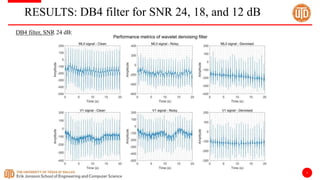
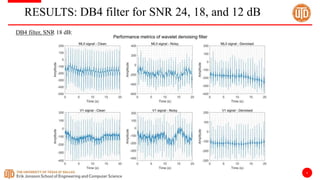
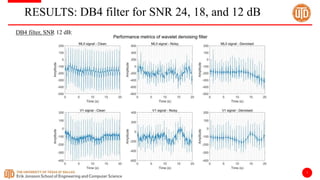
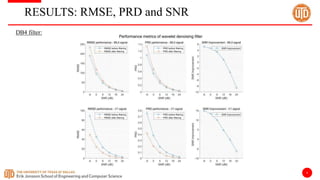
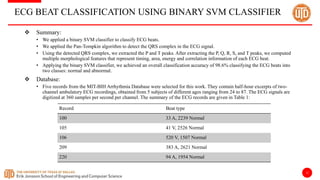
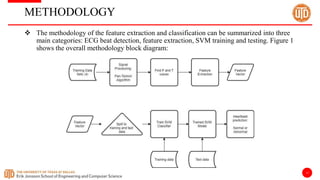
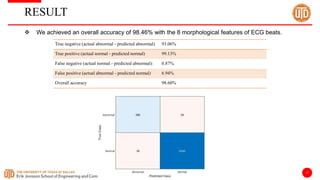
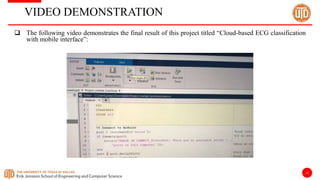
The document discusses a cloud-based ECG classification system that utilizes wavelet transform for ECG noise filtering and a binary SVM classifier for ECG beat classification, achieving a high accuracy of 98.6%. It details the methodology, data sources, and implementation involving an Android application for real-time ECG signal analysis. The application allows healthcare providers to access and analyze ECG signals from Bluetooth-enabled devices efficiently.