PCR and RT PCR.pptx
- 1. DNA AMPLIFICATION – PCR,RT-PCR PRESENTED BY:- Y.Rajkumar Sohanlal
- 2. DNA AMPLIFICATION The production of multiple copies of a DNA. OR Repeated copying of a piece of DNA known as DNA amplification
- 3. HISTORY OF PCR YEAR SCIENTIST WORK 1966 Thomas Bork Thermus aquaticus 1983 kary Mullis Concept of PCR 1985 Saiki Application of PCR 1985 Cetus Corporation Taq Polymerase
- 4. PCR PCR is a rapid and versatile in vitro method for amplifying defined target DNA sequence with the source of DNA . It is known as polymerase chain reaction.
- 5. Requirements
- 7. For selective amplification some prior known DNA sequence is required , called as primer . Rules for primer binding To avoid the chances of primer binding other than desired location certain rules must be followed – Primer length Not very short or long As short primer may bind to non -target sites and long primer will slow down pace . PRIMER
- 8. NATURE OF SEQUENCE Inverted repeats should be avoided . Melting temperature – Tm value for two primer used togather should not differ by > 5 ͦc ͦͦ
- 9. Steps of PCR
- 10. DENATURATION
- 11. PRIMER ANNEALING
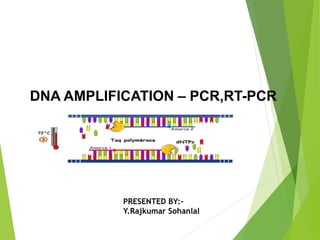
- 12. AMPLIFICATION
- 13. Taq polymerase used in DNA synthesis is obtained from microorganism Thermus aquaticus as it is stable upto 94⁰C Other than this Pfu (Pyrococcus furiosus), Bst E (Bacillus stearothermophilus) , Tth (Thermus thermophillus)
- 14. MODIFICATION IN PCR TECHNOLOGY PCR TYPE PROCEDURE Asymmetric PCR One strand is prefered over the other Reverse transcriptase PCR c-DNA is obtained from RNA then amplification is completed Hot start PCR it reduces non – specific amplification Inverse PCR used to amplify DNA with only one known sequence Anchored PCR Only one primer is used instead of two primer
- 16. INVERSE PCR
- 17. RT - PCR Real time PCR Based on general principle of PCR Also quantify target DNA molecule Depends upon detection and quantification of a fluorescent reporter . SYBR Green is used as fluorescent dye which binds in groove of ds DNA and moniters the total amount of DNA
- 20. PRINCIPLE OF RT - PCR
- 21. RT-PCR RT-PCR uses reverse transcriptase to convert m-RNA into double stranded DNA and then the gene without any introns can be amplified by regular PCR. RT-PCR has completed in two steps.
- 22. REAGENTS Magnesium chloride- 5-2.5 mm Buffer- pH 8.3-8.8 dNTP- 20-200µm Primers -0.1-0.5 µm Taq polymerase – 1- 2.5 µm Reverse transcriptase – 1-2.5 units Primary RNA - < 1µg
- 23. FIRST STEP Firstly reverse transcriptase recognises the 3’end of primers containing repeated thymines, this enzyme work at 70⁰C. It synthesizes a DNA strand that is complementary to the m-RNA. Then the RNA strand is replaced with another DNA strand leaving a double stranded DNA i.e. cDNA.
- 24. Second step Next the cDNA is amplified using a normal PCR reaction containing appropriate primers (one usually recognizes the poly(A) tail) Taq Polymerase and nucleotides. RT-PCR MACHINE (Thermocycler)
- 25. ADVANTAGES The advantages of this technique is evident when trying to use PCR to amplify a gene from eukaryotic DNA. Eukaryotes have introns which are present in genome but not in the mature m-RNA, some extremely long, which interrupt the coding segments. After transcription, the primary RNA transcript is processed to remove all the introns, hence becoming mRNA. When these genes are expressed in prokaryotic cells for protein production, the RNA produced directly from transcription need not undergo splicing as the transcript contains only exons.
- 28. REFERENCE Freeman WM, Walker SJ, (Jan. 1999).’Quantitative RT-PCR: pitfalls and potential” Bio Techniques. Mackay, Ian (2007). Real-time PCR in Microbiology : From Diagnosis to characterization. Norfolk, England: Caister Academic Press. Pp. Taylor S, Wakem M, Dijkman G, Alsarraj M, Nguyen M (April 2010). "A practical approach to RT-qPCR-Publishing data that conform to the MIQE guidelines". Methods. Bagasra, O. Protocols for the in situ PCR-amplification and detection of mRNA and DNA sequences. Nat Protoc 2, 2782–2795 (2007). https://doi.org/10.1038/nprot.2007.395