Chapter 16: Molecular Basis of Inheritance
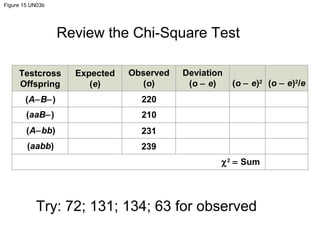
- 1. Figure 15.UN03b Testcross Offspring Expected (e) Observed (o) Deviation (o − e) (o − e)2 (o − e)2 /e (A−B−) (aaB−) (A−bb) (aabb) 220 210 231 239 χ2 = Sum Review the Chi-Square Test Try: 72; 131; 134; 63 for observed
- 3. Chapter 16 The Molecular Basis of Inheritance
- 4. In 1953, James Watson and Francis Crick introduced an elegant double-helical model for the structure of deoxyribonucleic acid, or DNA
- 5. The Search for the Genetic Material: Scientific Inquiry • When Morgan’s group showed that genes are located on chromosomes, the two components of chromosomes—DNA and protein—became candidates for the genetic material • The key factor in determining the genetic material was choosing appropriate experimental organisms • The role of DNA in heredity was first discovered by studying bacteria and the viruses that infect them
- 6. Evidence That DNA Can Transform Bacteria • The discovery of the genetic role of DNA began with research by Frederick Griffith in 1928 • Griffith worked with two strains of a bacterium, a pathogenic “S” strain and a harmless “R” strain • When he mixed heat-killed remains of the pathogenic strain with living cells of the harmless strain, some living cells became pathogenic • He called this phenomenon transformation, now defined as a change in genotype and phenotype due to assimilation of foreign DNA
- 7. Figure 16.2 Living S cells (pathogenic control) Experiment Results Living R cells (nonpathogenic control) Heat-killed S cells (nonpathogenic control) Mouse dies Mouse healthy Mouse healthy Mouse dies Mixture of heat- killed S cells and living R cells Living S cells
- 8. • In 1944, Oswald Avery, Maclyn McCarty, and Colin MacLeod announced that the transforming substance was DNA • Their conclusion was based on experimental evidence that only DNA worked in transforming harmless bacteria into pathogenic bacteria • Many biologists remained skeptical, mainly because little was known about DNA
- 9. Copyright ©The McGraw-Hill Companies, Inc. Permission required for reproduction or display 9-11 Avery et al also conducted the following experiments To further verify that DNA, and not a contaminant (RNA or protein), is the genetic material
- 10. 1. Which of the following results from Griffith’s experiment is an example of transformation? a. Mouse dies after being injected with living S cells. b. Mouse is healthy after being injected with living R cells. c. Mouse is healthy after being injected with heat-killed S cells. d. Mouse dies after being injected with a mixture of heat-killed S and living R cells. e. In blood samples from the mouse in “d”, living S cells were found.
- 11. 1. Which of the following results from Griffith’s experiment is an example of transformation? a. Mouse dies after being injected with living S cells. b. Mouse is healthy after being injected with living R cells. c. Mouse is healthy after being injected with heat-killed S cells. d. Mouse dies after being injected with a mixture of heat-killed S and living R cells. e. In blood samples from the mouse in “d”, living S cells were found.
- 12. • In 1952, Alfred Hershey and Martha Chase performed experiments showing that DNA is the genetic material of a phage known as T2 • To determine the source of genetic material in the phage, they designed an experiment showing that only one of the two components of T2 (DNA or protein) enters an E. coli cell during infection • They concluded that the injected DNA of the phage provides the genetic information
- 13. Figure 16.3 Phage head DNA Tail sheath Tail fiber Genetic material Bacterial cell 100nm Evidence That Viral DNA Can Program Cells
- 14. Figure 16.4 Experiment Batch 1: Radioactive sulfur (35 S) in phage protein Batch 2: Radioactive phosphorus (32 P) in phage DNA Labeled phages infect cells. Agitation frees outside phage parts from cells. Centrifuged cells form a pellet. Radioactivity (phage protein) found in liquid Pellet Centrifuge Radioactive protein Radioactive DNA Radioactivity (phage DNA) found in pellet Centrifuge Pellet 1 2 4 3 4
- 15. Additional Evidence That DNA Is the Genetic Material • In 1950, Erwin Chargaff reported that DNA composition varies from one species to the next A & T same ratio; G & C same ratio, but % GC varies from species to species (Chargaff’s rule) • This evidence of diversity made DNA a more credible candidate for the genetic material • By the 1950s, it was already known that DNA is a polymer of nucleotides, each consisting of a nitrogenous base, a sugar, and a phosphate group
- 16. Figure 16.5 5′ end Thymine (T) Adenine (A) Cytosine (C) Guanine (G) Nitrogenous bases Sugar– phosphate backbone 3′ end Nitrogenous base Sugar (deoxyribose)DNA nucleotide Phosphate
- 17. Maurice Wilkins and Rosalind Franklin were using a technique called X-ray crystallography to study molecular structure Franklin’s X-ray diffraction photograph of DNA Rosalind Franklin
- 18. • Franklin’s X-ray crystallographic images of DNA enabled Watson to deduce that DNA was helical • The X-ray images also enabled Watson to deduce the width of the helix and the spacing of the nitrogenous bases • The width suggested that the DNA molecule was made up of two strands, forming a double helix, anti-parallel in nature B-DNA 2 nm wide; 3.6 angstrom per unit; 10 units per
- 19. Figure 16.7 (a) Key features of DNA structure (b) Partial chemical structure 0.34 nm 3′ end 5′ endT T T A A A C C C G G G AT 1 nm TA C G CG AT 3.4 nm CG CG C G C G 3′ end 5′ end Hydrogen bond T A G C A T C G (c) Space-filling model
- 20. • Watson and Crick built models of a double helix to conform to the X-rays and chemistry of DNA • Franklin had concluded that there were two antiparallel sugar-phosphate backbones, with the nitrogenous bases paired in the molecule’s interior • At first, Watson and Crick thought the bases paired like with like (A with A, and so on), but such pairings did not result in a uniform width • Instead, pairing a purine with a pyrimidine resulted in a uniform width consistent with the X-ray
- 21. Figure 16.UN02 Purine + purine: too wide Pyrimidine + pyrimidine: too narrow Purine + pyrimidine: width consistent with X-ray data
- 22. Adenine (A) Thymine (T) Guanine (G) Cytosine (C) Sugar Sugar Sugar Sugar •Watson and Crick reasoned that the pairing was more specific, dictated by the base structures •They determined that adenine paired only with thymine, and guanine paired only with cytosine
- 23. The Basic Principle: Base Pairing to a Template Strand • Since the two strands of DNA are complementary, each strand acts as a template for building a new strand in replication • In DNA replication, the parent molecule unwinds, and two new daughter strands are built based on base-pairing rules •Watson and Crick noted that the specific base pairing suggested a possible copying mechanism for genetic material
- 24. Figure 16.9-3 (a) Parental molecule (b) Separation of parental strands into templates A A A T T T G G C C A A A T T T G G C C A A A T T T G G C C A A A T T T G G C C (c) Formation of new strands complementary to template strands
- 25. • Watson and Crick’s semiconservative model of replication predicts that when a double helix replicates, each daughter molecule will have one old strand (derived or “conserved” from the parent molecule) and one newly made strand • Competing models were the conservative model and the dispersive model • Meselson and Stahl provided the supporting scientific evidence
- 26. Figure 16.10 (a) Conservative model (b) Semiconserva- tive model (c) Dispersive model Parent cell First replication Second replication
- 27. • Experiments by Meselson and Stahl supported the semiconservative model • They labeled the nucleotides of the old strands with a heavy isotope of nitrogen, while any new nucleotides were labeled with a lighter isotope • The first replication produced a band of hybrid DNA, eliminating the conservative model • A second replication produced both light and hybrid DNA, eliminating the dispersive model and supporting the semiconservative model
- 28. Figure 16.11 Bacteria cultured in medium with 15 N (heavy isotope) Experiment Results Conclusion Bacteria transferred to medium with 14 N (lighter isotope) DNA sample centrifuged after first replication DNA sample centrifuged after second replication Less dense More dense Predictions: First replication Second replication Conservative model Semiconservative model Dispersive model 1 2 43
- 29. DNA Replication: A Closer Look • The copying of DNA is remarkable in its speed and accuracy • More than a dozen enzymes and other proteins participate in DNA replication Video: DNA Replication
- 30. 2. What is the %T in wheat DNA? a. approximately 22% b. approximately 23% c. approximately 28% d. approximately 45%
- 31. 2. What is the %T in wheat DNA? a. approximately 22% b. approximately 23% c. approximately 28% d. approximately 45%
- 32. Getting Started: Origins of Replication • Replication begins at special sites called origins of replication, where the two DNA strands are separated, opening up a replication “bubble” • A eukaryotic chromosome may have hundreds or even thousands of origins of replication • Replication proceeds in both directions from each origin, until the entire molecule is copied • At the end of each replication bubble is a replication fork, a Y-shaped region where new DNA strands are elongating
- 33. Figure 16.12 Origin of replication 0.5µm 0.25µm Bacterial chromosome Two daughter DNA molecules Replication bubble Parental (template) strand Daughter (new) strand Replication fork Double- stranded DNA molecule (a) Origin of replication in an E. coli cell (b) Origins of replication in a eukaryotic cell Origin of replication Eukaryotic chromosome Double-stranded DNA molecule Parental (template) strand Daughter (new) strand Replication fork Bubble Two daughter DNA molecules
- 34. Elongating a New DNA Strand • Enzymes called DNA polymerases catalyze the elongation of new DNA at a replication fork • Each nucleotide that is added to a growing DNA strand is a nucleoside triphosphate • The rate of elongation is about 500 nucleotides per second in bacteria and 50 per second in human cells
- 35. LE 16-13 New strand 5′ end Phosphate Base Sugar Template strand 3′ end 5′ end 3′ end 5′ end 3′ end 5′ end 3′ end Nucleoside triphosphate DNA polymerase Pyrophosphate
- 36. Antiparallel Elongation • The antiparallel structure of the double helix (two strands oriented in opposite directions) affects replication • DNA polymerases add nucleotides only to the free 3′ end of a growing strand; therefore, a new DNA strand can elongate only in the 5′ to 3′ direction
- 37. • Along one template strand of DNA, called the leading strand, DNA polymerase can synthesize a complementary strand continuously, moving toward the replication fork • To elongate the other new strand, called the lagging strand, DNA polymerase must work in the direction away from the replication fork • The lagging strand is synthesized as a series of segments called Okazaki fragments, which are joined together by DNA ligase
- 38. LE 16-14 Parental DNA 5′ 3′ Leading strand 3′ 5′ 3′ 5′ Okazaki fragments Lagging strand DNA pol III Template strand Leading strand Lagging strand DNA ligase Template strand Overall direction of replication
- 39. Priming DNA Synthesis • DNA polymerases cannot initiate synthesis of a polynucleotide; they can only add nucleotides to the 3′ end • The initial nucleotide strand is a short one called an RNA or DNA primer • An enzyme called primase can start an RNA chain from scratch • Only one primer is needed to synthesize the leading strand, but for the lagging strand each Okazaki fragment must be primed separately
- 40. Figure 16.16 Overview 1 2 1 1 2 1 2 1 1 2 5′ 3′ 3′ 5′ 5′ 3′ 3′ 3′ 3′ 5′ 5′ 5′ 5′ 3′ 3′ 5′ 3′ 5′ 3′ 5′ 3′ 5′ 3′ 5′ 3′ 5′ 3′ 1 2 5 6 4 3 Origin of replication Lagging strand Leading strand Lagging strand Leading strand Overall directions of replication RNA primer for fragment 2 Okazaki fragment 2 DNA pol III makes Okazaki fragment 2. DNA pol I replaces RNA with DNA. DNA ligase forms bonds between DNA fragments. Overall direction of replication Okazaki fragment 1 DNA pol III detaches. RNA primer for fragment 1 Template strand DNA pol III makes Okazaki fragment 1. Origin of replication Primase makes RNA primer. 5′
- 41. Other Proteins That Assist DNA Replication • Helicase untwists the double helix and separates the template DNA strands at the replication fork • Single-strand binding protein binds to and stabilizes single-stranded DNA until it can be used as a template • Topoisomerase corrects “overwinding” ahead of replication forks by breaking, swiveling, and rejoining DNA strands
- 42. • Primase synthesizes an RNA primer at the 5′ ends of the leading strand and the Okazaki fragments • DNA pol III continuously synthesizes the leading strand and elongates Okazaki fragments • DNA pol I removes primer from the 5′ ends of the leading strand and Okazaki fragments, replacing primer with DNA and adding to adjacent 3′ ends • DNA ligase joins the 3′ end of the DNA that replaces the primer to the rest of the leading strand and also joins the lagging strand fragments
- 43. Figure 16.17 Overview 5′ 3′ Lagging strand Leading strand Leading strand Lagging strand Leading strand Leading strand template Origin of replication Overall directions of replication 5′ 3′ 5′ 3′ 5′ 5′ 3′ 3′ 3′ Single-strand binding proteins Helicase Parental DNA DNA pol III Primer Primase Lagging strand Lagging strand template DNA pol III DNA pol I 5′ DNA ligase 123 4 5
- 45. The DNA Replication Machine as a Stationary Complex • The proteins that participate in DNA replication form a large complex, a DNA replication “machine” Called the Replisome • The DNA replication machine is probably stationary during the replication process • Recent studies support a model in which DNA polymerase molecules “reel in” parental DNA and “extrude” newly made daughter DNA molecules
- 46. Table 16.1
- 47. Proofreading and Repairing DNA • DNA polymerases proofread newly made DNA, replacing any incorrect nucleotides • In mismatch repair of DNA, repair enzymes correct errors in base pairing • In nucleotide excision repair, enzymes cut out and replace damaged stretches of DNA
- 48. LE 16-17 DNA ligase DNA polymerase DNA ligase seals the free end of the new DNA to the old DNA, making the strand complete. Repair synthesis by a DNA polymerase fills in the missing nucleotides. A nuclease enzyme cuts the damaged DNA strand at two points and the damaged section is removed. Nuclease A thymine dimer distorts the DNA molecule.
- 49. Replicating the Ends of DNA Molecules • Limitations of DNA polymerase create problems for the linear DNA of eukaryotic chromosomes • The usual replication machinery provides no way to complete the 5′ ends, so repeated rounds of replication produce shorter DNA molecules
- 50. Figure 16.20 Ends of parental DNA strands Lagging strand Parental strand RNA primer Last fragment Next-to-last fragment Lagging strand Leading strand Removal of primers and replacement with DNA where a 3′ end is available 3′ 5′ 5′ Second round of replication Further rounds of replication New lagging strand New leading strand Shorter and shorter daughter molecules 5′ 3′ 3′ 5′ 3′ 5′ 3′
- 51. • Eukaryotic chromosomal DNA molecules have at their ends nucleotide sequences called telomeres • Telomeres do not prevent the shortening of DNA molecules, but they do postpone the erosion of genes near the ends of DNA molecules • It has been proposed that the shortening of telomeres is connected to aging
- 53. • If chromosomes of germ cells became shorter in every cell cycle, essential genes would eventually be missing from the gametes they produce • An enzyme called telomerase catalyzes the lengthening of telomeres in germ cells using an enzyme with an RNA template
- 54. Concept 16.3: A chromosome consists of a DNA molecule packed together with proteins • The bacterial chromosome is a double- stranded, circular DNA molecule associated with a small amount of protein • Eukaryotic chromosomes have linear DNA molecules associated with a large amount of protein • In a bacterium, the DNA is “supercoiled” and found in a region of the cell called the nucleoid
- 55. • In the eukaryotic cell, DNA is precisely combined with proteins in a complex called chromatin • Chromosomes fit into the nucleus through an elaborate, multilevel system of packing
- 56. Figure 16.22 DNA, the double helix Histones Nucleosomes, or “beads on a string” (10-nm fiber) 30-nm fiber Looped domains (300-nm fiber) Metaphase chromosome DNA double helix (2 nm in diameter) Nucleosome (10 nm in diameter) 30-nm fiber Loops Scaffold Histones Histone tail H1 300-nm fiber Chromatid (700 nm) Replicated chromosome (1,400 nm)
- 58. • Most chromatin is loosely packed in the nucleus during interphase and condenses prior to mitosis • Loosely packed chromatin is called euchromatin • During interphase a few regions of chromatin (centromeres and telomeres) are highly condensed into heterochromatin • Dense packing of the heterochromatin makes it difficult for the cell to express genetic information coded in these regions
- 59. 3. Which of the following statements is false when comparing prokaryotes with eukaryotes? A) The prokaryotic chromosome is circular, whereas eukaryotic chromosomes are linea B) Prokaryotic chromosomes have a single origin of replication, whereas eukaryotic chromosomes have many. C) The rate of elongation during DNA replication is higher in prokaryotes than in eukaryotes. D) Prokaryotes produce Okazaki fragments during DNA replication, but eukaryotes do not. E) Eukaryotes have telomeres, and prokaryotes do not.
- 60. 3. Which of the following statements is false when comparing prokaryotes with eukaryotes? A) The prokaryotic chromosome is circular, whereas eukaryotic chromosomes are linea B) Prokaryotic chromosomes have a single origin of replication, whereas eukaryotic chromosomes have many. C) The rate of elongation during DNA replication is higher in prokaryotes than in eukaryotes. D) Prokaryotes produce Okazaki fragments during DNA replication, but eukaryotes do not. E) Eukaryotes have telomeres, and prokaryotes do not.
Editor's Notes
- Figure 15.UN03b Skills exercise: using the chi-square test (part 2)
- Figure 16.2 Inquiry: Can a genetic trait be transferred between different bacterial strains?
- Answer: D
- Answer: D
- Figure 16.3 A virus infecting a bacterial cell
- Figure 16.4 Inquiry: Is protein or DNA the genetic material of phage T2?
- Figure 16.5 The structure of a DNA strand
- Figure 16.7 The structure of the double helix
- Figure 16.UN02 In-text figure, purines and pyrimidines, p. 318
- Figure 16.9-3 A model for DNA replication: the basic concept (step 3)
- Figure 16.10 Three alternative models of DNA replication
- Figure 16.11 Inquiry: Does DNA replication follow the conservative, semiconservative, or dispersive model?
- Answer: C
- Answer: C
- Figure 16.12 Origins of replication in E. coli and eukaryotes
- Figure 16.16 Synthesis of the lagging strand
- Figure 16.17 A summary of bacterial DNA replication
- Table 16.1 Bacterial DNA replication proteins and their functions
- Figure 16.20 Shortening of the ends of linear DNA molecules
- Figure 16.21 Telomeres
- Figure 16.22 Exploring chromatin packing in a eukaryotic chromosome
- Figure 16.23 “Painting” chromosomes