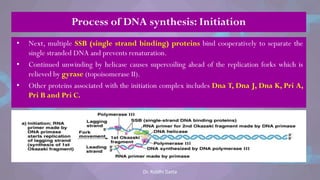
The document covers the process of DNA replication, detailing models such as conservative, semi-conservative, and dispersive, with a focus on the semi-conservative model supported by the Meselson-Stahl experiment. It discusses the role of DNA-dependent DNA polymerases, their functions, and the stages of DNA synthesis: initiation, elongation, and termination. Additionally, it highlights the various proteins involved in the replication process and their specific roles.























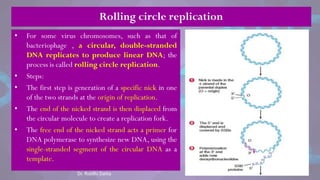
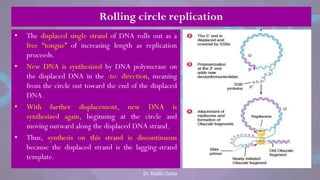
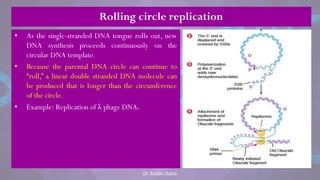


![• Each eukaryotic chromosome consists
of one linear DNA double helix.
• Eukaryotic chromosomes replicate efficiently
and quickly because DNA replication is
initiated at many origins of replication
throughout the genome.
• Eventually, each replication fork runs into an
adjacent replication fork, initiated at an
adjacent origin of replication.
• The stretch of DNA from the origin of
replication to the two termini of replication
(where adjacent replication forks fuse) on
each side of the origin is called a replicon
or replication unit.
• [The E.coli genome consists of one replicon.]
Replication of linear double stranded DNA in eukaryotes
Dr. Riddhi Datta](https://image.slidesharecdn.com/dnareplication-200204184727/85/Basics-of-DNA-Replication-47-320.jpg)