NCI systems epidemiology 03012019
•
0 likes•202 views
Need for systems approaches to dissect environment and genetics in disease
Report
Share
Report
Share
Download to read offline
Recommended
Chirag patel unite for sight 041418

Talk at Unite for Sight on 4/14/18 - https://www.uniteforsight.org/conference/
EWAS and the exposome: Mt Sinai in Brescia 052119

Talk from Brescia, Italy, on the analyzing the complex exposome.
Bioinformatics Strategies for Exposome 100416

Talk given at International Society of Exposure Research, Utrecht, NL., 2016
Repurposing large datasets to dissect exposomic (and genomic) contributions i...

Talk at CDC Public Health Genomics on 2/22/16
Biomedical Informatics 706: Precision Medicine with exposures

Lecture for Zak Kohane's precision medicine course at HMS
Recommended
Chirag patel unite for sight 041418

Talk at Unite for Sight on 4/14/18 - https://www.uniteforsight.org/conference/
EWAS and the exposome: Mt Sinai in Brescia 052119

Talk from Brescia, Italy, on the analyzing the complex exposome.
Bioinformatics Strategies for Exposome 100416

Talk given at International Society of Exposure Research, Utrecht, NL., 2016
Repurposing large datasets to dissect exposomic (and genomic) contributions i...

Talk at CDC Public Health Genomics on 2/22/16
Biomedical Informatics 706: Precision Medicine with exposures

Lecture for Zak Kohane's precision medicine course at HMS
Data analytics to support exposome research course slides

We present new publicly available tools to bootstrap your own data-driven investigations to correlate the environment with phenotype. Course materials here: http://www.chiragjpgroup.org/exposome-analytics-course/
Informatics and data analytics to support for exposome-based discovery

International Society of Exposure Science 10/20/2015
Methods to enhance the validity of precision guidelines emerging from big data

A talk on the use (and misuse) of using observational big data streams for medical guideline development.
Correlation globes of the exposome 2016

Talk given at International Society of Exposure Research, Utrecht, NL., 2016
Searching for predictors of male fecundity

What semen-realted phenotypes are prognostic of male fecundity? Check out our recent work with the LIFE study here.
Japanese Environmental Children's Study and Data-driven E

Talk at IEA in Tokyo, Japan for the JECS collaborators
dkNET Webinar: Illuminating The Druggable Genome With Pharos 10/23/2020

Abstract
Pharos (https://pharos.nih.gov/) is an integrated web-based informatics platform for the analysis of data aggregated by the Illuminating the Druggable Genome (IDG) Knowledge Management Center, an NIH Common Fund initiative. The current version of Pharos (as of October 2019) spans 20,244 proteins in the human proteome, 19,880 disease and phenotype associations, and 226,829 ChEMBL compounds. This resource not only collates and analyzes data from over 60 high-quality resources to generate these types, but also uses text indexing to find less apparent connections between targets, and has recently begun to collaborate with institutions that generate data and resources. Proteins are ranked according to a knowledge-based classification system, which can help researchers to identify less studied “dark” targets that could be potentially further illuminated. This is an important process for both drug discovery and target validation, as more knowledge can accelerate target identification, and previously understudied proteins can serve as novel targets in drug discovery. In this webinar, Dr. Tudor Oprea will introduce how to use Pharos to find targets of interest for drug discovery.
The top 3 key questions that Pharos can answer:
1. What are the novel drug targets that may play a role in a specific disease?
2. What are the diseases that are related directly or indirectly to a drug target?
3. Find researchers that are related directly or indirectly to a drug target.
Presenter: Tudor Oprea, MD, PhD, Professor of Medicine, Chief of Translational Informatics Division & Internal Medicine, University of New Mexico
dkNET Webinar Information: https://dknet.org/about/webinar
Open zika presentation 

These slides provide a complete and upto date analysis of the project and complement the recent article in PLOS Neglected Tropical Diseases
Identification of PFOA linked metabolic diseases by crossing databases

The increasing amount of biological data makes possible their interpretation more accurate and richer than never before. Various way of representations and interpretations of the links between those data have been applied or developed consequently to these new elements which can be taken into account in diagnostics and soon in personalized medicine. The aim of this student project was to cross data coming from various databases to be able to link Perfluorooctaoic Acid (PFOA) to one or more human phenotypes and metabolic diseases. Our approach makes possible an easy and confident interpretation on the data kept and also allow us to rank diseases linked according to their risk of correlation to a specific set of proteins.
Real-Time Genome Sequencing of Resistant Bacteria Provides Precision Infectio...

http://www.fao.org/about/meetings/wgs-on-food-safety-management/en/
Real-Time Genome Sequencing of Resistant Bacteria Provides Precision Infection Control in an Institutional Setting. Presentation from the Technical Meeting on the impact of Whole Genome Sequencing (WGS) on food safety management and GMI-9, 23-25 May 2016, Rome, Italy.
Using In Silico Tools in Repurposing Drugs for Neglected and Orphan Diseases

Presentation given at the AAPS meeting 2016 in Denver in a session on repurposing. I describes several recent published studies and unpublished work.
dkNET Webinar: Population-Based Approaches to Investigate Endocrine Communica...

Abstract
Mechanisms of inter-organ signaling have been established as hallmarks of nearly every pathophysiologic condition, where many exist as related and complex diseases. While significant work has been focused on understanding how individual cell types contribute and respond to specific perturbations related to common, complex disease, an equally-important but relatively less-explored question involves how relationships between organs are altered in the context of an integrated living organism. Current technical advances, such as proteomic analysis of plasma or conditioned media, have allowed for a more unbiased visualization and discovery of additional inter-tissue signaling molecules. However, one important feature which is lacking from these approaches is the ability to gain insight as to the function, mechanisms of action and target tissue(s) of relevant molecules. To begin to address these constraints, we initially developed a correlation-based bioinformatics framework which uses multi-tissue gene expression and/or proteomic data, as well as publicly available resources to statistically rank and functionally annotate endocrine proteins involved in tissue cross-talk. Using this approach, we identified many known and experimentally validated several novel inter-tissue circuits. This was this first study to directly link an endocrine-focused bioinformatics pipeline from population data directly to experimentally-validated mechanisms of inter-tissue communication. While these validations provide strong support for exploiting natural variation to discover new modes of communication, these serve as simple proof-of-principle studies and, thus, have promising potential for expansion. Some of these will be discussed during the presentation.
Presenter: Marcus Seldin, Ph.D. Assistant Professor, Biological Chemistry, University of California Irvine
Upcoming webinars schedule: https://dknet.org/about/webinar
Data Visualization in Biomedical Sciences: More than Meets the Eye

In science, data visualization serves two primary purposes. The first is to explore data sets interactively and the second is to communicate discoveries. However, the requirements for visualizations employed in these activities are very different. Therefore, the software tools used for these purposes are typically disconnected, creating significant challenges for reproducibility and effective communication of discoveries in data-driven biomedical science. In this presentation, I will address how a new approach to creating data visualization tools can connect data analysts and other stakeholders inside and outside the scientific community. I will introduce and demonstrate the "Vistories" approach that was motivated by these question.
Presented at the 5th Cancer Research UK Big Data Analytics Conference on Data Visualization.
Human Disease Ontology Project presented at ISB's Biocurator meeting April 2014

The Human Disease Ontology (DO), organized as a directed acyclic graph, represents a knowledge base of inherited, environmental, infectious diseases (http://www.disease-ontology.org). DO's textual definition model incorporates a semi-structured format describing the disease etiology built to capture the complex nature of human disease etiology within a is_a hierarchy. DO includes disease concepts for cancer, metabolic disease, infectious disease, mental disorders, genetic disease and syndromes. DO contains disease definitions, external references to resources including ICD, NCI-metathesaurus, SNOMED, MeSH and OMIM and extended relationships that conform to OBO guidelines. DO provides a central ‘switchboard’ for connecting resources, datasets, and computational tools that include disease terms or relationships.
Introduction to Gene Mining Part A: BLASTn-off!

In this lesson, students will learn to use bioinformatics portals and tools to mine plant versions of human genes. Student handout and teacher resource materials are available at www.Araport.org, Teaching Resources (Community tab). Suitable for grades 9-12 or first year undergraduate students.
Introduction to Gene Mining: Part B: How similar are plant and animal version...

In this lesson, students will navigate BLASTp and www.Araport.org to determine whether plant and animal versions of genes and proteins are homologous. Student handout and teacher resources are available at www.Araport.org, teacher resources page (under Community). Suitable for grades 9-12 or first year undergraduate students.
More Related Content
What's hot
Data analytics to support exposome research course slides

We present new publicly available tools to bootstrap your own data-driven investigations to correlate the environment with phenotype. Course materials here: http://www.chiragjpgroup.org/exposome-analytics-course/
Informatics and data analytics to support for exposome-based discovery

International Society of Exposure Science 10/20/2015
Methods to enhance the validity of precision guidelines emerging from big data

A talk on the use (and misuse) of using observational big data streams for medical guideline development.
Correlation globes of the exposome 2016

Talk given at International Society of Exposure Research, Utrecht, NL., 2016
Searching for predictors of male fecundity

What semen-realted phenotypes are prognostic of male fecundity? Check out our recent work with the LIFE study here.
Japanese Environmental Children's Study and Data-driven E

Talk at IEA in Tokyo, Japan for the JECS collaborators
dkNET Webinar: Illuminating The Druggable Genome With Pharos 10/23/2020

Abstract
Pharos (https://pharos.nih.gov/) is an integrated web-based informatics platform for the analysis of data aggregated by the Illuminating the Druggable Genome (IDG) Knowledge Management Center, an NIH Common Fund initiative. The current version of Pharos (as of October 2019) spans 20,244 proteins in the human proteome, 19,880 disease and phenotype associations, and 226,829 ChEMBL compounds. This resource not only collates and analyzes data from over 60 high-quality resources to generate these types, but also uses text indexing to find less apparent connections between targets, and has recently begun to collaborate with institutions that generate data and resources. Proteins are ranked according to a knowledge-based classification system, which can help researchers to identify less studied “dark” targets that could be potentially further illuminated. This is an important process for both drug discovery and target validation, as more knowledge can accelerate target identification, and previously understudied proteins can serve as novel targets in drug discovery. In this webinar, Dr. Tudor Oprea will introduce how to use Pharos to find targets of interest for drug discovery.
The top 3 key questions that Pharos can answer:
1. What are the novel drug targets that may play a role in a specific disease?
2. What are the diseases that are related directly or indirectly to a drug target?
3. Find researchers that are related directly or indirectly to a drug target.
Presenter: Tudor Oprea, MD, PhD, Professor of Medicine, Chief of Translational Informatics Division & Internal Medicine, University of New Mexico
dkNET Webinar Information: https://dknet.org/about/webinar
Open zika presentation 

These slides provide a complete and upto date analysis of the project and complement the recent article in PLOS Neglected Tropical Diseases
Identification of PFOA linked metabolic diseases by crossing databases

The increasing amount of biological data makes possible their interpretation more accurate and richer than never before. Various way of representations and interpretations of the links between those data have been applied or developed consequently to these new elements which can be taken into account in diagnostics and soon in personalized medicine. The aim of this student project was to cross data coming from various databases to be able to link Perfluorooctaoic Acid (PFOA) to one or more human phenotypes and metabolic diseases. Our approach makes possible an easy and confident interpretation on the data kept and also allow us to rank diseases linked according to their risk of correlation to a specific set of proteins.
Real-Time Genome Sequencing of Resistant Bacteria Provides Precision Infectio...

http://www.fao.org/about/meetings/wgs-on-food-safety-management/en/
Real-Time Genome Sequencing of Resistant Bacteria Provides Precision Infection Control in an Institutional Setting. Presentation from the Technical Meeting on the impact of Whole Genome Sequencing (WGS) on food safety management and GMI-9, 23-25 May 2016, Rome, Italy.
Using In Silico Tools in Repurposing Drugs for Neglected and Orphan Diseases

Presentation given at the AAPS meeting 2016 in Denver in a session on repurposing. I describes several recent published studies and unpublished work.
dkNET Webinar: Population-Based Approaches to Investigate Endocrine Communica...

Abstract
Mechanisms of inter-organ signaling have been established as hallmarks of nearly every pathophysiologic condition, where many exist as related and complex diseases. While significant work has been focused on understanding how individual cell types contribute and respond to specific perturbations related to common, complex disease, an equally-important but relatively less-explored question involves how relationships between organs are altered in the context of an integrated living organism. Current technical advances, such as proteomic analysis of plasma or conditioned media, have allowed for a more unbiased visualization and discovery of additional inter-tissue signaling molecules. However, one important feature which is lacking from these approaches is the ability to gain insight as to the function, mechanisms of action and target tissue(s) of relevant molecules. To begin to address these constraints, we initially developed a correlation-based bioinformatics framework which uses multi-tissue gene expression and/or proteomic data, as well as publicly available resources to statistically rank and functionally annotate endocrine proteins involved in tissue cross-talk. Using this approach, we identified many known and experimentally validated several novel inter-tissue circuits. This was this first study to directly link an endocrine-focused bioinformatics pipeline from population data directly to experimentally-validated mechanisms of inter-tissue communication. While these validations provide strong support for exploiting natural variation to discover new modes of communication, these serve as simple proof-of-principle studies and, thus, have promising potential for expansion. Some of these will be discussed during the presentation.
Presenter: Marcus Seldin, Ph.D. Assistant Professor, Biological Chemistry, University of California Irvine
Upcoming webinars schedule: https://dknet.org/about/webinar
Data Visualization in Biomedical Sciences: More than Meets the Eye

In science, data visualization serves two primary purposes. The first is to explore data sets interactively and the second is to communicate discoveries. However, the requirements for visualizations employed in these activities are very different. Therefore, the software tools used for these purposes are typically disconnected, creating significant challenges for reproducibility and effective communication of discoveries in data-driven biomedical science. In this presentation, I will address how a new approach to creating data visualization tools can connect data analysts and other stakeholders inside and outside the scientific community. I will introduce and demonstrate the "Vistories" approach that was motivated by these question.
Presented at the 5th Cancer Research UK Big Data Analytics Conference on Data Visualization.
Human Disease Ontology Project presented at ISB's Biocurator meeting April 2014

The Human Disease Ontology (DO), organized as a directed acyclic graph, represents a knowledge base of inherited, environmental, infectious diseases (http://www.disease-ontology.org). DO's textual definition model incorporates a semi-structured format describing the disease etiology built to capture the complex nature of human disease etiology within a is_a hierarchy. DO includes disease concepts for cancer, metabolic disease, infectious disease, mental disorders, genetic disease and syndromes. DO contains disease definitions, external references to resources including ICD, NCI-metathesaurus, SNOMED, MeSH and OMIM and extended relationships that conform to OBO guidelines. DO provides a central ‘switchboard’ for connecting resources, datasets, and computational tools that include disease terms or relationships.
What's hot (20)
Data analytics to support exposome research course slides

Data analytics to support exposome research course slides
Informatics and data analytics to support for exposome-based discovery

Informatics and data analytics to support for exposome-based discovery
Methods to enhance the validity of precision guidelines emerging from big data

Methods to enhance the validity of precision guidelines emerging from big data
Japanese Environmental Children's Study and Data-driven E

Japanese Environmental Children's Study and Data-driven E
dkNET Webinar: Illuminating The Druggable Genome With Pharos 10/23/2020

dkNET Webinar: Illuminating The Druggable Genome With Pharos 10/23/2020
Identification of PFOA linked metabolic diseases by crossing databases

Identification of PFOA linked metabolic diseases by crossing databases
Real-Time Genome Sequencing of Resistant Bacteria Provides Precision Infectio...

Real-Time Genome Sequencing of Resistant Bacteria Provides Precision Infectio...
Using In Silico Tools in Repurposing Drugs for Neglected and Orphan Diseases

Using In Silico Tools in Repurposing Drugs for Neglected and Orphan Diseases
dkNET Webinar: Population-Based Approaches to Investigate Endocrine Communica...

dkNET Webinar: Population-Based Approaches to Investigate Endocrine Communica...
Data Visualization in Biomedical Sciences: More than Meets the Eye

Data Visualization in Biomedical Sciences: More than Meets the Eye
Human Disease Ontology Project presented at ISB's Biocurator meeting April 2014

Human Disease Ontology Project presented at ISB's Biocurator meeting April 2014
Similar to NCI systems epidemiology 03012019
Introduction to Gene Mining Part A: BLASTn-off!

In this lesson, students will learn to use bioinformatics portals and tools to mine plant versions of human genes. Student handout and teacher resource materials are available at www.Araport.org, Teaching Resources (Community tab). Suitable for grades 9-12 or first year undergraduate students.
Introduction to Gene Mining: Part B: How similar are plant and animal version...

In this lesson, students will navigate BLASTp and www.Araport.org to determine whether plant and animal versions of genes and proteins are homologous. Student handout and teacher resources are available at www.Araport.org, teacher resources page (under Community). Suitable for grades 9-12 or first year undergraduate students.
Fixing the leaks in the pipeline from public genomics data to the clinic

A talk about improving reproducibility, simplifying genomic machine learning, and using the resulting predictors to improve power in clinical trials.
Impact tech: Opportunities in Clean Meat and Cellular Agriculture by Liz Specht

Slides from the Impact.tech seminar on Opportunities in Clean Meat and Cellular Agriculture. The seminar was taught by Liz Specht, a Senior Scientist with the Good Food Institute. The Good Food Institute is a non-profit organization advancing plant-based and clean meat food technology.
Cellular agriculture is an interdisciplinary branch of science at the intersection of medicine and farming. Cellular agriculture capitalizes on breakthroughs in tissue-engineering, material sciences, bioengineering, and synthetic biology to design new ways of producing existing agricultural products like milk, meat, fragrances, and rhino horn from cells and microorganisms [instead of whole animals].
Mining Phenotypes: How to set up a reverse genetics experiment with an Arabid...

In this lesson, students will mine data from Araport.org to design and propose a reverse genetics experiment using a known Arabidopsis mutant. They will select a treatment to reveal phenotypic dfifferences between wild type and mutant Arabidopsis. Student handout and teacher resources are available at www.Araport.org, teacher resources. Suitable for grades 9-12 or first year undergraduate students.
Genetic Mapping

Biology Molecular
University of Haifa. (2015, August 13). Big data and the social character of genes. ScienceDaily. Retrieved September 12, 2015 from www.sciencedaily.com/releases/2015/08/150813084035.htm
Lawrence Berkeley National Laboratory. (2015, August 4). Keeping algae from stressing out. ScienceDaily. Retrieved September 12, 2015 from www.sciencedaily.com/releases/2015/08/150804144013.htm
Martínez Sánchez, Lina María. Biología Molecular. 8.ed. Medellín. UPB. Facultad de Medicina.
9212017 9222017 gmo online documention lethality review MANTICORE

Manticore Group, MT. Reviewed searchable online documents on the lethality research on GMOs. These were the ones of value found at time of the review on the subject. A notorious Oregon Lab report originally the subject of the review was not discover-able online at this time that referenced lab work commissioned by Russian interests in Oregon on the subject.
Genomics in Society: Genomics, Cellular Networks, Preventive Medicine, and So...

10.10.06
Guest Lecture
UCSD Medical and Pharmaceutical Students Foundations of Human Biology--Lecture #41
Title: Genomics in Society: Genomics, Cellular Networks, Preventive Medicine, and Society
La Jolla, CA
Standards for public health genomic epidemiology - Biocuration 2015

A presentation introducing genomic epidemiology and its application in public health. It also explains the need for standards to support the Canadian Integrated Rapid Infectious Disease Analysis platform which implements genomic epidemiology analyses for detection and investigation of infectious disease outbreaks caused by food-borne pathogens.
Genetic Technologies Biotech Showcase Presentation - October

Genetic Technologies Limited (ASX: GTG; Nasdaq: GENE), a diversified molecular diagnostics company. GTG offers cancer predictive testing and assessment tools to help physicians proactively manage patient health. The Company’s lead products GeneType for Breast Cancer and GeneType for Colorectal Cancer are clinically validated risk assessment tests and are first in class. Genetic Technologies is developing a pipeline of risk assessment products. Learn more at GENETechinfo.com.
Genetic Engineering in Indian Agriculture - How Bad is it

Genetic Engineering in Indian Agriculture - How Bad is it
Forage Agriculture's Future in a Changing Climate

Presented to the 2018 Crop Science Society of America,
Baltimore, Maryland, November 6, 2018
Impact.tech: Opportunities in Plant-based Food Technologies by Liz Specht

Slides from the Impact.tech seminar on Opportunities in Plant-based Food Technologies. The seminar was taught by Liz Specht, a Senior Scientist with the Good Food Institute. The Good Food Institute is a non-profit organization advancing plant-based and clean meat food technology.
The plant-based foods sector has experienced tremendous growth and innovation as plant-based alternatives to animal products are increasingly adopted into the diets of mainstream consumers seeking healthier or more sustainable options. These products have come a long way in replicating the taste, texture, and mouthfeel of their animal-based counterparts. However, there is still ample room for food technology and product development to enable greater inroads into mainstream markets. The seminar discussed opportunities all across the product development pipeline - from genetic mapping to develop better plant protein crop strains, to novel protein isolation and functionalization methods, to mechanical processing and formulation to better replicate the structure and flavor of meat.
Similar to NCI systems epidemiology 03012019 (20)
Introduction to Gene Mining: Part B: How similar are plant and animal version...

Introduction to Gene Mining: Part B: How similar are plant and animal version...
Fixing the leaks in the pipeline from public genomics data to the clinic

Fixing the leaks in the pipeline from public genomics data to the clinic
Impact tech: Opportunities in Clean Meat and Cellular Agriculture by Liz Specht

Impact tech: Opportunities in Clean Meat and Cellular Agriculture by Liz Specht
Mining Phenotypes: How to set up a reverse genetics experiment with an Arabid...

Mining Phenotypes: How to set up a reverse genetics experiment with an Arabid...
9212017 9222017 gmo online documention lethality review MANTICORE

9212017 9222017 gmo online documention lethality review MANTICORE
Genomics in Society: Genomics, Cellular Networks, Preventive Medicine, and So...

Genomics in Society: Genomics, Cellular Networks, Preventive Medicine, and So...
Standards for public health genomic epidemiology - Biocuration 2015

Standards for public health genomic epidemiology - Biocuration 2015
Genetic Technologies Biotech Showcase Presentation - October

Genetic Technologies Biotech Showcase Presentation - October
Genetic Engineering in Indian Agriculture - How Bad is it

Genetic Engineering in Indian Agriculture - How Bad is it
application_of_bioinformatics_in_various_fields.ppt

application_of_bioinformatics_in_various_fields.ppt
application_of_bioinformatics_in_various_fields.ppt

application_of_bioinformatics_in_various_fields.ppt
Impact.tech: Opportunities in Plant-based Food Technologies by Liz Specht

Impact.tech: Opportunities in Plant-based Food Technologies by Liz Specht
Recently uploaded
Travis Hills' Endeavors in Minnesota: Fostering Environmental and Economic Pr...

Travis Hills of Minnesota developed a method to convert waste into high-value dry fertilizer, significantly enriching soil quality. By providing farmers with a valuable resource derived from waste, Travis Hills helps enhance farm profitability while promoting environmental stewardship. Travis Hills' sustainable practices lead to cost savings and increased revenue for farmers by improving resource efficiency and reducing waste.
Richard's aventures in two entangled wonderlands

Since the loophole-free Bell experiments of 2020 and the Nobel prizes in physics of 2022, critics of Bell's work have retreated to the fortress of super-determinism. Now, super-determinism is a derogatory word - it just means "determinism". Palmer, Hance and Hossenfelder argue that quantum mechanics and determinism are not incompatible, using a sophisticated mathematical construction based on a subtle thinning of allowed states and measurements in quantum mechanics, such that what is left appears to make Bell's argument fail, without altering the empirical predictions of quantum mechanics. I think however that it is a smoke screen, and the slogan "lost in math" comes to my mind. I will discuss some other recent disproofs of Bell's theorem using the language of causality based on causal graphs. Causal thinking is also central to law and justice. I will mention surprising connections to my work on serial killer nurse cases, in particular the Dutch case of Lucia de Berk and the current UK case of Lucy Letby.
ISI 2024: Application Form (Extended), Exam Date (Out), Eligibility

The Indian Statistical Institute (ISI) has extended its application deadline for 2024 admissions to April 2. Known for its excellence in statistics and related fields, ISI offers a range of programs from Bachelor's to Junior Research Fellowships. The admission test is scheduled for May 12, 2024. Eligibility varies by program, generally requiring a background in Mathematics and English for undergraduate courses and specific degrees for postgraduate and research positions. Application fees are ₹1500 for male general category applicants and ₹1000 for females. Applications are open to Indian and OCI candidates.
20240520 Planning a Circuit Simulator in JavaScript.pptx

Evaporation step counter work. I have done a physical experiment.
(Work in progress.)
Shallowest Oil Discovery of Turkiye.pptx

The Petroleum System of the Çukurova Field - the Shallowest Oil Discovery of Türkiye, Adana
Earliest Galaxies in the JADES Origins Field: Luminosity Function and Cosmic ...

We characterize the earliest galaxy population in the JADES Origins Field (JOF), the deepest
imaging field observed with JWST. We make use of the ancillary Hubble optical images (5 filters
spanning 0.4−0.9µm) and novel JWST images with 14 filters spanning 0.8−5µm, including 7 mediumband filters, and reaching total exposure times of up to 46 hours per filter. We combine all our data
at > 2.3µm to construct an ultradeep image, reaching as deep as ≈ 31.4 AB mag in the stack and
30.3-31.0 AB mag (5σ, r = 0.1” circular aperture) in individual filters. We measure photometric
redshifts and use robust selection criteria to identify a sample of eight galaxy candidates at redshifts
z = 11.5 − 15. These objects show compact half-light radii of R1/2 ∼ 50 − 200pc, stellar masses of
M⋆ ∼ 107−108M⊙, and star-formation rates of SFR ∼ 0.1−1 M⊙ yr−1
. Our search finds no candidates
at 15 < z < 20, placing upper limits at these redshifts. We develop a forward modeling approach to
infer the properties of the evolving luminosity function without binning in redshift or luminosity that
marginalizes over the photometric redshift uncertainty of our candidate galaxies and incorporates the
impact of non-detections. We find a z = 12 luminosity function in good agreement with prior results,
and that the luminosity function normalization and UV luminosity density decline by a factor of ∼ 2.5
from z = 12 to z = 14. We discuss the possible implications of our results in the context of theoretical
models for evolution of the dark matter halo mass function.
Salas, V. (2024) "John of St. Thomas (Poinsot) on the Science of Sacred Theol...

I Introduction
II Subalternation and Theology
III Theology and Dogmatic Declarations
IV The Mixed Principles of Theology
V Virtual Revelation: The Unity of Theology
VI Theology as a Natural Science
VII Theology’s Certitude
VIII Conclusion
Notes
Bibliography
All the contents are fully attributable to the author, Doctor Victor Salas. Should you wish to get this text republished, get in touch with the author or the editorial committee of the Studia Poinsotiana. Insofar as possible, we will be happy to broker your contact.
Unveiling the Energy Potential of Marshmallow Deposits.pdf

Unveiling the Energy Potential of Marshmallow Deposits: A Revolutionary
Breakthrough in Sustainable Energy Science
Toxic effects of heavy metals : Lead and Arsenic

Heavy metals are naturally occuring metallic chemical elements that have relatively high density, and are toxic at even low concentrations. All toxic metals are termed as heavy metals irrespective of their atomic mass and density, eg. arsenic, lead, mercury, cadmium, thallium, chromium, etc.
Deep Behavioral Phenotyping in Systems Neuroscience for Functional Atlasing a...

Functional Magnetic Resonance Imaging (fMRI) provides means to characterize brain activations in response to behavior. However, cognitive neuroscience has been limited to group-level effects referring to the performance of specific tasks. To obtain the functional profile of elementary cognitive mechanisms, the combination of brain responses to many tasks is required. Yet, to date, both structural atlases and parcellation-based activations do not fully account for cognitive function and still present several limitations. Further, they do not adapt overall to individual characteristics. In this talk, I will give an account of deep-behavioral phenotyping strategies, namely data-driven methods in large task-fMRI datasets, to optimize functional brain-data collection and improve inference of effects-of-interest related to mental processes. Key to this approach is the employment of fast multi-functional paradigms rich on features that can be well parametrized and, consequently, facilitate the creation of psycho-physiological constructs to be modelled with imaging data. Particular emphasis will be given to music stimuli when studying high-order cognitive mechanisms, due to their ecological nature and quality to enable complex behavior compounded by discrete entities. I will also discuss how deep-behavioral phenotyping and individualized models applied to neuroimaging data can better account for the subject-specific organization of domain-general cognitive systems in the human brain. Finally, the accumulation of functional brain signatures brings the possibility to clarify relationships among tasks and create a univocal link between brain systems and mental functions through: (1) the development of ontologies proposing an organization of cognitive processes; and (2) brain-network taxonomies describing functional specialization. To this end, tools to improve commensurability in cognitive science are necessary, such as public repositories, ontology-based platforms and automated meta-analysis tools. I will thus discuss some brain-atlasing resources currently under development, and their applicability in cognitive as well as clinical neuroscience.
The use of Nauplii and metanauplii artemia in aquaculture (brine shrimp).pptx

Although Artemia has been known to man for centuries, its use as a food for the culture of larval organisms apparently began only in the 1930s, when several investigators found that it made an excellent food for newly hatched fish larvae (Litvinenko et al., 2023). As aquaculture developed in the 1960s and ‘70s, the use of Artemia also became more widespread, due both to its convenience and to its nutritional value for larval organisms (Arenas-Pardo et al., 2024). The fact that Artemia dormant cysts can be stored for long periods in cans, and then used as an off-the-shelf food requiring only 24 h of incubation makes them the most convenient, least labor-intensive, live food available for aquaculture (Sorgeloos & Roubach, 2021). The nutritional value of Artemia, especially for marine organisms, is not constant, but varies both geographically and temporally. During the last decade, however, both the causes of Artemia nutritional variability and methods to improve poorquality Artemia have been identified (Loufi et al., 2024).
Brine shrimp (Artemia spp.) are used in marine aquaculture worldwide. Annually, more than 2,000 metric tons of dry cysts are used for cultivation of fish, crustacean, and shellfish larva. Brine shrimp are important to aquaculture because newly hatched brine shrimp nauplii (larvae) provide a food source for many fish fry (Mozanzadeh et al., 2021). Culture and harvesting of brine shrimp eggs represents another aspect of the aquaculture industry. Nauplii and metanauplii of Artemia, commonly known as brine shrimp, play a crucial role in aquaculture due to their nutritional value and suitability as live feed for many aquatic species, particularly in larval stages (Sorgeloos & Roubach, 2021).
Leaf Initiation, Growth and Differentiation.pdf

Leaf initiation, growth and differentiation, genetic control of leaf development.
3D Hybrid PIC simulation of the plasma expansion (ISSS-14)

3D Particle-In-Cell (PIC) algorithm,
Plasma expansion in the dipole magnetic field.
What is greenhouse gasses and how many gasses are there to affect the Earth.

What are greenhouse gasses how they affect the earth and its environment what is the future of the environment and earth how the weather and the climate effects.
Nucleic Acid-its structural and functional complexity.

This presentation explores a brief idea about the structural and functional attributes of nucleotides, the structure and function of genetic materials along with the impact of UV rays and pH upon them.
Comparing Evolved Extractive Text Summary Scores of Bidirectional Encoder Rep...

Comparing Evolved Extractive Text Summary Scores of Bidirectional Encoder Rep...University of Maribor
Slides from:
11th International Conference on Electrical, Electronics and Computer Engineering (IcETRAN), Niš, 3-6 June 2024
Track: Artificial Intelligence
https://www.etran.rs/2024/en/home-english/Recently uploaded (20)
Travis Hills' Endeavors in Minnesota: Fostering Environmental and Economic Pr...

Travis Hills' Endeavors in Minnesota: Fostering Environmental and Economic Pr...
ISI 2024: Application Form (Extended), Exam Date (Out), Eligibility

ISI 2024: Application Form (Extended), Exam Date (Out), Eligibility
20240520 Planning a Circuit Simulator in JavaScript.pptx

20240520 Planning a Circuit Simulator in JavaScript.pptx
Earliest Galaxies in the JADES Origins Field: Luminosity Function and Cosmic ...

Earliest Galaxies in the JADES Origins Field: Luminosity Function and Cosmic ...
Lateral Ventricles.pdf very easy good diagrams comprehensive

Lateral Ventricles.pdf very easy good diagrams comprehensive
Salas, V. (2024) "John of St. Thomas (Poinsot) on the Science of Sacred Theol...

Salas, V. (2024) "John of St. Thomas (Poinsot) on the Science of Sacred Theol...
Unveiling the Energy Potential of Marshmallow Deposits.pdf

Unveiling the Energy Potential of Marshmallow Deposits.pdf
In silico drugs analogue design: novobiocin analogues.pptx

In silico drugs analogue design: novobiocin analogues.pptx
Deep Behavioral Phenotyping in Systems Neuroscience for Functional Atlasing a...

Deep Behavioral Phenotyping in Systems Neuroscience for Functional Atlasing a...
The use of Nauplii and metanauplii artemia in aquaculture (brine shrimp).pptx

The use of Nauplii and metanauplii artemia in aquaculture (brine shrimp).pptx
3D Hybrid PIC simulation of the plasma expansion (ISSS-14)

3D Hybrid PIC simulation of the plasma expansion (ISSS-14)
What is greenhouse gasses and how many gasses are there to affect the Earth.

What is greenhouse gasses and how many gasses are there to affect the Earth.
Nucleic Acid-its structural and functional complexity.

Nucleic Acid-its structural and functional complexity.
Comparing Evolved Extractive Text Summary Scores of Bidirectional Encoder Rep...

Comparing Evolved Extractive Text Summary Scores of Bidirectional Encoder Rep...
NCI systems epidemiology 03012019
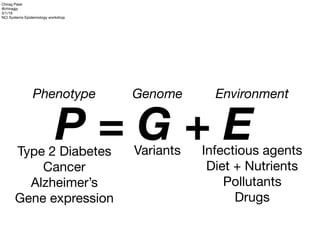
- 1. P = G + EType 2 Diabetes Cancer Alzheimer’s Gene expression Phenotype Genome Variants Environment Infectious agents Diet + Nutrients Pollutants Drugs Chirag Patel @chiragjp 3/1/19 NCI Systems Epidemiology workshop
- 2. Lakhani et al, Nature Genetics 2019http://apps.chiragjpgroup.org/catch/ 56K twins and 700K siblings in a massive health insurance cohort point to complex and elusive variation in 560 phenotypes https://rdcu.be/boZeV Chirag Patel @chiragjp 3/1/19 NCI Systems Epidemiology workshop
- 3. 56K twins and 700K siblings in a massive health insurance cohort point to complex and elusive variation in 560 phenotypes 560phenotypes Nature Genetics 2019http://apps.chiragjpgroup.org/catch/ h2=0.3 (🧬) c2=0.1 (🏠) ? https://rdcu.be/boZeV Chirag Patel @chiragjp 3/1/19 NCI Systems Epidemiology workshop
- 4. 🐘 Chirag Patel @chiragjp 3/1/19 NCI Systems Epidemiology workshop
- 5. Elephant in the room (full of system epidemiologists): Is all cancer risk random? Tomasetti and Vogelstein Science 2015 Chirag Patel @chiragjp 3/1/19 NCI Systems Epidemiology workshop
- 6. • What techniques, cohorts, and data measurement modalities to capture non-genetic variation in populations? • Does it matter? • Is it possible to develop approaches to partition genetic vs non-genetic vs random chance in disease variation? • What is “error” vs. true non-genetic signal? Chirag Patel @chiragjp 3/1/19 NCI Systems Epidemiology workshop
