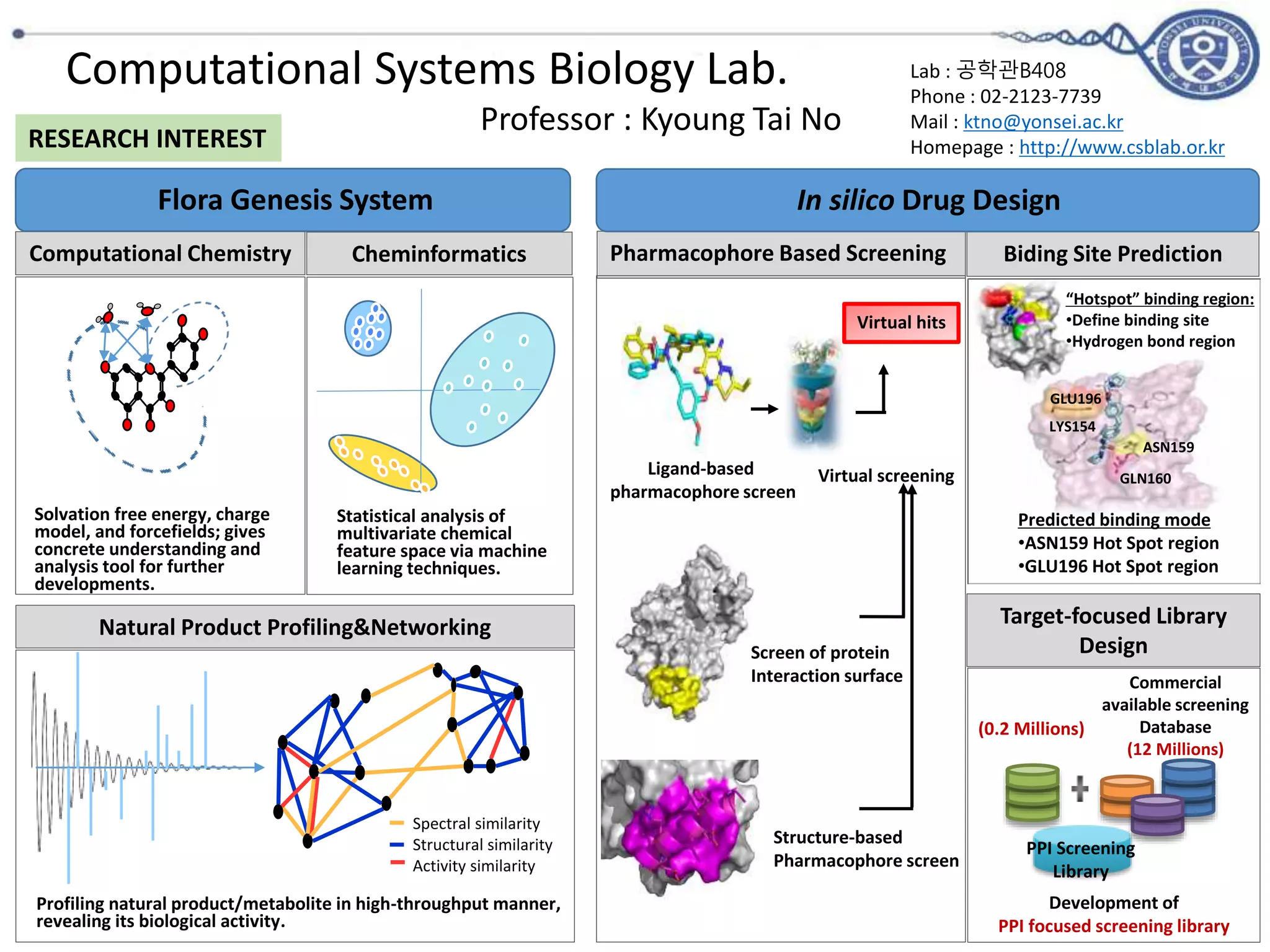
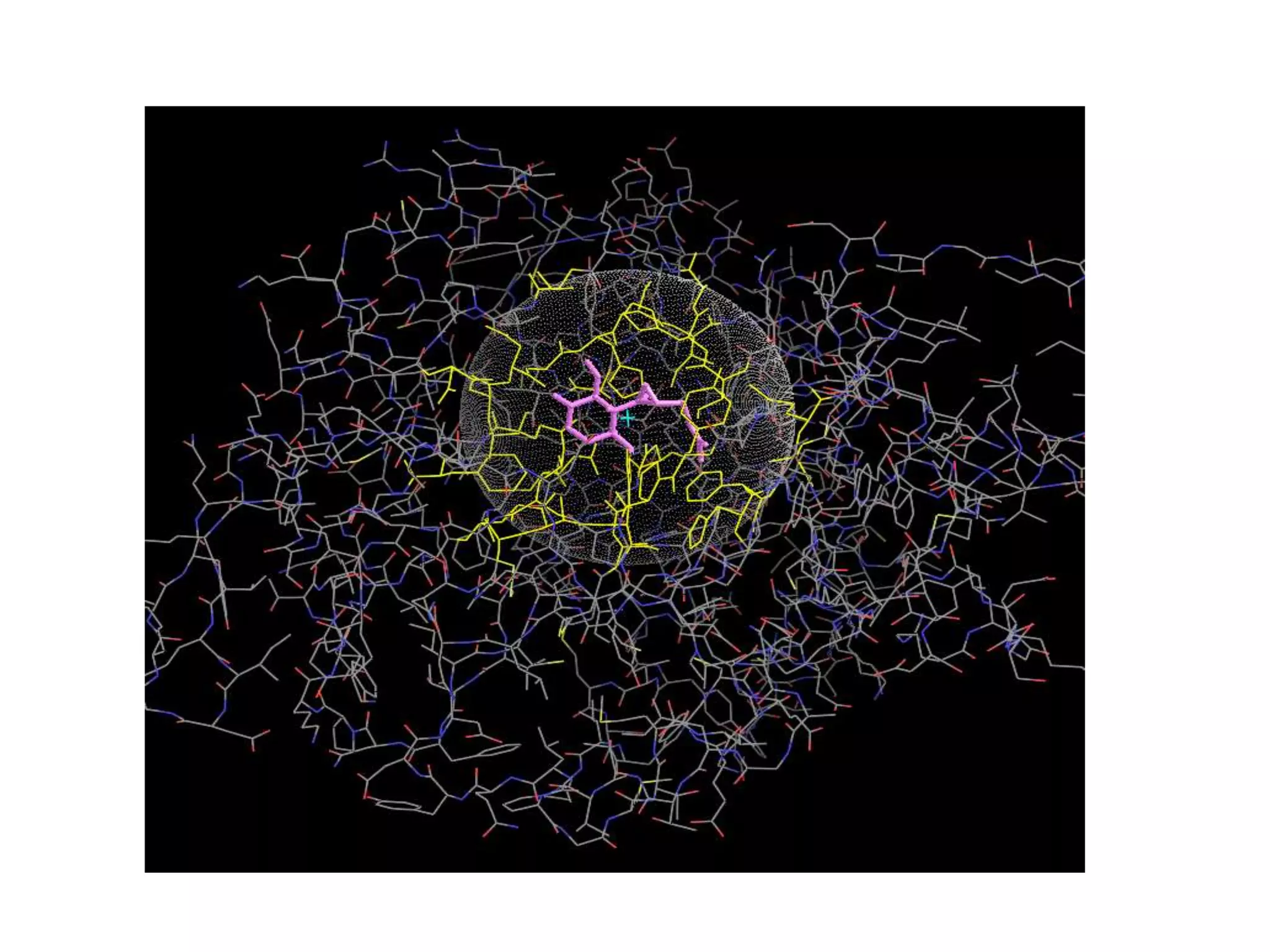
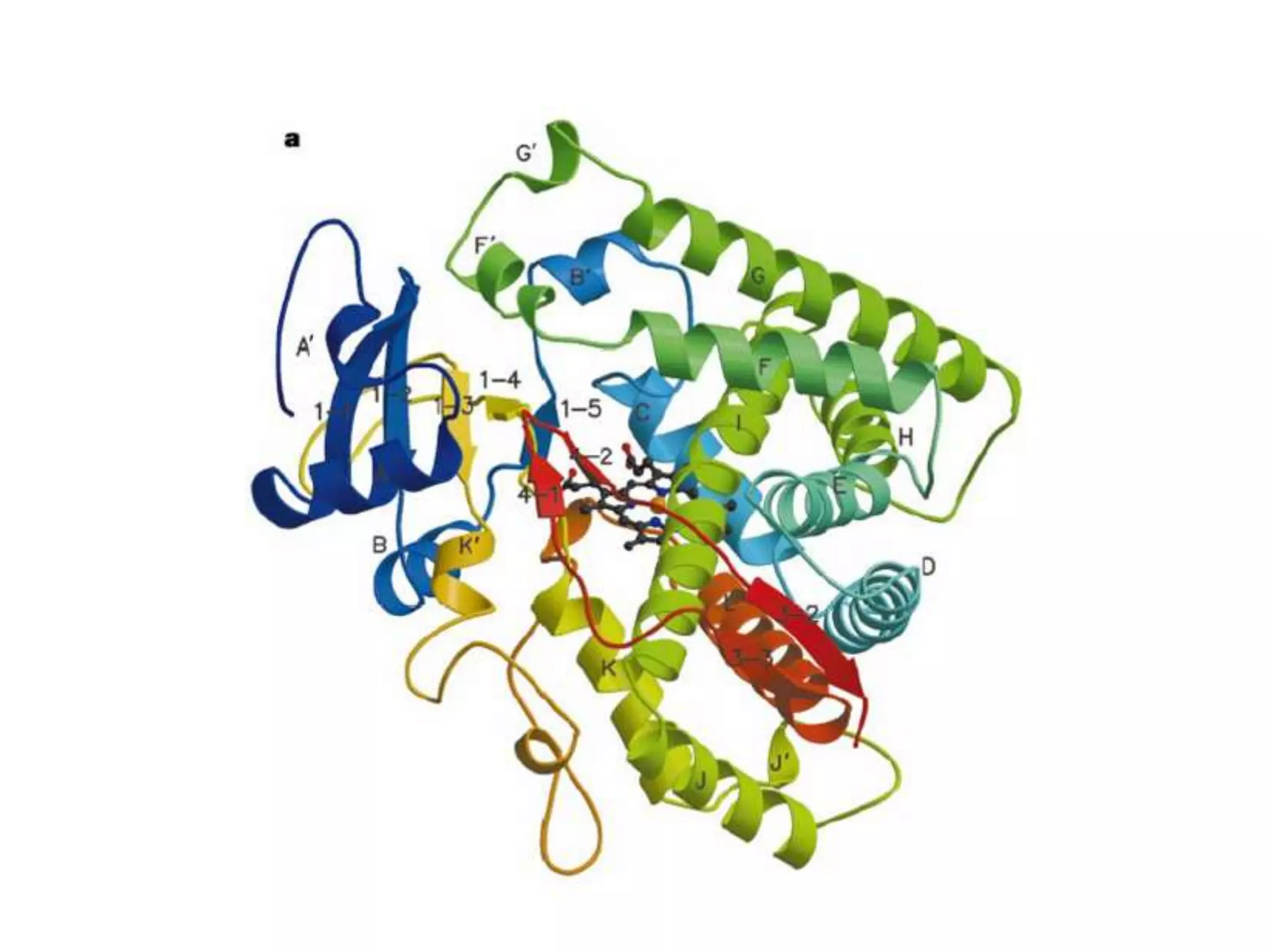
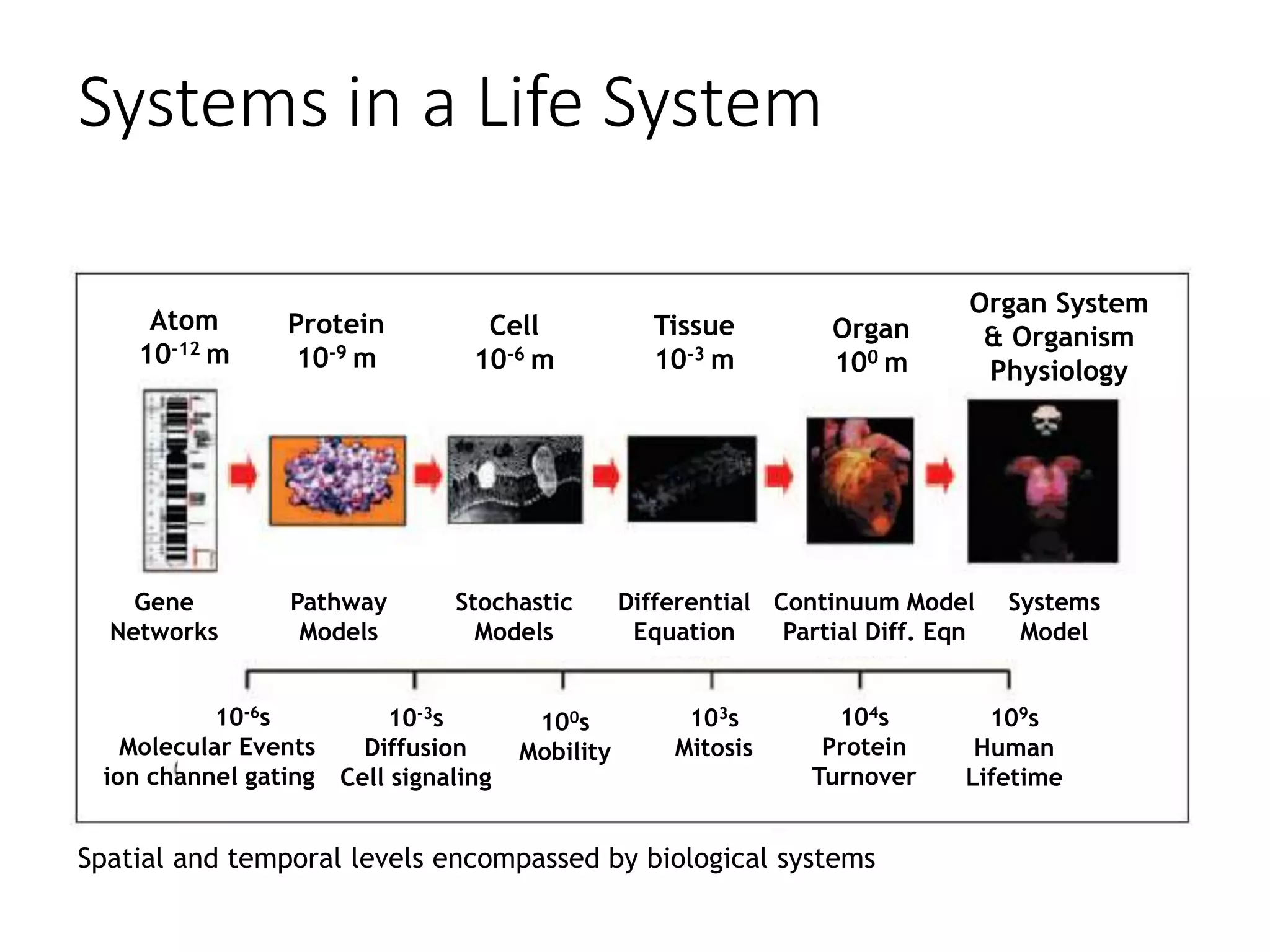
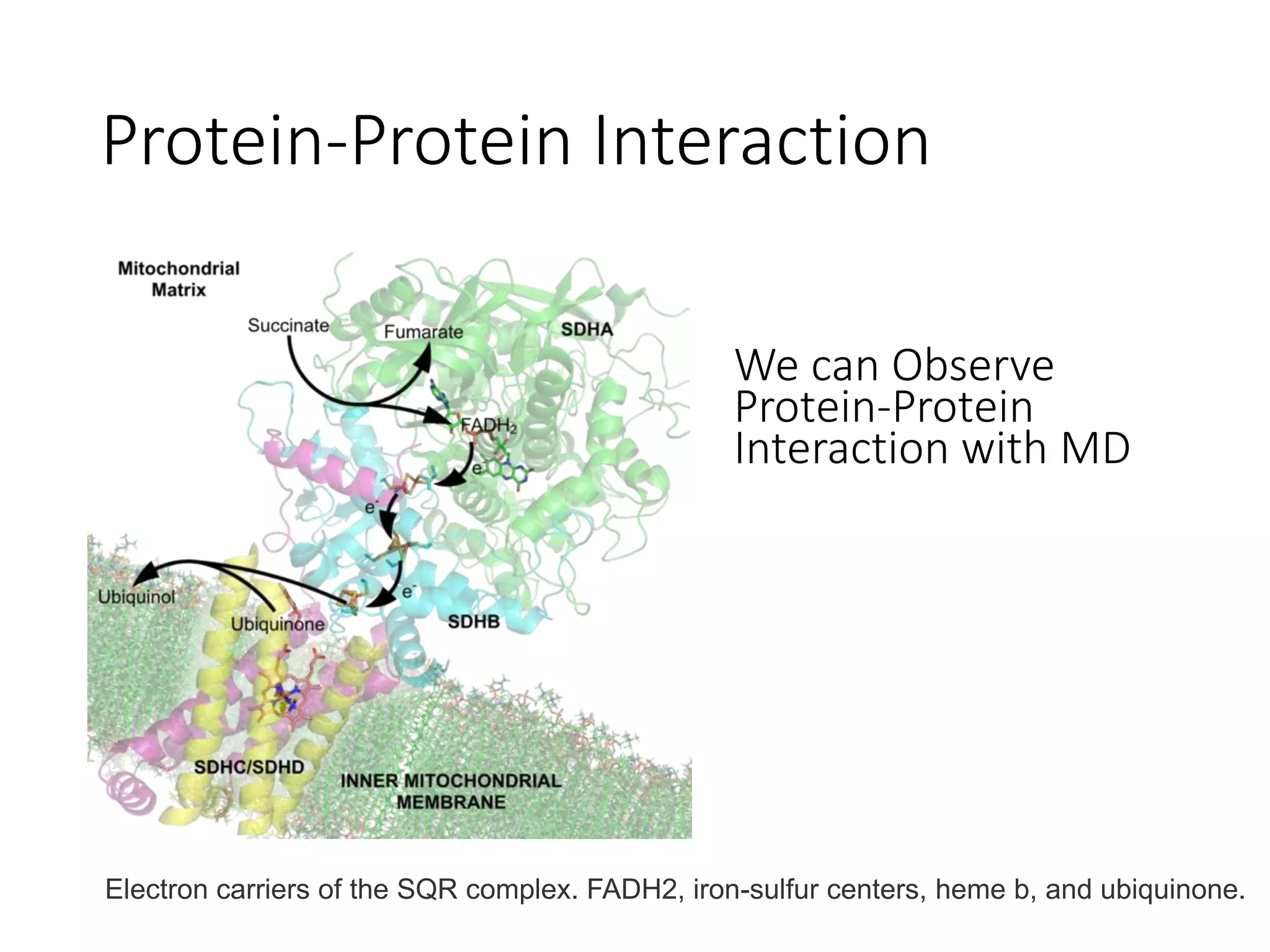
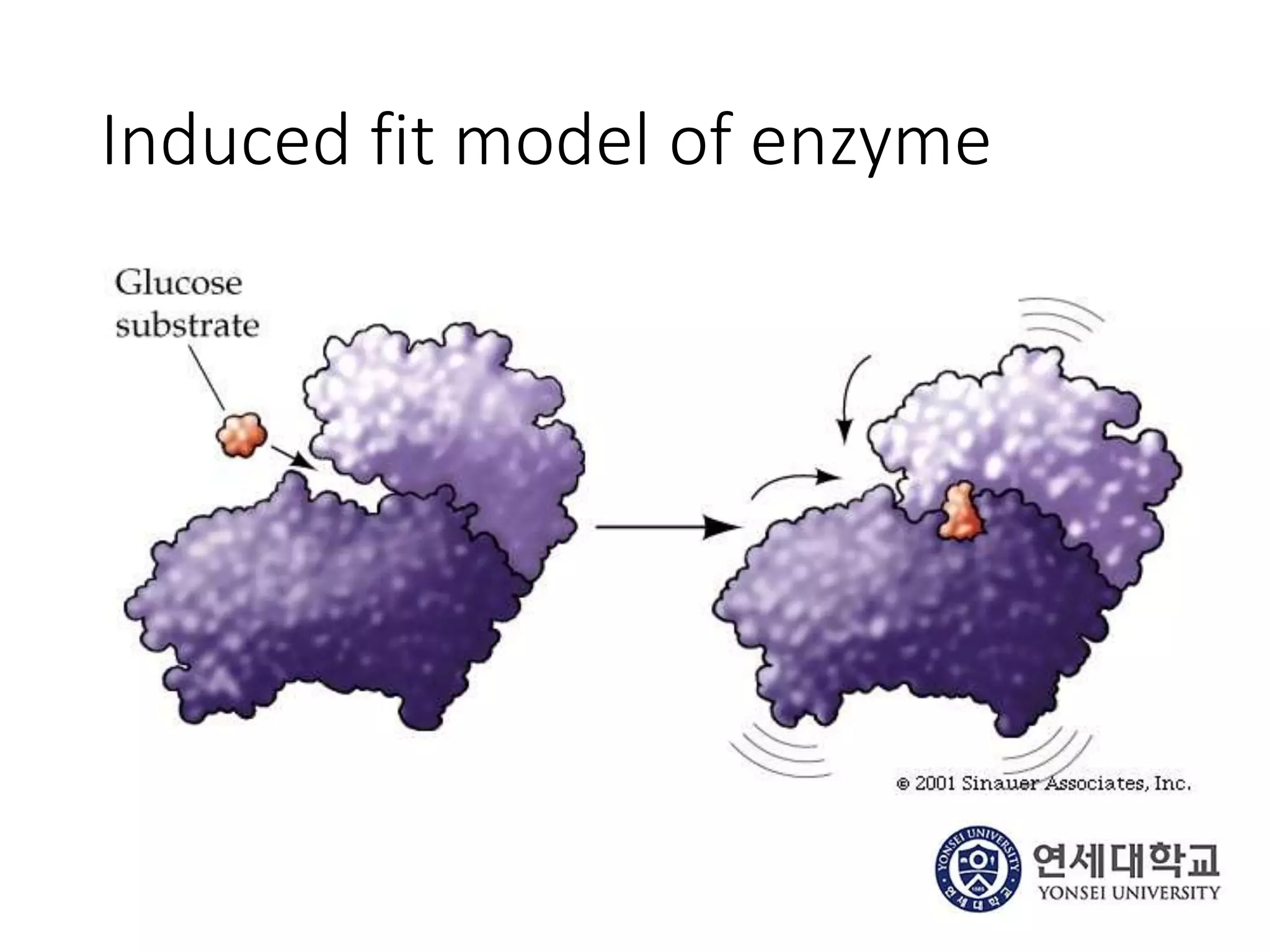
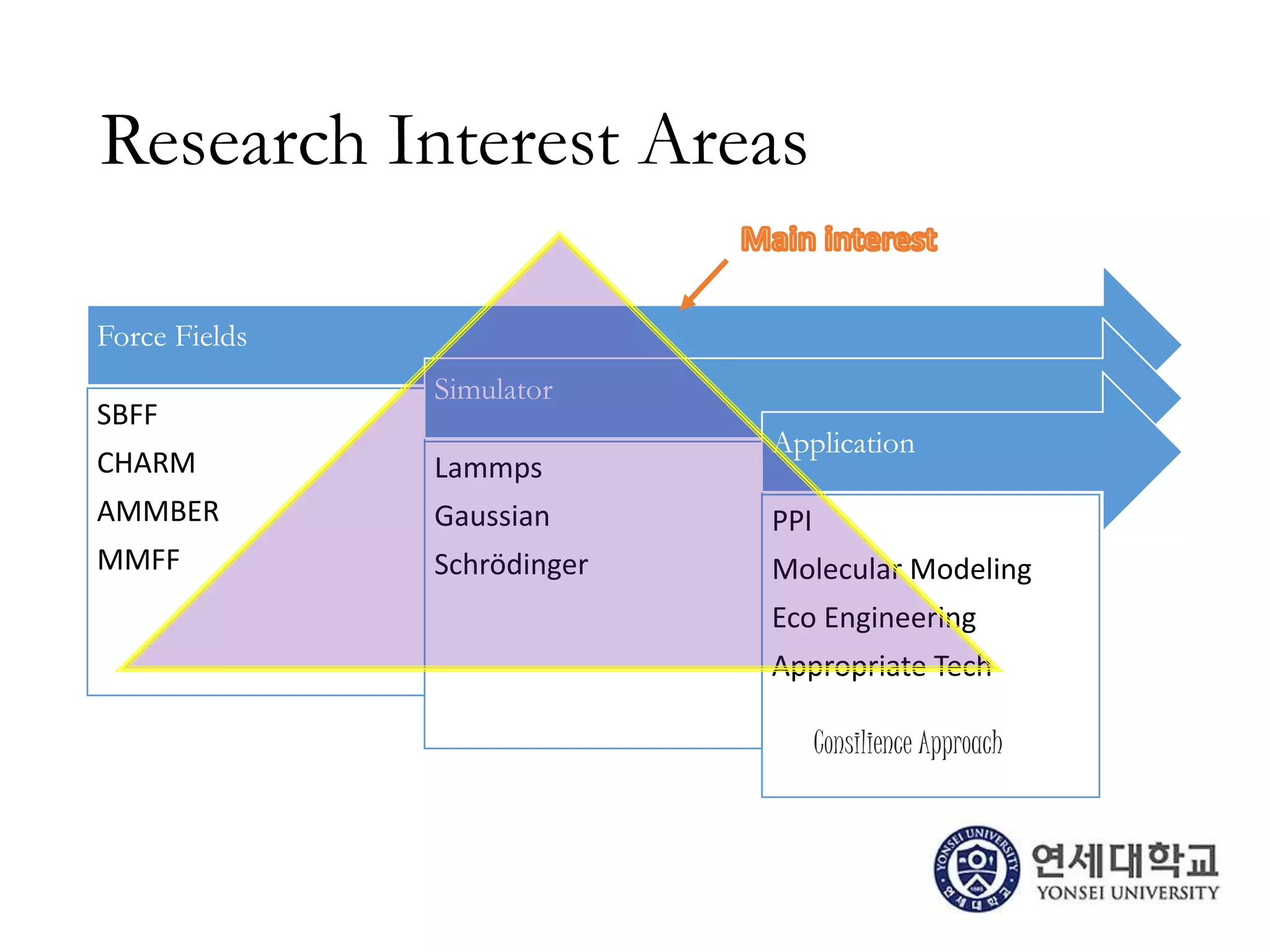
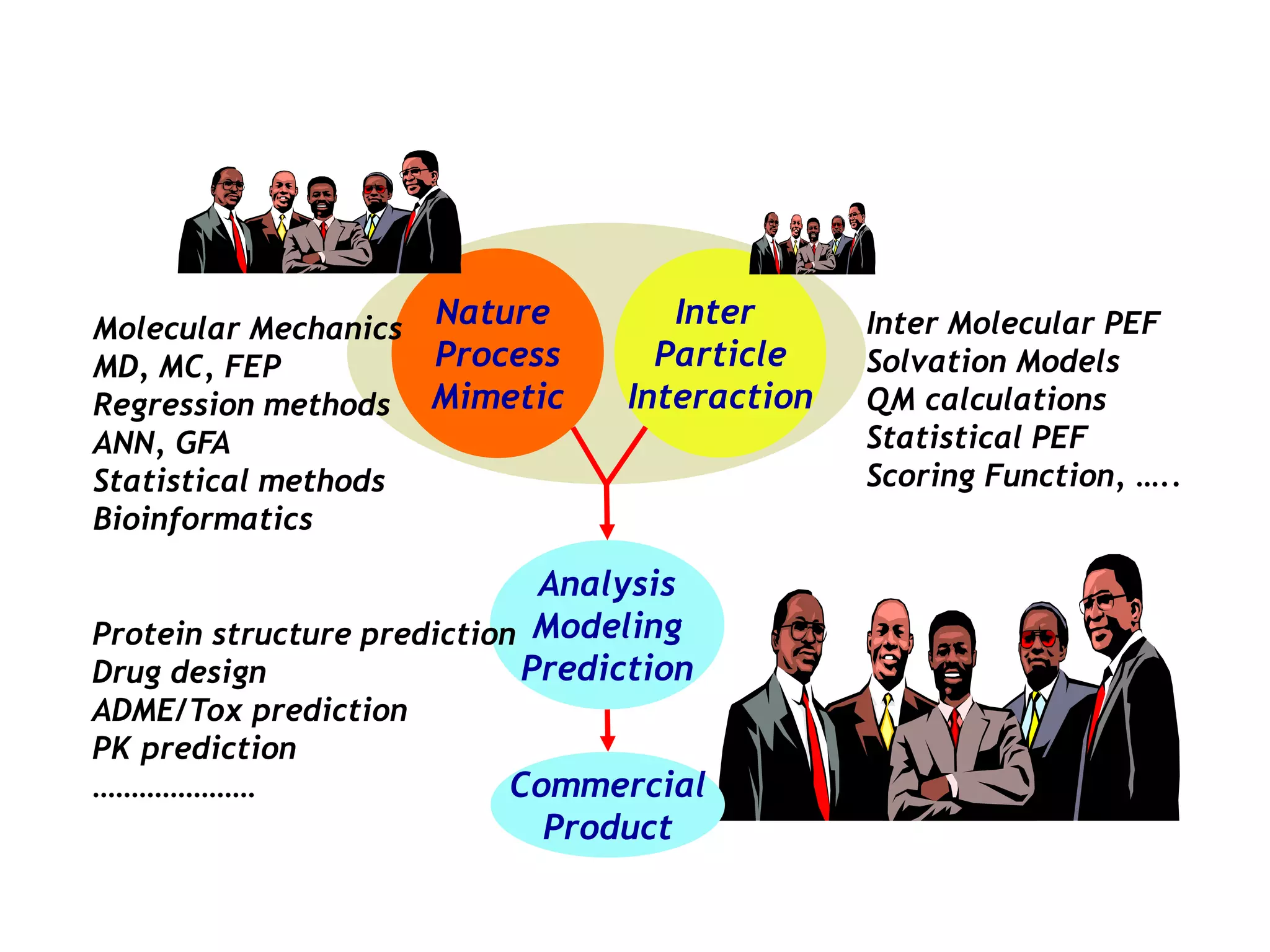
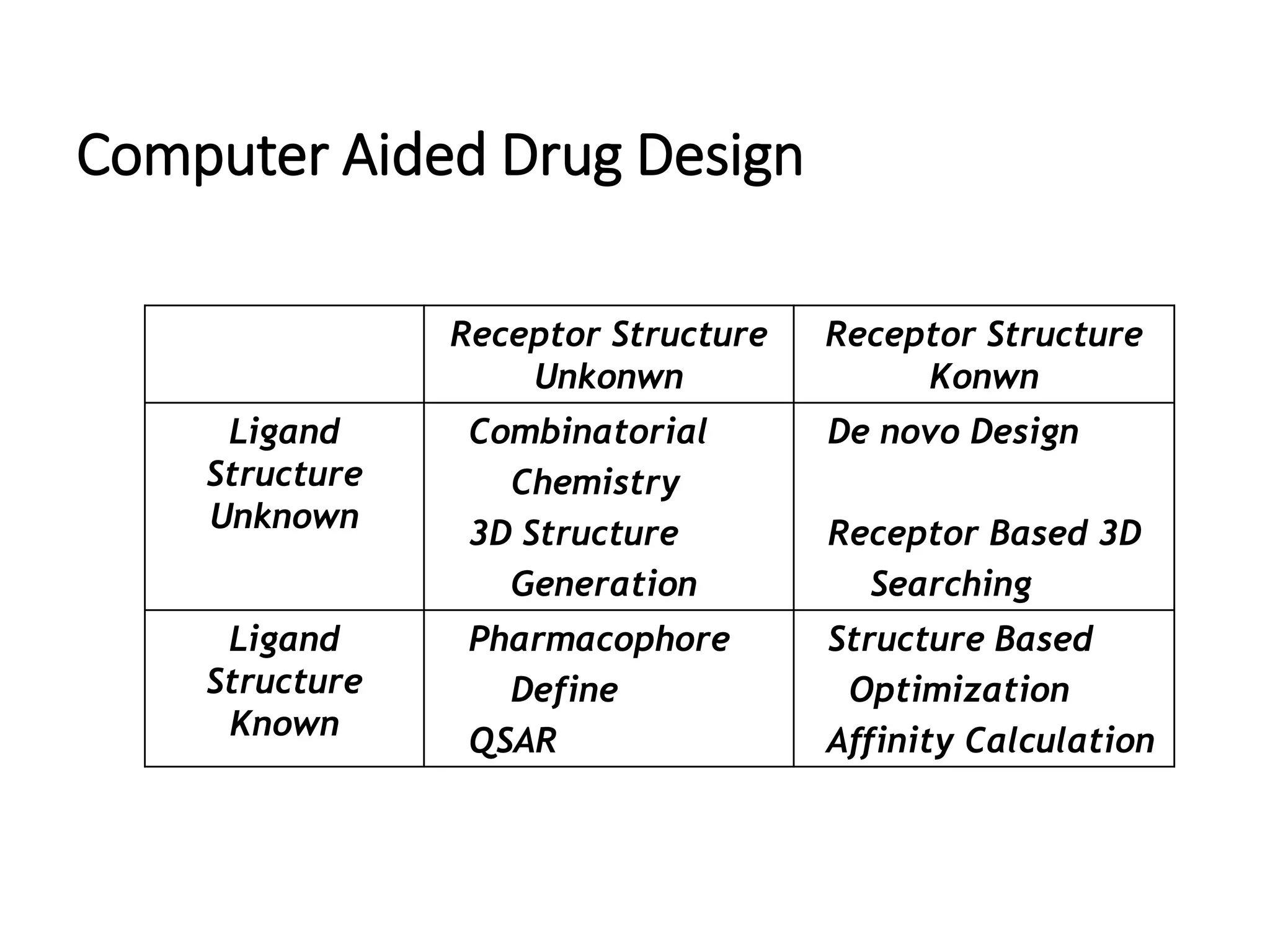
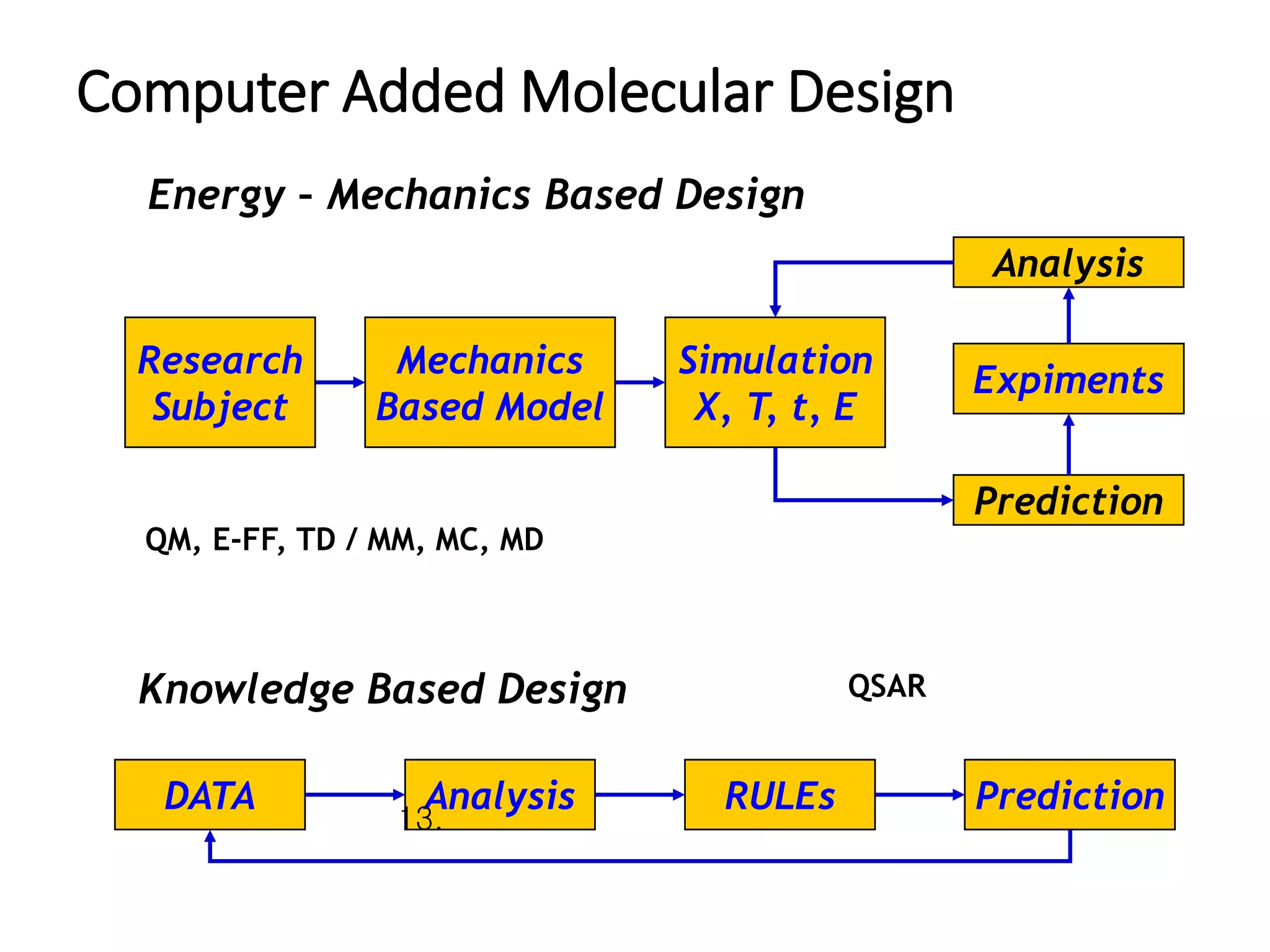
The document outlines the activities and research interests of the Bioinformatics & Molecular Design Research Center, focusing on molecular dynamics simulations, enzyme interactions, and drug design methodologies. It highlights computational techniques used for analyzing molecular behaviors, including various force fields and modeling systems, alongside the applications of high-performance computing in biological research. Additionally, it discusses specific case studies and advancements in molecular modeling and computer-aided drug development.