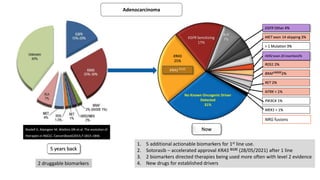
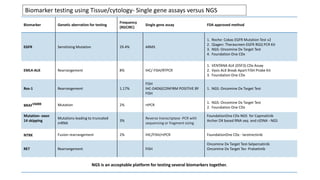
1. Molecular testing in lung cancer has identified several new biomarkers in recent years, including ALK, ROS1, MET exon 14 skipping, NTRK fusions, and KRAS G12C mutations.
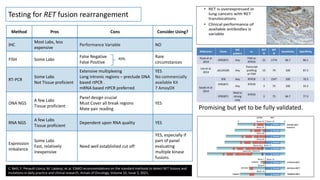
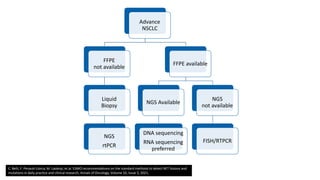
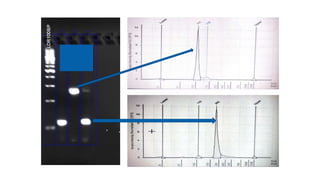
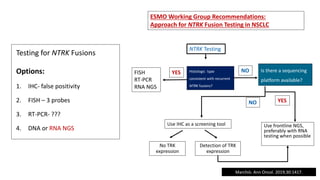
2. Next-generation sequencing has become a widely used testing method that allows for parallel testing of multiple genes and biomarkers. RNA sequencing is generally preferred over DNA sequencing for detection of fusions.
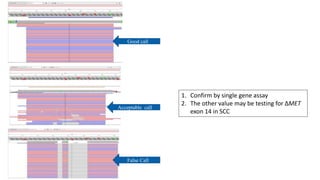
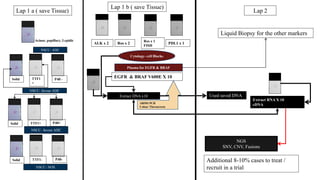
3. Single gene assays remain important options, especially for established biomarkers like EGFR mutations, but targeted NGS panels provide a more comprehensive approach and have been validated in large testing programs.