Embed presentation
Download to read offline






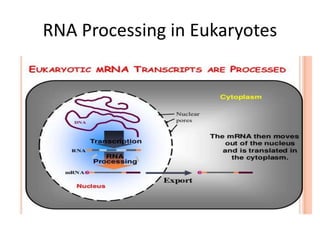





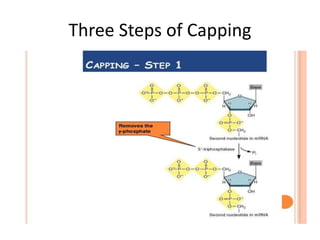
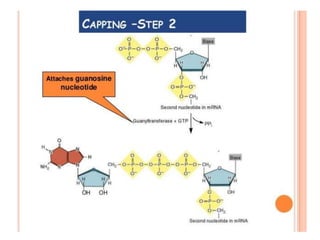
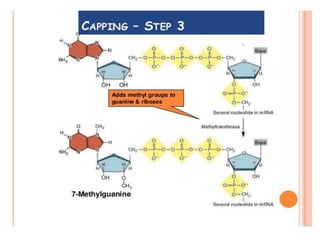


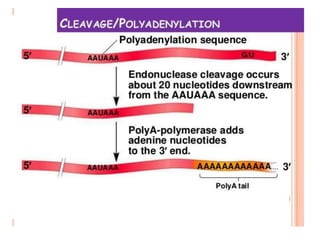


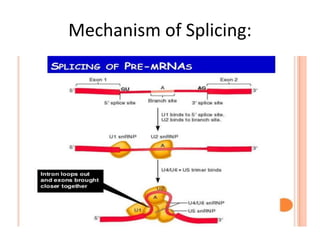
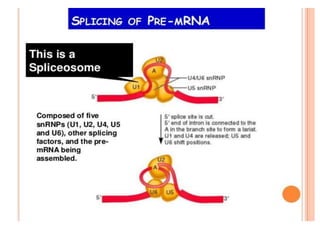
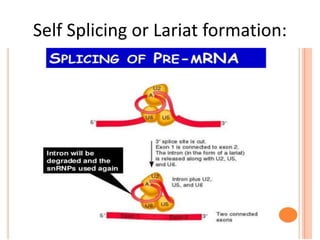







The document presented by Shahzaib Khurshid discusses the necessity of RNA processing in both prokaryotes and eukaryotes. It covers various aspects such as hnrna, snrna, 5' capping, poly A tail addition, and the mechanism of RNA splicing including spliceosome functions. Additionally, it addresses the steps involved in RNA processing and invites questions on the topic.






























