Shahenaz seminar i
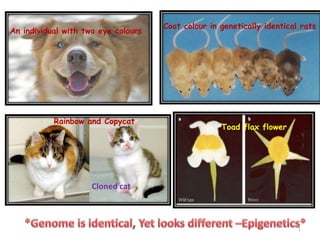
- 1. An individual with two eye colours Coat colour in genetically identical rats Cloned cat Rainbow and Copycat Toad flax flower 1
- 2. 2 University of Horticultural Sciences, Bagalkot Shahenaz begum Khaja hussain Sr MSc (UHS16PGM740) Dept. of Biotechnology and Crop Improvement
- 3. Basic concept in Epigenomics Epigenetic modifications Epigenomics of modifications Comparison of epigenetics with genetics Applications of epigenetics Research findings Conclusion
- 5. Term “EPIGENETICS” coined by – Waddington (1942) He is known as “Father of Epigenetics’’. 5
- 6. 6 Epigenetics literally means “above” Or “ on top of” genetics: It refers to external modification to DNA that turns genes ‘on’ or ‘off’. ‘’The study of changes in gene function that are heritable and donot entail a change in DNA sequence’’ - Corrent wu and morris(2001) Epigenomics : Study of the complete set of epigenetic modifications on the genetic material of a cell .
- 7. 7 DNA Histone protein Chromatin It is a multitude of chemical compounds that can tell the genome ‘What to do’ Epigenome
- 8. 8
- 9. 9 Methylation DNA methylation abundant in plants and vertebrates “Heritable epigenetic enzymatic modification resulting from the addition of a methyl group in the cyclic carbon-5 of cytosine occurs via enzyme called DNA methyl transferase.” . DNA methylation-DNMT Histone methylation-HMT = Methyl group DNA methyltransferase CYTOSINE 5’METHYLCYTIDINE
- 10. 10 DNA methylation usually inhibits transcription of eukaryotic genes particularly when it occurs in vicinity of promoter. CpG islands occur near the many promoters genes . These CpG islands are commonly 1000- 2000bp in length and contain high number of CpG sites.
- 11. 11 • Ex: DNMT 1Maintenance methylase • Ex: DNMT 3a, DNMT 3b Denovo methylase
- 13. 13
- 14. Enzymes Methylation sensitive enzyme Ex: HpaII EcoRI, Aat II, Acc II Methylation insensitive enzyme Ex : MspI( CCGG) Sphl(CGTAC/g) and Bbul(CGTAC/G) and Agarose gel banding pattern Unmethylated cytosine base Unmethylated and methylated cytosine base 14
- 15. 15
- 16. 16 Condensed and transcriptionally Inactive Chromatin Less Condensed (Due to acetylation of histone or other remodeling) making it transcriptionally active Switching off Switching on
- 17. 17
- 18. 18 Multimeric protein complexes have being identified that function as “cellular memory keys” that “lock” gene expression states and enable their inheritance over many cell mitotic or meiotic division cycles
- 20. 20 Modifications Role in transcription Modification site Acetylation Activation H3(K9,K14,K18,K56) H4(K5,K8,K12,K16) H2B(K6,K7,K16,K17) Strahl and allis(2000) Methylation Activation H3(K4me2,K4me3,K36me 3,K79me2) Strahl and allis(2000) Methylation Repression H3(K9me3,K27me3 and H4(K20me3) Balazs(2014) Phosphorylation Activation H3(S10) Strahl and allis(2000)
- 21. 21 1) Chromatinimmunoprecipitation(ChIP) i) Cross-linked ChIP(X ChIP) ii) Native ChIP(N ChIP) i)ChIP-Chip ii)ChIP-seq iii) ChIP-qPCR 2. Massspectrometry
- 22. 22 A non-coding RNA (ncRNA) is a functional RNA molecule that is transcribed from DNA but not translated into proteins. Epigenetic related ncRNAs include miRNA, siRNA. In general, ncRNAs function to regulate gene expression at the transcriptional and post-transcriptional level.
- 23. 23 Non coding RNA Infrastructural non coding RNA Regulatory non coding RNA non coding siRNA non coding miRNA
- 24. Single stranded 20- to 25 nucleotide RNA processed from a stem-loop region of a longer transcript by Dicer-like enzymes Bartel. , 2004 MicroRNA (miRNA) 24 siRNAs Double stranded 20-22 nucleotide short RNA molecules Produced from long double stranded RNA by Dicer Bartel. , 2004
- 25. 25 Properties Epigenetics Genetics Type of variation Usually does not change in DNA sequence Involves change in DNA sequence Origin of variation Random changes due to imperfection of DNA methylation pattern Usually random Unit of variation Activity of gene (Up or Down) DNA bases , sequences Heritability Varies 100%
- 26. 26
- 27. 27
- 29. Objective: To know the role of “Epigenetic regulation” for drought tolerance in Tomato by miRNA analysis in drought tolerant and drought sensitive tomato root and up ground tissues. 29
- 30. Total RNA isolation (miRNA isolation kit) Identification of tomato miRNA by deep sequencing Validation of miRNA expression by qRT PCR Library construction and sequencing Illumina HiSeq2000 Platform) Target predictiont (psRNATarget server), Plant Material – Drought Sensitive- L. esculentum (L.M.I-56), Drought Tolerant L. esculentum var. cerasiformae(PI187002) Functional annotation and Gene ontology Drought treatment-5% PEG-Sensitive and Tolerant Roots and Leaves-4 DNA Samples Control Condition –Sensitive and Tolerant Leaves and Roots -4 DNA Samples miRNA Processed 30
- 31. Figure1: Distribution of tomato miRNA across root and upground tissues in tolerant and sensitive genotypes 31
- 32. Fig 2:Tomato miRNA distribution in all samples of Drought sensitive and drought tolerant tissues 32
- 33. Fig3: Comparisons of conserved miRNA expressing changes after drought exposure33
- 34. Fig 4 :qRT-PCR Validation of randomly selected drought-responsive eight miRNA in tolerant root tissues 34
- 35. Figure 5 : Functional Annotation of GO biological process related with drought stress in tomato by AGRIGO 35
- 36. Drought responsive miRNA differentially expressed in drought tolerant and drought sensitive genotypes exhibit tissue specific expression pattern. Majority of these miRNAs were involved in fruit, shoot, seed and root devlopment and few miRNAs were involved in regulation of drought and development responsive genes like RP, HD-ZIP, MYB, NAC and PSII root and upground tissues. 36
- 37. Objectives: 1.High quality of Denovo assembly of Apple genome with the already available Apple genome 2.DNA methylation data analysis to know their Epigenetic marks which contribute in Apple fruit development and other agronomically important traits. 37
- 38. 38 GDDH13 Fruit Material and Methodology followed Haploid Plant -GDDH13-Leaves and Fruits Haploid Plant - GDDH 18-Fruits DNA Isolation- GDDH 13 (Leaves and Fruits) GDDH 18 (Fruits) @ 3 days before pollination and 9 DAP Sodium Bisulfite treatment (Bisulfite Kit) Whole genome bisulfite sequencing (To determine cytosine methylation status) Identification of Differentially methylated regions (DMR) Between Leaves and Fruits of GDDH13 Between Fruits of GDDH13 and GDDH18 @ -3DAP and 9DAP Functional annotation of DMR (Reference Annotated Apple genome and Arabidopsis)
- 39. Fig.6 DNA Methylation distribution on all 17 chromosomes DNA Methylation distriubtion and recombination rates on Chromosome 11 39
- 40. 40 Comparison of Methylation level across gene rich regions, Transposable elements and HODOR (Type of TE) *TSS-Transcriptional Start Site TES-Transcriptional End Site Fig.7..Methylation within the gene rich regions is less compared to TE Transposable element-HODOR showed maximum methylation (90% for CG, 65% for CHG and 3% for CHH)
- 41. 41 Comparison of global Methylation sites (CG, CHG and CHH) of Arabidopsis, Soybean and Apple genomes Fig.8 Higher methylation for CG (86%) sites than CHG (74%) and CHH (5.4%) sites Content of Methylation of Soybean and Apple is almost same and less in Arabidopsis
- 42. 42 Differentially Methylated Regions (DMR) between Leaves and Fruits of GDDH13 Fig .9 Total of 1,017 significant DMR identified between leaves and fruits Maximum methylation was for CHH (86%) with 875 DMR followed by CHG sites with approximately 100 DMR Increased methylation was seen for developing Fruits than Leaves. 294 DMR were present in the Promoter regions
- 43. 43 Functional analysis of DMR Table 1..11 DMR were mostly involved In fruit size regulation/fruit devlopment
- 44. 44 Parent Haploid Haploid Isogenic lines -3DAP---197 DMR observed between GDDH13 (Large Fruits) and GDDH18 (Small fruits) 47 DMR located in promoter region (fig 11) 9DAP- 148 DMR were identified between GDDH13 (Large Fruits) and GDDH18 (Small fruits) 53 were located in promoter region Majority of the DMR showing decreased methylation in small fruited genotype (GDDH18) and increased methylation in Larger fruits genotype(GDDH18) 22 gene rich- DMR played a role in fruit size regulation- SPL13, ACS8, Cytochrome P456 etc
- 45. Methylation plays an important role in Fruit size regulation Higher methylation occurs in CG sites than others Limited methylation occurs in the promoter regions Increased methylation occurs in larger fruits 45
- 46. Some Tomatoes do not ripen and remain green These tomatoes have a heritable cytosine hyper methylation on the CNR, NOR and RIN gene promoter, which inhibits the expression of these genes leading to no ripening. When the unripe tomato is injected with the enzymes that inhibit the methylase enzymes-Lead to ripening of the tomato indicating the strong regulation of epigenetics in Tomato ripening process 46 Objective: Role of DNA methylation in tomato ripening related genes such as RIN, NOR and CNR & their association with ripening/non-ripening.
- 47. 47 Figure. 12
- 48. 48
- 49. 49
- 50. Role of Epigenetic regulations in biological process in plants Phenylpropanoid synthesis N & S assimilation Light perception Photosynthesis Flowering time Sugar metabolism Cell elongation Cold responses Starch mobilization Hormone signaling 50
- 51. 51
- 52. 52
Editor's Notes
- 50