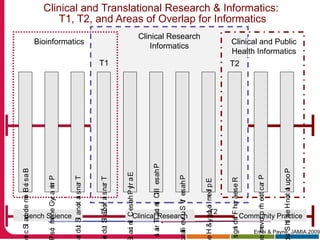
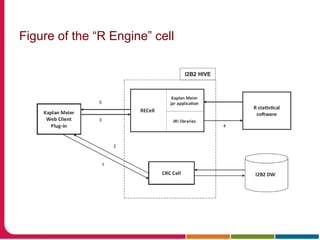
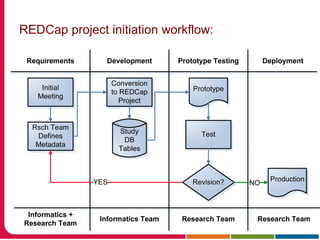
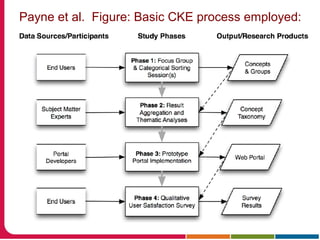
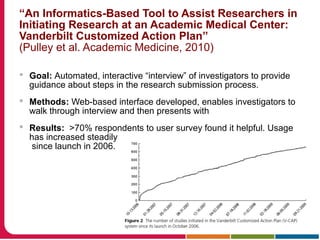
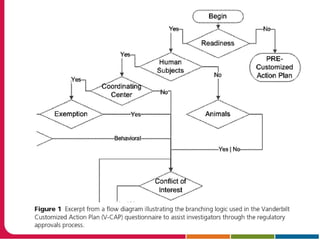
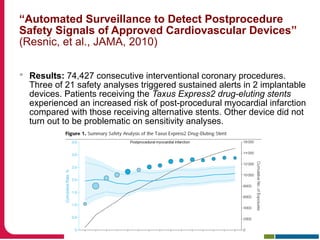
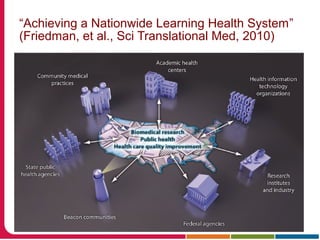
This document discusses Peter Embi's presentation on clinical research informatics. The presentation included summaries of 22 research papers on topics like data warehousing and knowledge discovery, researcher support and resources, and recruitment informatics. It also discussed ongoing efforts to integrate informatics approaches and resources to support clinical and translational research.


![Source of Content for Session
Literature review:
Initial search by MESH terms:
("Biomedical Research"[Mesh] NOT "Genetic
Research"[Mesh]) NOT "Translational Research"[Mesh]) AND
"Informatics"[Mesh] AND "2009/01/10"[PDat] :
"2011/03/09"[Pdat]
Resulted in 87 articles; 35 were CRI relevant
Additional 51 articles through:
Recommendations from selected colleagues
Other keyword searches using terms like:
Clinical Trials, Clinical Research, Informatics, Translational, Data
Warehouse
Result = 87 total CRI relevant
From those, ~22 will be presented](https://image.slidesharecdn.com/embicri-review-2011-160821061801/85/Peter-Embi-s-2011-AMIA-CRI-Year-in-Review-3-320.jpg)