Recommended
The UKMenCar4 study: carriage studies, high-resolution genomics and disease control
http://www.meningitis.org/conference2015Professor Martin Maiden @ MRF's Meningitis & Septicaemia in Children & Adults...

Professor Martin Maiden @ MRF's Meningitis & Septicaemia in Children & Adults...Meningitis Research Foundation
First use of genomics to inform national meningococcal immunisation policy
http://www.meningitis.org/conference2015Dr Jay Lucidarme @ MRF's Meningitis & Septicaemia in Children & Adults 2015 

Dr Jay Lucidarme @ MRF's Meningitis & Septicaemia in Children & Adults 2015 Meningitis Research Foundation
Recommended
The UKMenCar4 study: carriage studies, high-resolution genomics and disease control
http://www.meningitis.org/conference2015Professor Martin Maiden @ MRF's Meningitis & Septicaemia in Children & Adults...

Professor Martin Maiden @ MRF's Meningitis & Septicaemia in Children & Adults...Meningitis Research Foundation
First use of genomics to inform national meningococcal immunisation policy
http://www.meningitis.org/conference2015Dr Jay Lucidarme @ MRF's Meningitis & Septicaemia in Children & Adults 2015 

Dr Jay Lucidarme @ MRF's Meningitis & Septicaemia in Children & Adults 2015 Meningitis Research Foundation
ASHG 2016 PresentationA Retrospective Analysis of Exome Sequencing Cases Using the GenePool™ Genomi...

A Retrospective Analysis of Exome Sequencing Cases Using the GenePool™ Genomi...Antoaneta Vladimirova
Presentation from the ECDC expert consultation on Whole Genome Sequencing organised by the European Centre of Disease Prevention and Control - Stockholm, 19 November 2015Overview of the ECDC whole genome sequencing strategy

Overview of the ECDC whole genome sequencing strategyEuropean Centre for Disease Prevention and Control (ECDC)
Selection of genes to include in genomic studies of disease
remains a difficult task. Current methods rely on expert opinion
or manual search engine use. With these methods, the
process and result are neither repeatable nor scalable. To
remedy this situation, we created the Informative Genetic
Content (IGC) system, which enables the algorithmic selection
of genes for inclusion in such studies, given one or more
diseases to target.
The IGC system stands on three components: a database
associating diseases with genes and other diseases, an
algorithm to rank the genes under consideration for inclusion in
a panel, and a module that clusters genes by families of
diseases. The first component, the database, maps diseases
to associated genes and scores each of these mappings
according to the strength of the relationship. The database also
maps diseases to other diseases, such that groups of diseases
or hierarchical relationships between diseases can be
identified. The second component enables the ranking of
candidate genes when multiple diseases are of interest. The
algorithm accounts for the common situation where two or
more diseases are associated with the same gene with varying
strengths of association, weighting and combining the scores
across the diseases associated with each gene. The final
component, the gene clustering module, groups genes by
pathogenic pathways, should the user want to consider
targeting a broader family of diseases affected by a closely
related set of genes.
We validated the IGC system through comparisons of our
automated gene selections with expertly curated gene panel
designs. We found a high degree of overlap between the IGC’s
gene selection and the gene lists chosen by experts,
supporting the viability of our system.
Together with the scalability and repeatability enabled by its
automation, the IGC system greatly improves the gene panel
selection process and therefore advances targeted genomic
studies.Algorithmically Optimized Gene Selection for Targeted Clinical Sequencing Panels

Algorithmically Optimized Gene Selection for Targeted Clinical Sequencing PanelsThermo Fisher Scientific
More Related Content
What's hot
ASHG 2016 PresentationA Retrospective Analysis of Exome Sequencing Cases Using the GenePool™ Genomi...

A Retrospective Analysis of Exome Sequencing Cases Using the GenePool™ Genomi...Antoaneta Vladimirova
Presentation from the ECDC expert consultation on Whole Genome Sequencing organised by the European Centre of Disease Prevention and Control - Stockholm, 19 November 2015Overview of the ECDC whole genome sequencing strategy

Overview of the ECDC whole genome sequencing strategyEuropean Centre for Disease Prevention and Control (ECDC)
What's hot (20)
Andrew Hudgens - Antimicrobial Resistance Surveillance and Reporting at Food ...

Andrew Hudgens - Antimicrobial Resistance Surveillance and Reporting at Food ...
Interpreting genomic variation and phylogenetic trees to understand disease t...

Interpreting genomic variation and phylogenetic trees to understand disease t...
A Retrospective Analysis of Exome Sequencing Cases Using the GenePool™ Genomi...

A Retrospective Analysis of Exome Sequencing Cases Using the GenePool™ Genomi...
Real-Time Genome Sequencing of Resistant Bacteria Provides Precision Infectio...

Real-Time Genome Sequencing of Resistant Bacteria Provides Precision Infectio...
Bioinformatics-driven discovery of EGFR mutant Lung Cancer

Bioinformatics-driven discovery of EGFR mutant Lung Cancer
ClinVar: Aggregating Data to Improve Variant Interpretation - Melissa Landrum

ClinVar: Aggregating Data to Improve Variant Interpretation - Melissa Landrum
Overview of the ECDC whole genome sequencing strategy

Overview of the ECDC whole genome sequencing strategy
Dr. David Baumert - Swine Production Infection Chain

Dr. David Baumert - Swine Production Infection Chain
Genomics in Society: Genomics, Preventive Medicine, and Society

Genomics in Society: Genomics, Preventive Medicine, and Society
Monarch Initiative Poster - Rare Disease Symposium 2015

Monarch Initiative Poster - Rare Disease Symposium 2015
Whole genome sequencing as a starting point to understanding antimicrobial re...

Whole genome sequencing as a starting point to understanding antimicrobial re...
Dr. Jayaveeramuthu Nirmala - Vaccination as One of the Drivers of Influenza G...

Dr. Jayaveeramuthu Nirmala - Vaccination as One of the Drivers of Influenza G...
Similar to poster-final
Selection of genes to include in genomic studies of disease
remains a difficult task. Current methods rely on expert opinion
or manual search engine use. With these methods, the
process and result are neither repeatable nor scalable. To
remedy this situation, we created the Informative Genetic
Content (IGC) system, which enables the algorithmic selection
of genes for inclusion in such studies, given one or more
diseases to target.
The IGC system stands on three components: a database
associating diseases with genes and other diseases, an
algorithm to rank the genes under consideration for inclusion in
a panel, and a module that clusters genes by families of
diseases. The first component, the database, maps diseases
to associated genes and scores each of these mappings
according to the strength of the relationship. The database also
maps diseases to other diseases, such that groups of diseases
or hierarchical relationships between diseases can be
identified. The second component enables the ranking of
candidate genes when multiple diseases are of interest. The
algorithm accounts for the common situation where two or
more diseases are associated with the same gene with varying
strengths of association, weighting and combining the scores
across the diseases associated with each gene. The final
component, the gene clustering module, groups genes by
pathogenic pathways, should the user want to consider
targeting a broader family of diseases affected by a closely
related set of genes.
We validated the IGC system through comparisons of our
automated gene selections with expertly curated gene panel
designs. We found a high degree of overlap between the IGC’s
gene selection and the gene lists chosen by experts,
supporting the viability of our system.
Together with the scalability and repeatability enabled by its
automation, the IGC system greatly improves the gene panel
selection process and therefore advances targeted genomic
studies.Algorithmically Optimized Gene Selection for Targeted Clinical Sequencing Panels

Algorithmically Optimized Gene Selection for Targeted Clinical Sequencing PanelsThermo Fisher Scientific
Journal Club presentation. Infectious Diseases rotation (R2).
Clinical Microbiology Residency Program
KFHU-Alkhobar( Journal Club ) Procalcitonin as a diagnostic biomarker of sepsis: A tertiar...

( Journal Club ) Procalcitonin as a diagnostic biomarker of sepsis: A tertiar...Abdullatif Al-Rashed
Similar to poster-final (20)
Algorithmically Optimized Gene Selection for Targeted Clinical Sequencing Panels

Algorithmically Optimized Gene Selection for Targeted Clinical Sequencing Panels
Advanced biotechnological tools for human health care dr shiv om pratap

Advanced biotechnological tools for human health care dr shiv om pratap
Epstein-Barr virus genetic variants are associated with multiple sclerosis.

Epstein-Barr virus genetic variants are associated with multiple sclerosis.
Human Disease Ontology Project presented at ISB's Biocurator meeting April 2014

Human Disease Ontology Project presented at ISB's Biocurator meeting April 2014
A New Generation Of Mechanism-Based Biomarkers For The Clinic

A New Generation Of Mechanism-Based Biomarkers For The Clinic
NGS for Infectious Disease Diagnostics: An Opportunity for Growth 

NGS for Infectious Disease Diagnostics: An Opportunity for Growth
IOSR Journal of Pharmacy (IOSRPHR), www.iosrphr.org, call for paper, research...

IOSR Journal of Pharmacy (IOSRPHR), www.iosrphr.org, call for paper, research...
Gene Therapy - A Novel Approch in Medical Treatment

Gene Therapy - A Novel Approch in Medical Treatment
The 'omics' revolution: How will it improve our understanding of infections a...

The 'omics' revolution: How will it improve our understanding of infections a...
Establishment and analysis of a disease risk prediction model for chronic kid...

Establishment and analysis of a disease risk prediction model for chronic kid...
( Journal Club ) Procalcitonin as a diagnostic biomarker of sepsis: A tertiar...

( Journal Club ) Procalcitonin as a diagnostic biomarker of sepsis: A tertiar...
More from John Schrom
More from John Schrom (7)
poster-final
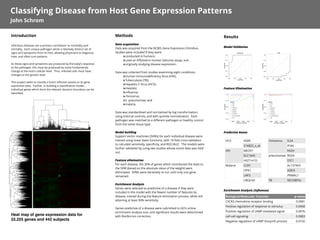
- 1. Heat map of gene expression data for 22,225 genes and 442 subjects Introduction Infectious diseases are a primary contributor to morbidity and mortality. Each unique pathogen elicits a relatively distinct set of signs and symptoms from its host, allowing physicians to diagnose, treat, and often cure patients. As these signs and symptoms are produced by the body’s response to the pathogen, this must be produced by some fundamental change at the host’s cellular level. Thus, infected cells must have changes at the genetic level. This project seeks to classify a host’s infection based on its gene expression data. Further, in building a classification model, individual genes which form the relevant decision boundary can be identified. Results Model Validation Feature Elimination Predictive Genes Enrichment Analysis (influenza) Methods Data acquisition Data was acquired from the NCBI’s Gene Expression Omnibus. Studies were included if they were: ‣conducted in humans; ‣used an Affymetrix Human Genome assay; and ‣originally studying disease expression. Data was collected from studies examining eight conditions: ‣Human Immunodeficiency Virus (HIV); ‣Tuberculosis (TB); ‣Hepatitis C Virus (HCV); ‣measles; ‣influenza; ‣rhinovirus; ‣S. pneumoniae; and ‣malaria. Data was standardized and normalized by log transformation, using internal controls, and with quintile normalization. Each pathogen was matched to a different pathogen or healthy control from the same tissue type. Model building Support vector machines (SVMs) for each individual disease were trained using linear basis functions, with 10-fold cross-validation to calculate sensitivity, specificity, and ROC/AUC. The models were further validated by using two studies whose entire data was held out. Feature elimination For each disease, the 30% of genes which contributed the least to the SVM (based on the absolute value of the weight) were eliminated. SVMs were iteratively re-run until only one gene remained. Enrichment Analysis Genes were selected as predictive of a disease if they were included in the model with the fewest number of features by disease, trained during the feature elimination process, while still attaining at least 90% sensitivity. Genes predictive of a disease were submitted to GO’s online enrichment analysis tool, and significant results were determined with Bonferroni correction. Classifying Disease from Host Gene Expression Patterns John Schrom HCV ASB9 216822_x_at HIV MEOX1 SLC16A5 HIST1H1D Malaria CUX1 ITPK1 LAP3 UBQLN2 rhinovirus IL24 IFI44 FA2H pneumoniae RGS4 STC1 AL137403 ADD3 PNMAL1 TB SECISBP2L Biological/Molecular Function p-value CXCR3 chemokine receptor binding 0.0081 Positive regulation of response to stimulus 0.0068 Positive regulation of cAMP-mediated signal 0.0076 cell-cell signaling 0.0083 Negative regulation of cAMP biosynth process 0.0102