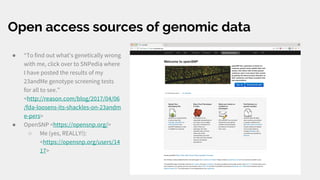
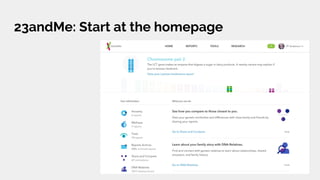
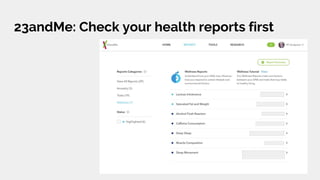
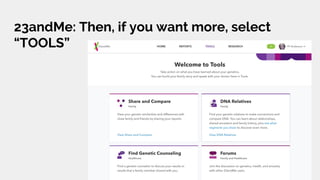
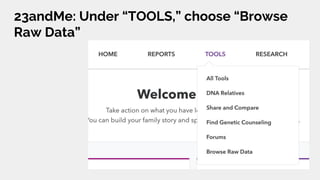
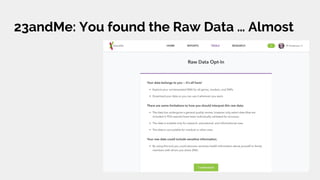
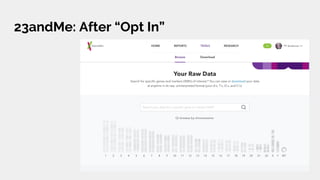
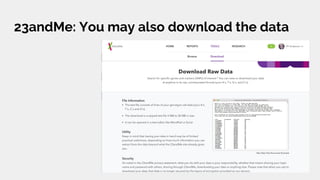
The document discusses the role of personal genomics in teaching genetic concepts and integrating genomic data into precision medicine, highlighting its accessibility and the resources available for educators. It emphasizes strategies for utilizing personal genomic data in a classroom setting, while also addressing the benefits, limitations, risks, and ethical considerations involved. Additionally, the document outlines various personal genomics services and tools that can facilitate self-tracking and data analysis.