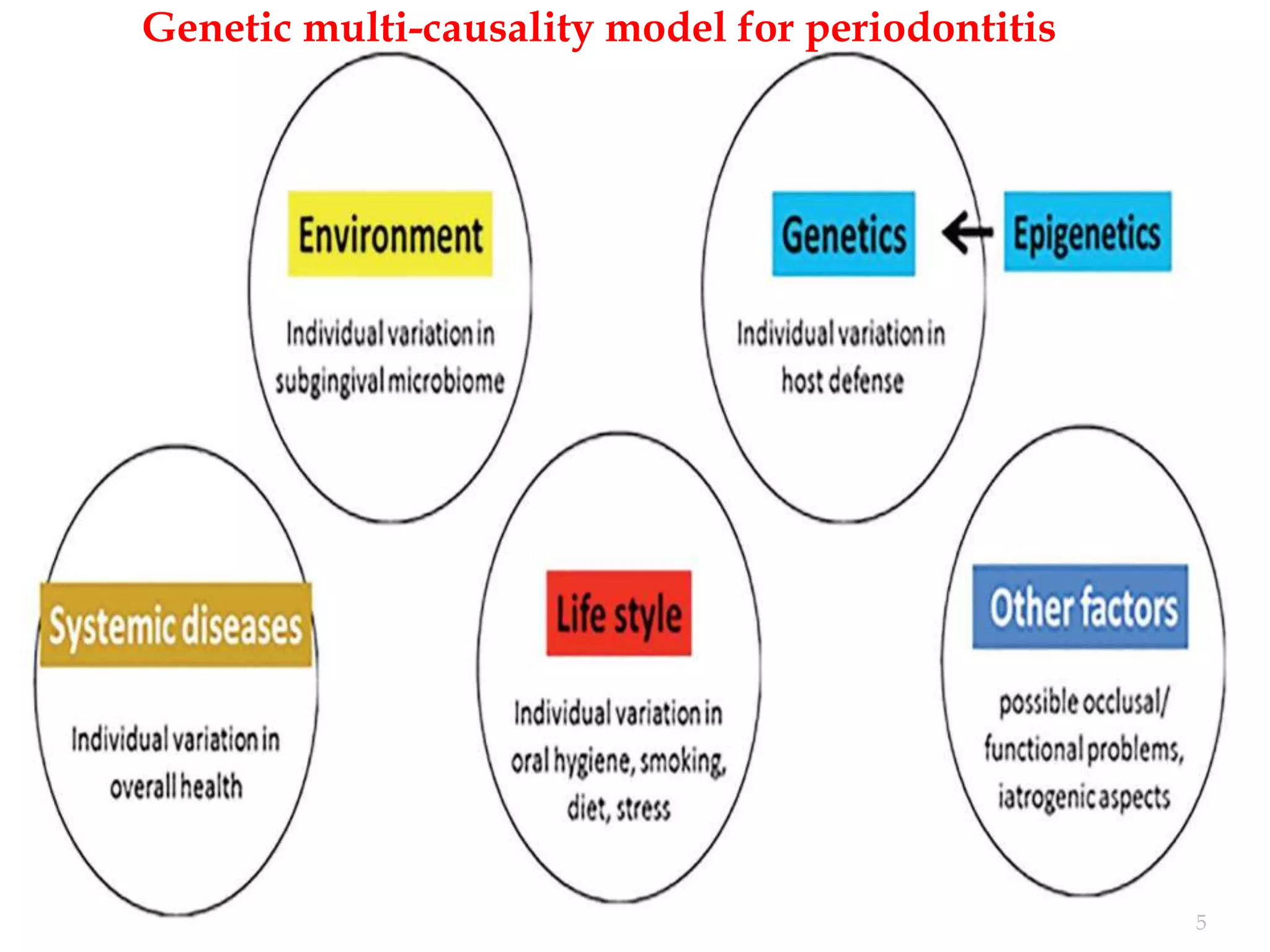
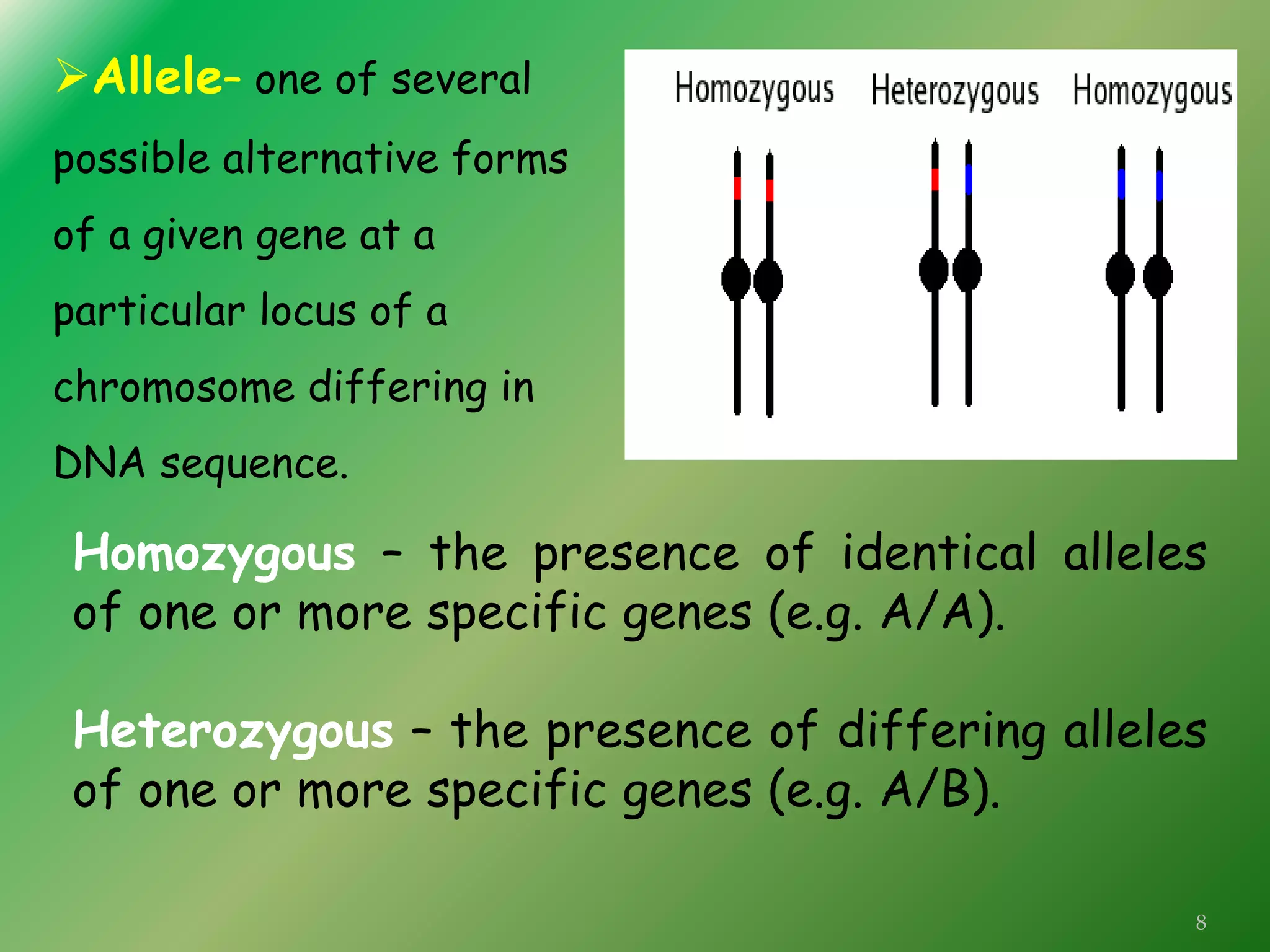
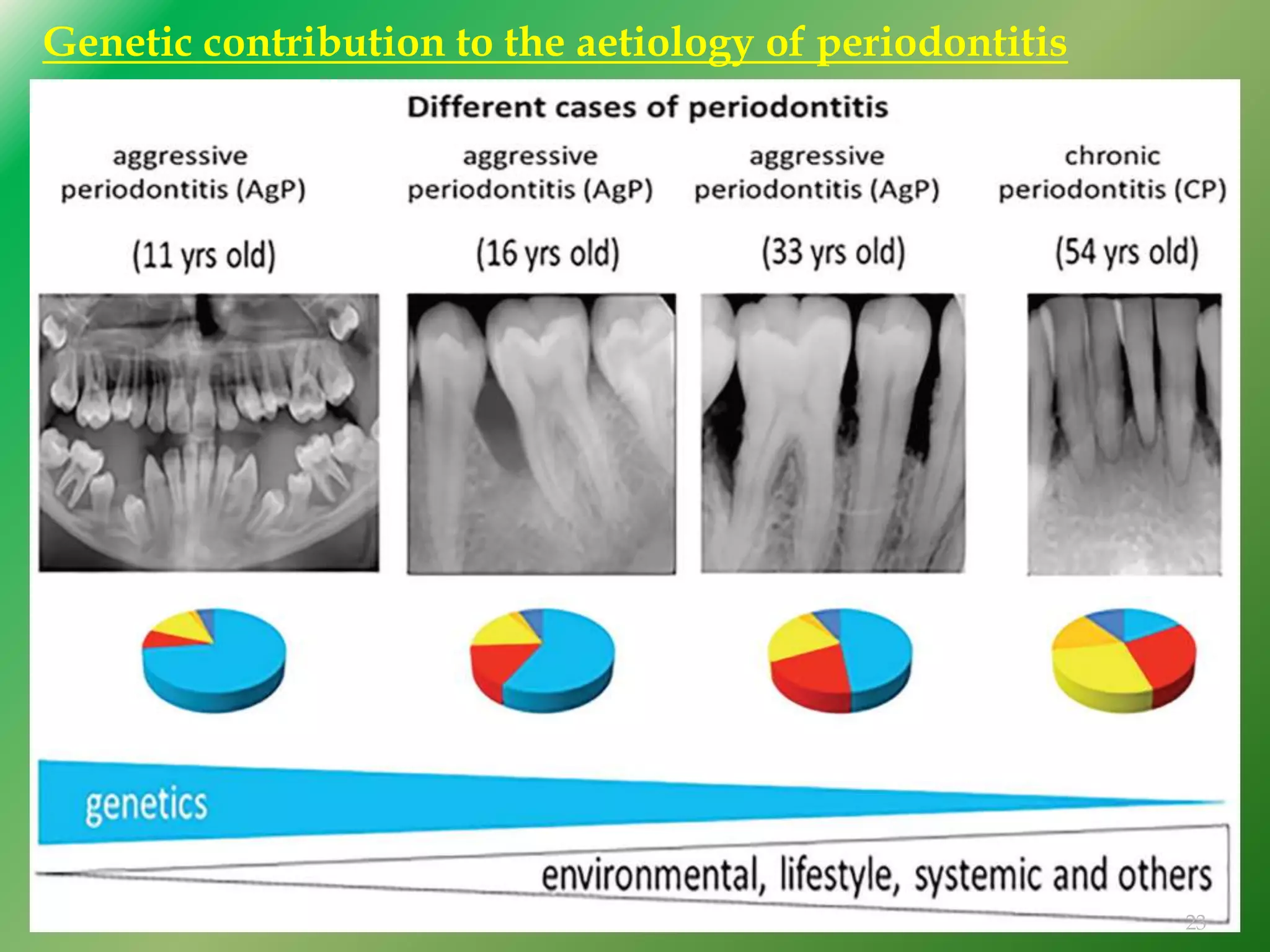
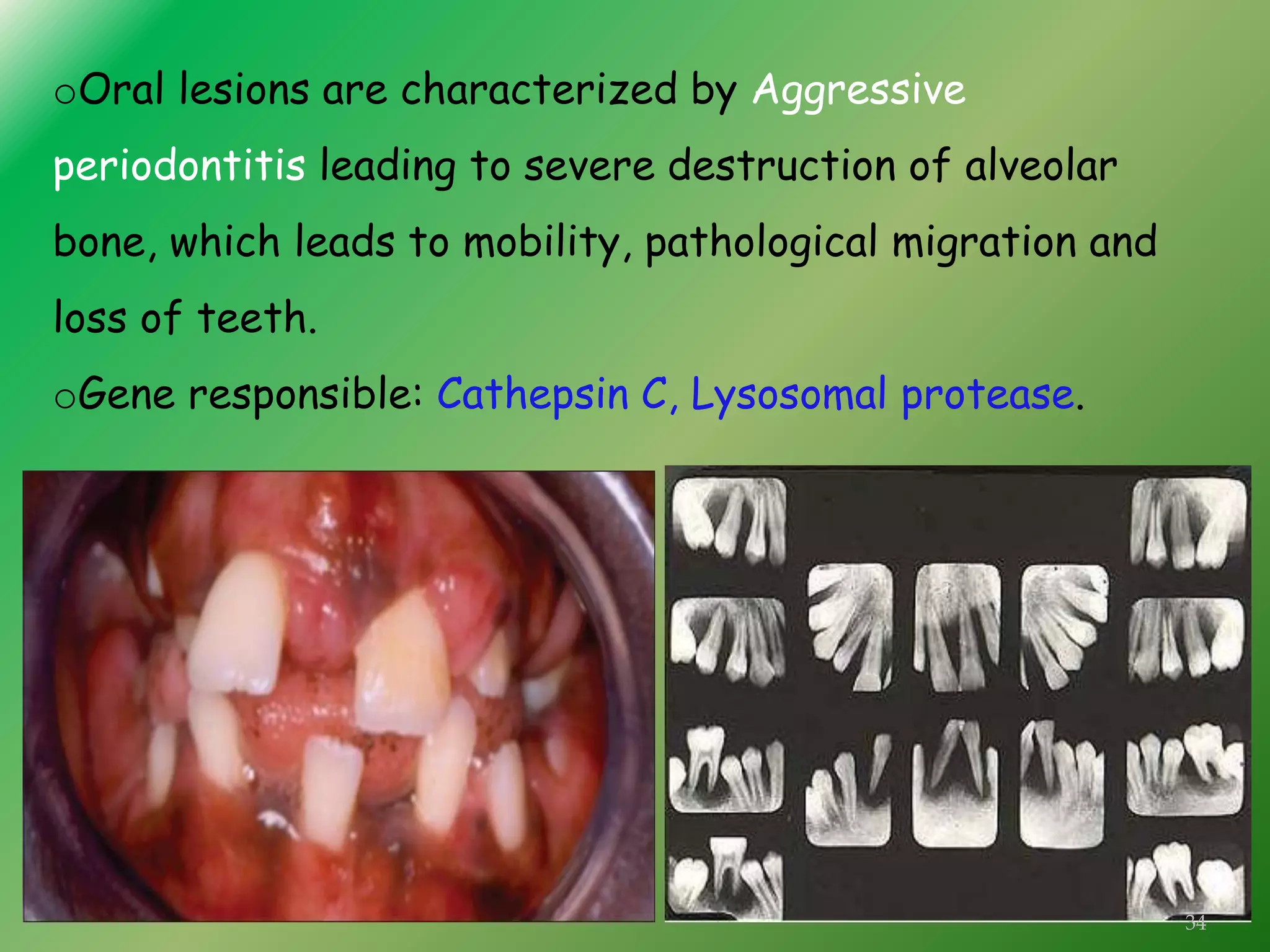
The document discusses the significant role of genetics in periodontitis, an inflammatory disease that is influenced by various genetic and environmental factors. It highlights the methods used in genetic studies, including segregation analysis, twin studies, linkage analysis, and genome-wide association studies, to understand the hereditary aspects of periodontitis. The findings emphasize the genetic predispositions associated with aggressive and chronic periodontitis, suggesting potential avenues for gene therapy and personalized treatment strategies.