Danielli model that led to the Singer- Nicolson model. Fluorescent antibody tagging • Within 40 minutes the red and green markers were mixed throughout the membrane of the fused cell. • This showed that membrane proteins are free to move within the membrane rather than being fixed in a peripheral layer. • red or green fluorescent markers attached to antibodies which would bind to membrane proteins • The membrane proteins of some cells were tagged with red markers and other cells with green markers. • The cells were fused together.
- 34. Our current model of the cell membrane is called the Singer-Nicholson fluid mosaic model http://commons.wikimedia.org/wiki/File:Cell_membrane_detailed_diagram_en.svg?uselang=en-gb 1.3.S3 Analysis of the falsification of the Davson-Danielli model that led to the Singer- Nicolson model. This model was first proposed in by Singer-Nicolson in 1972 Before then Davson-Danielli model was widely accepted …
- 35. 1.3.S2 Analysis of evidence from electron microscopy that led to the proposal of the Davson-Danielli model. The model: • A protein-lipid sandwich • Lipid bilayer composed of phospholipids (hydrophobic tails inside, hydrophilic heads outside) • Proteins coat outer surface • Proteins do not permeate the lipid bilayer http://www.cytochemistry.net/cell-biology/EMview.jpg PoreProteins Phospholipids This explains: Despite being very thin membranes are an effective barrier to the movement of certain substances. http://upload.wikimedia.org/wikipedia/commons/6/69/Davson_danielli_miguelferig.jpg The evidence: In high magnification electron micrographs membranes appeared as two dark parallel lines with a lighter coloured region in between. Proteins appear dark in electron micrographs and phospholipids appear light - possibly indicating proteins layers either side of a phospholipid core. Davson-Danielli Model
- 36. 1.3.S3 Analysis of the falsification of the Davson-Danielli model that led to the Singer- Nicolson model. http://www.cytochemistry.net/cell-biology/ffimage.jpg This technique involves rapid freezing of cells and then fracturing them. Interpreting the image: • The fracture occurs along lines of weakness, including the centre of membranes. • The fracture reveals an irregular rough surface inside the phospholipid bilayer • The globular structures were interpreted as trans- membrane proteins. Falsification of the Davson-Danielli model – freeze fracturing Conclusion: This is contrary to the Davson-Danielli model which only involves proteins coating the surface of the membrane. A new model is needed to explain the presence of as trans-membrane proteins.
- 37. Bibliography / Acknowledgments Jason de Nys