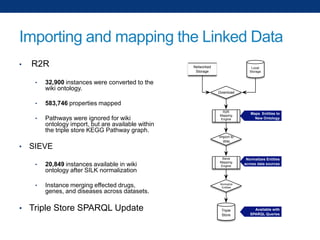
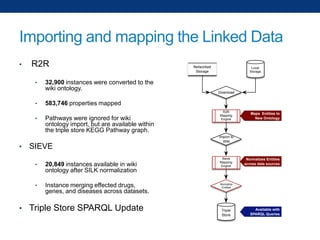
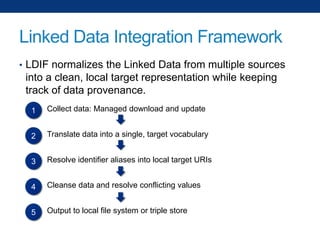
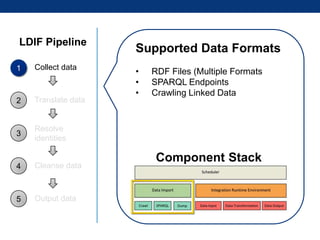
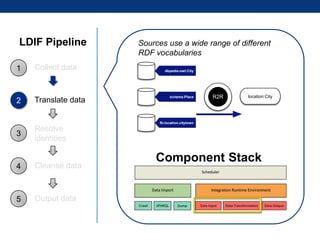
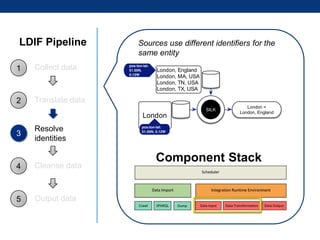
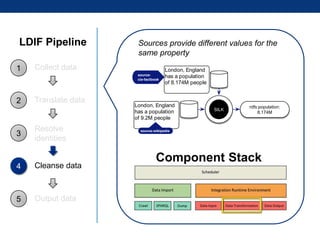
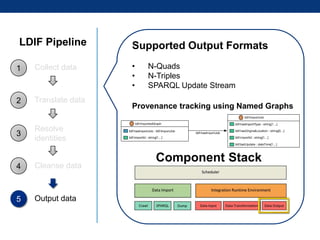
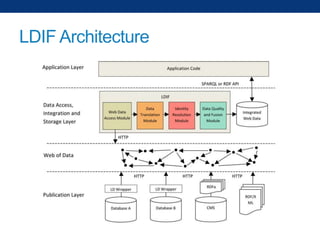
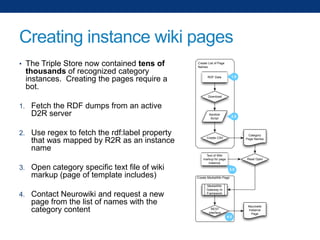
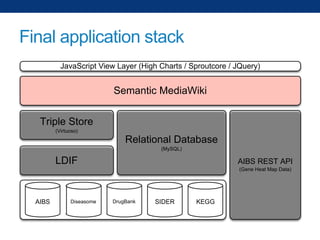
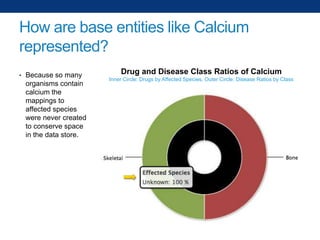
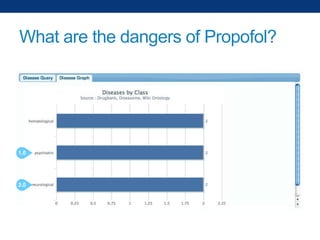
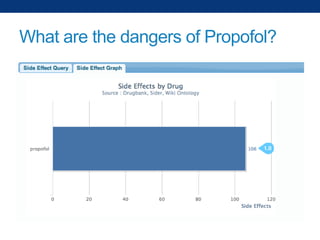
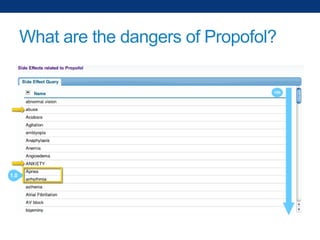
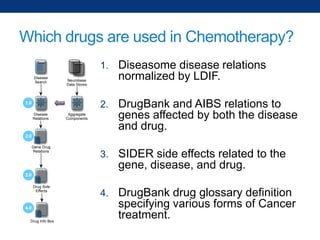
The document discusses the application of linked data and semantic technology to expand a neurobiology dataset, focusing on normalization and enrichment of gene data from various research sources. It highlights the Allen Institute for Brain Science's mission and introduces the Linked Data Integration Framework (LDIF) for managing diverse RDF datasets, addressing challenges such as conflicting identifiers and properties. The document also details the creation of a neurobiology authoring platform and the methodologies for integrating and visualizing this complex data.