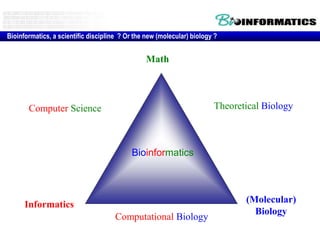
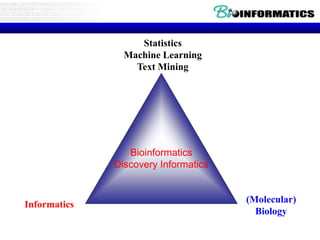
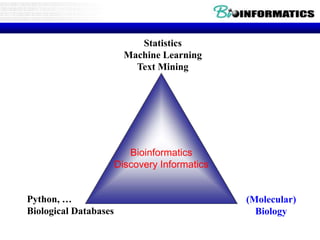
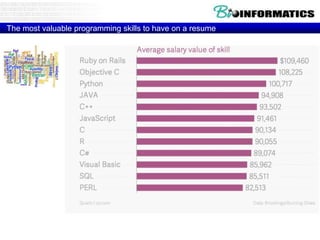
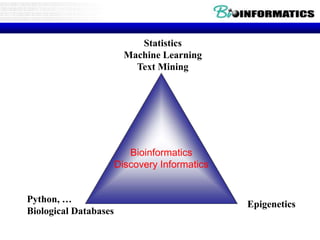
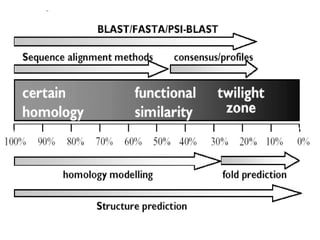
This document discusses biological databases and bioinformatics. It begins with an overview of bioinformatics as an interdisciplinary field combining biology, computer science, and information technology. It then discusses different types of biological databases, including those focused on sequences, pathways, protein structures, and gene expression. The document outlines some common uses of biological databases, including searching for annotations, identifying similar sequences through homology, searching for patterns, and making predictions. It also briefly discusses comparing data across databases. The summary provides a high-level overview of the key topics and uses of biological databases covered in the document.
























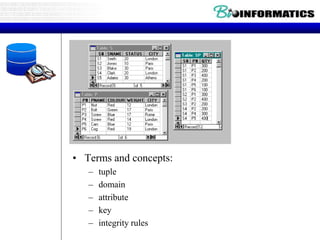



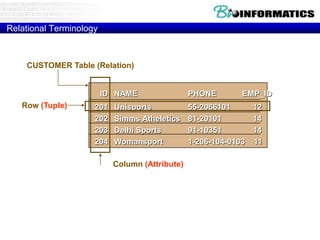

![Relational Database Terminology
Relational operators
• Relational
– select
rel WHERE boolean-xpr
– project
rel [ attr-specs ]
– join
rel JOIN rel
– divide by
rel DIVIDEBY rel
• Set-based
rel UNION rel
rel INTERSECT rel
rel MINUS rel
rel TIMES rel](https://image.slidesharecdn.com/20180220biologicaldatabasespart1vupload-180221090003/85/2018-02-20_biological_databases_part1_v_upload-40-320.jpg)