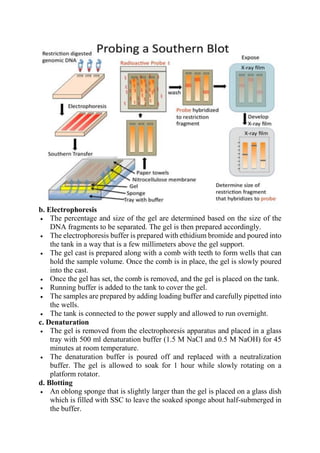
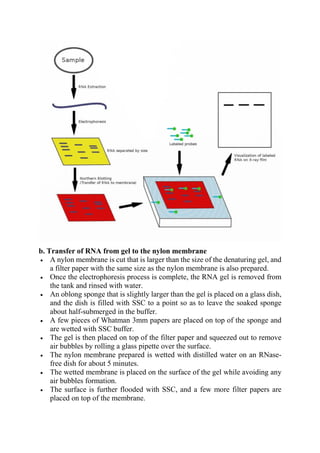
The document provides detailed notes on Southern, Western, and Northern blotting techniques used in biotechnology. Southern blotting involves transferring DNA fragments for analysis, while Western blotting focuses on protein detection using antibodies. Northern blotting is similar to Southern blotting but is used for RNA analysis, highlighting the principles, procedures, and applications of each technique.