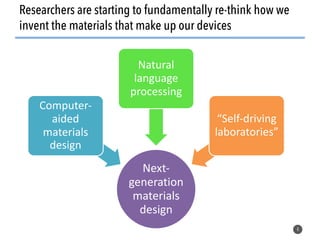
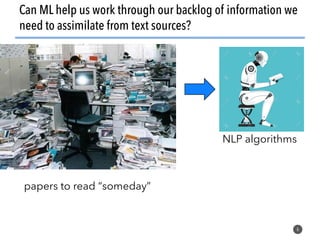
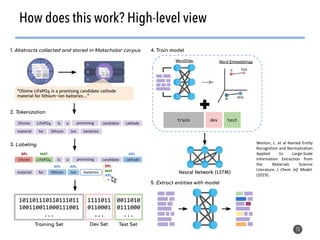
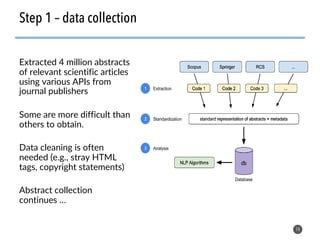
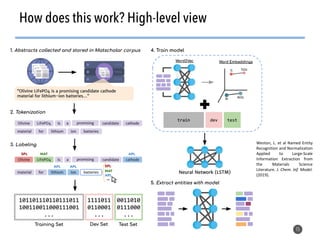
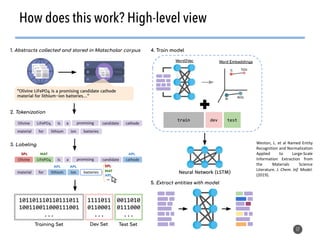
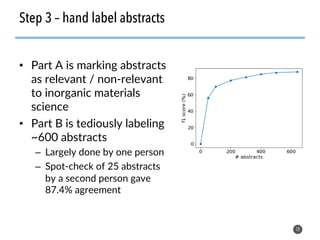
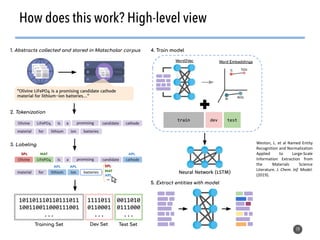
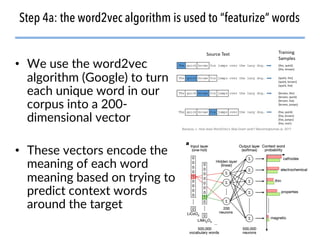
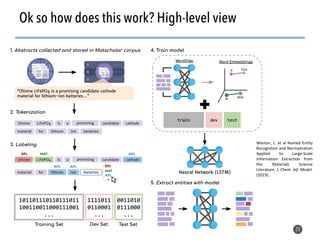
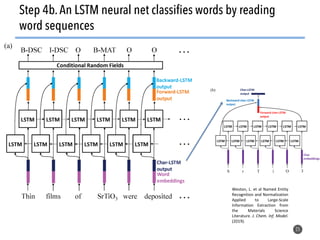
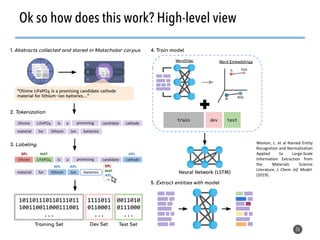
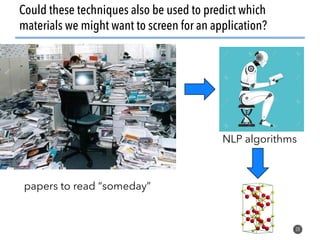
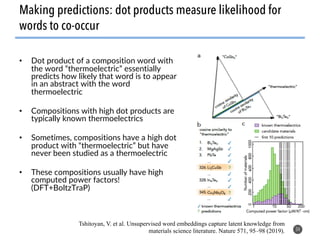
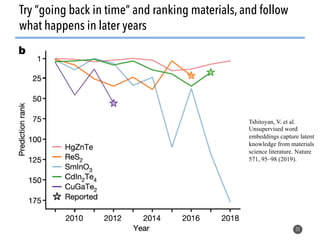
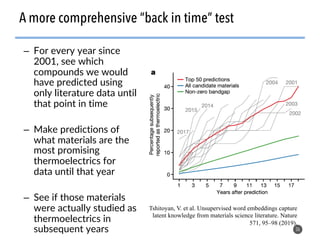
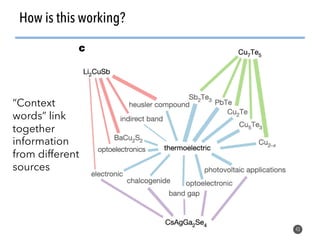
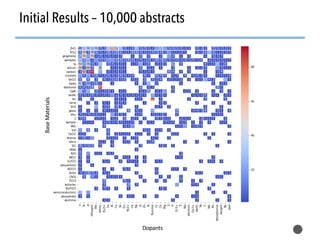
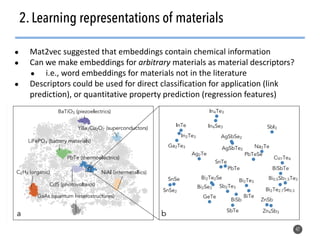
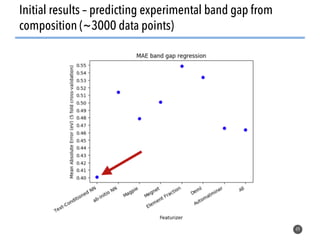
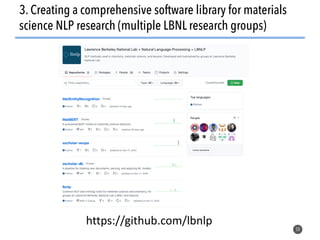
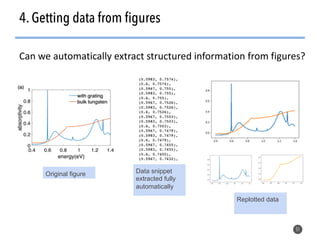
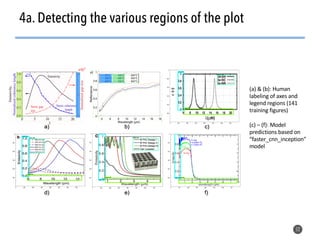
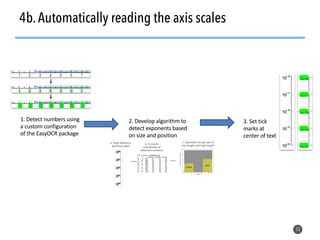
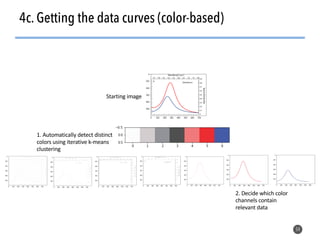
This document discusses natural language processing (NLP) techniques for materials design using information from millions of journal articles. It begins with an overview of how materials are typically discovered and optimized over decades before discussing how NLP could help address this challenge. The document then provides a high-level view of how NLP is used to extract and analyze information from millions of materials science abstracts, including data collection, tokenization, training machine learning models on labeled text, and using the models to automatically extract entities. Examples are given of how word embeddings can encode scientific concepts and relationships in ways that allow predicting promising new materials for applications like thermoelectrics. The talk concludes by discussing future directions for the NLP work.


![What constrains traditional approaches to materials design?
3
“[The Chevrel] discovery resulted from a lot of
unsuccessful experiments of Mg ions insertion
into well-known hosts for Li+ ions insertion, as
well as from the thorough literature analysis
concerning the possibility of divalent ions
intercalation into inorganic materials.”
-Aurbach group, on discovery of Chevrel cathode
for multivalent (e.g., Mg2+) batteries
Levi, Levi, Chasid, Aurbach
J. Electroceramics (2009)](https://image.slidesharecdn.com/jainimxtalk-210412051033/85/Materials-design-using-knowledge-from-millions-of-journal-articles-via-natural-language-processing-techniques-3-320.jpg)