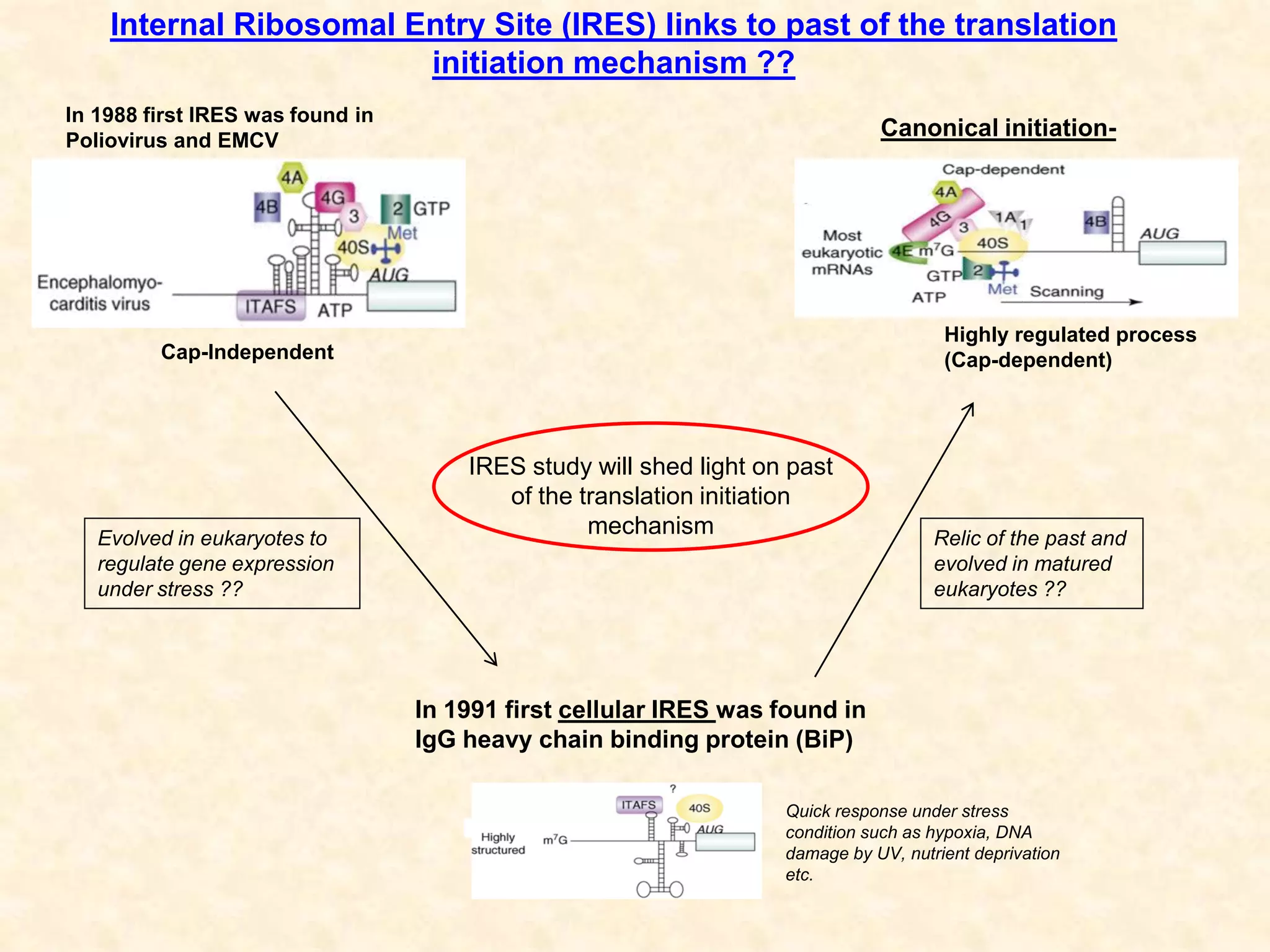
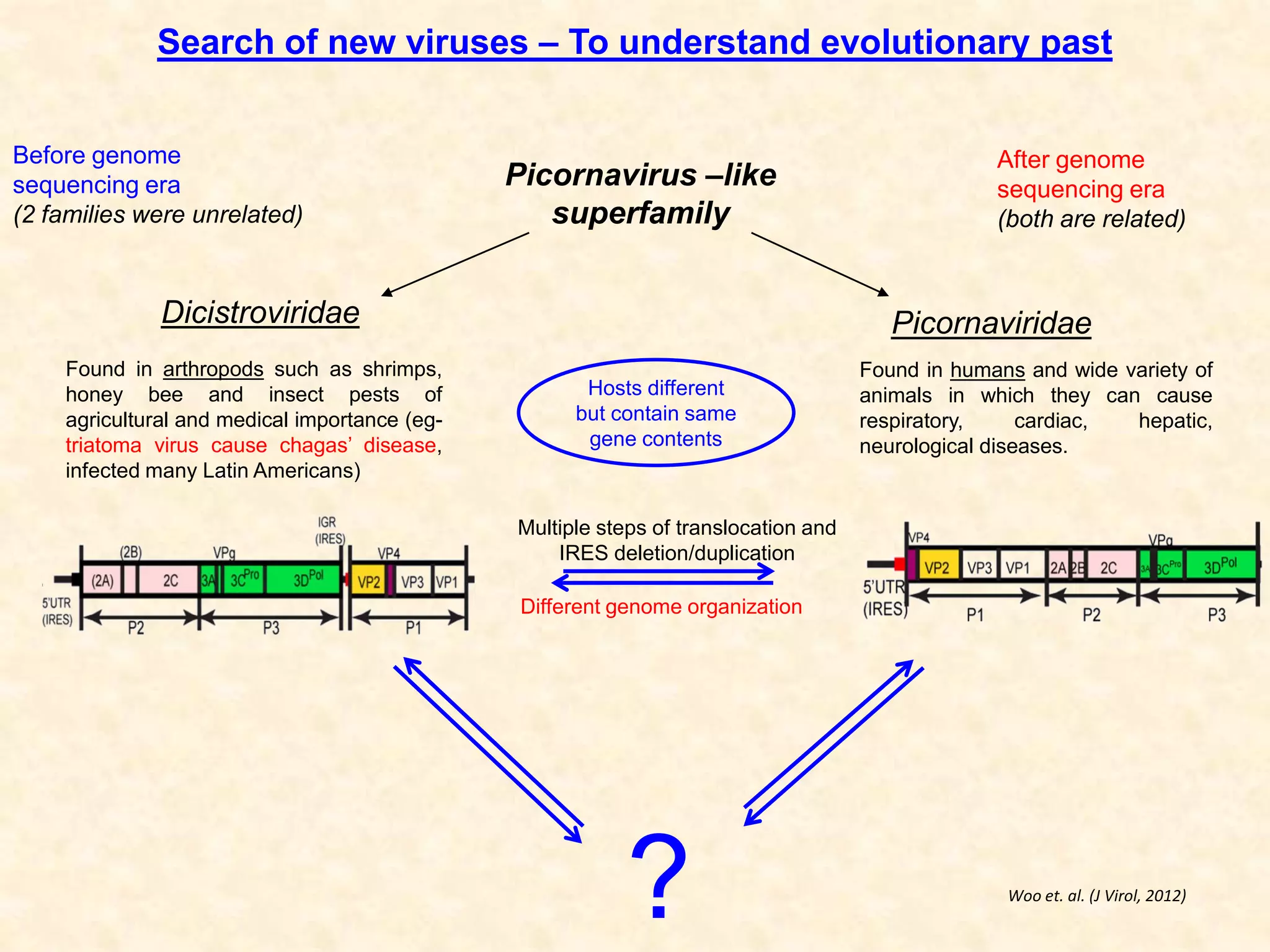
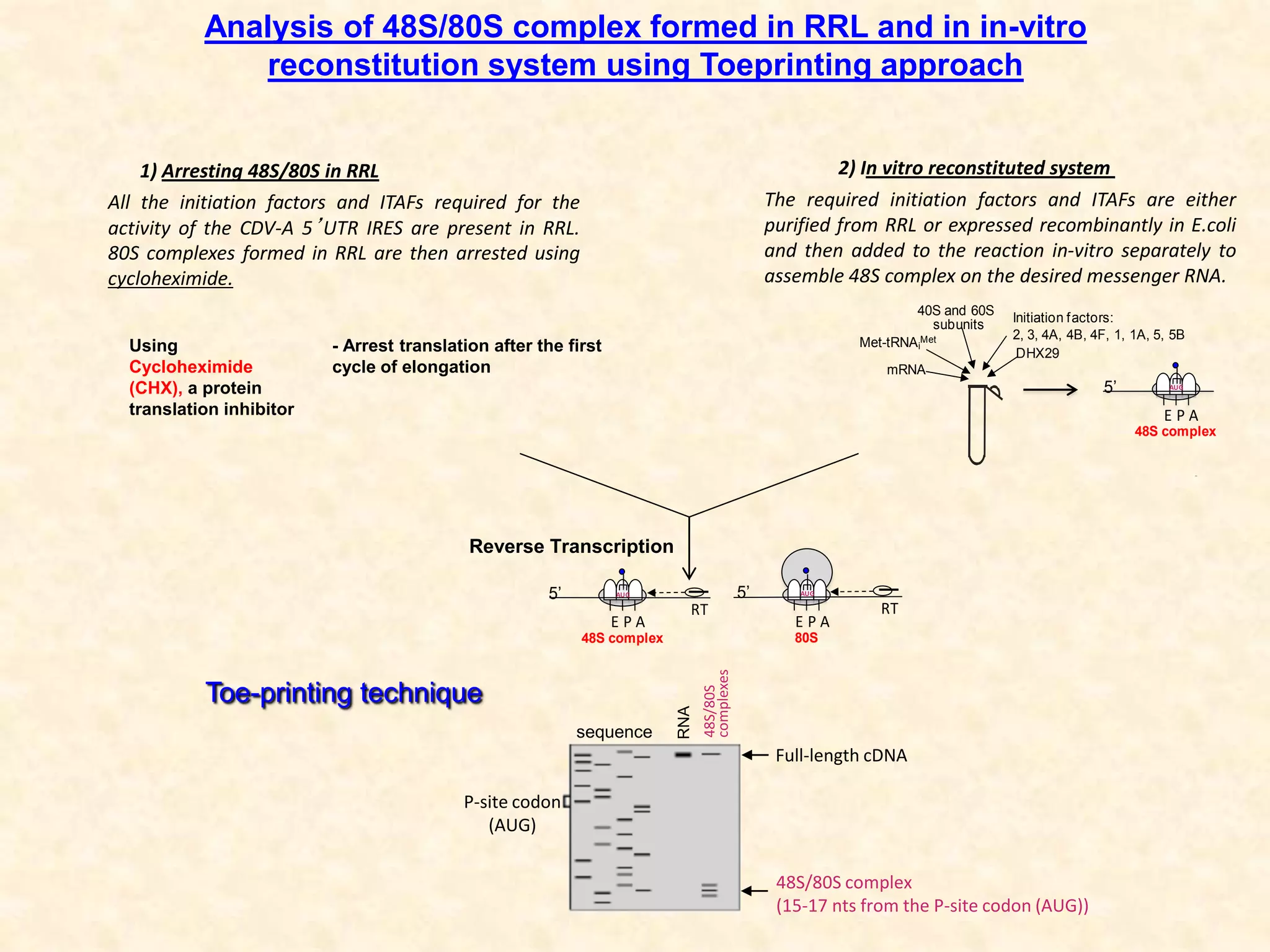
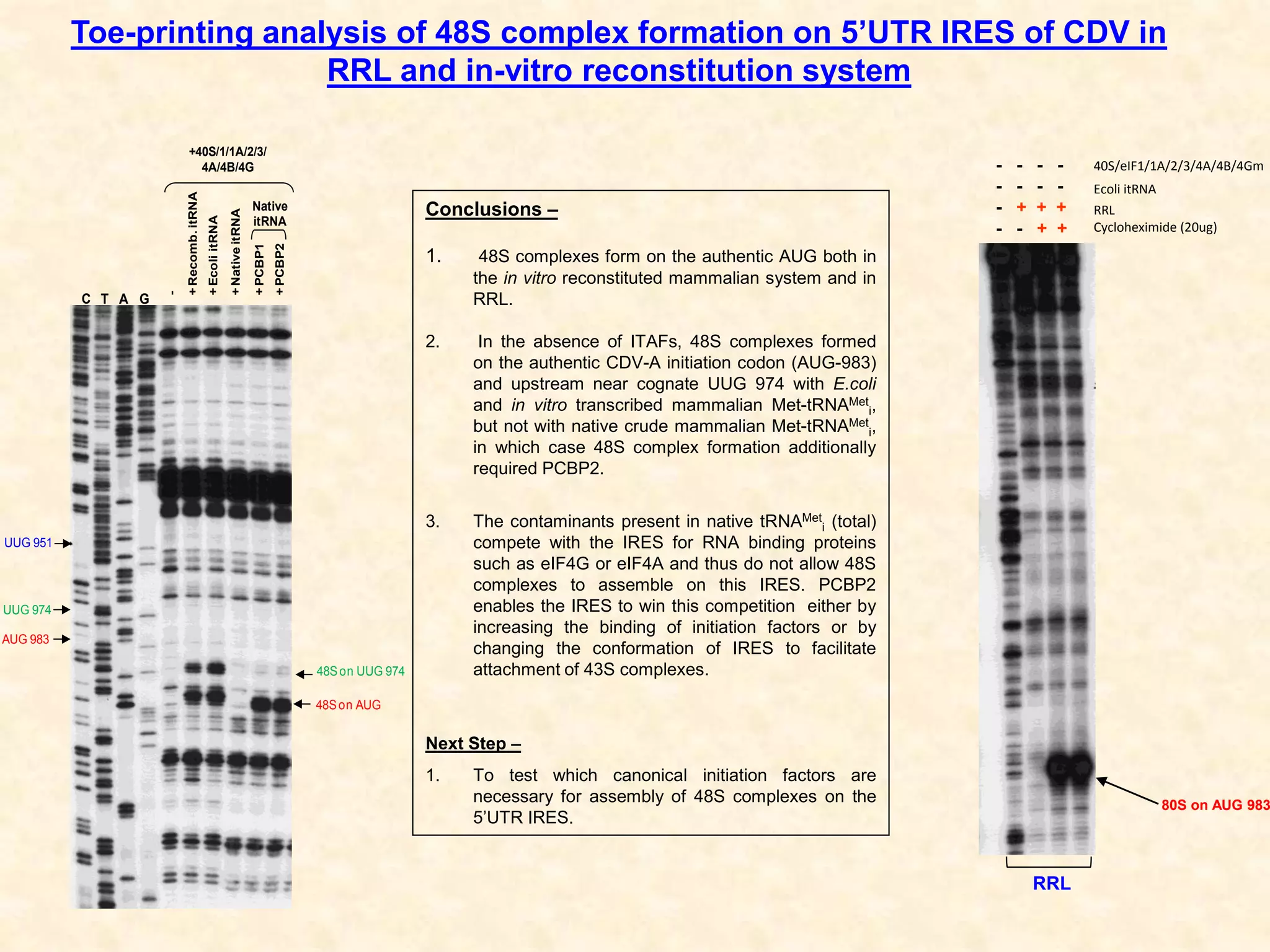
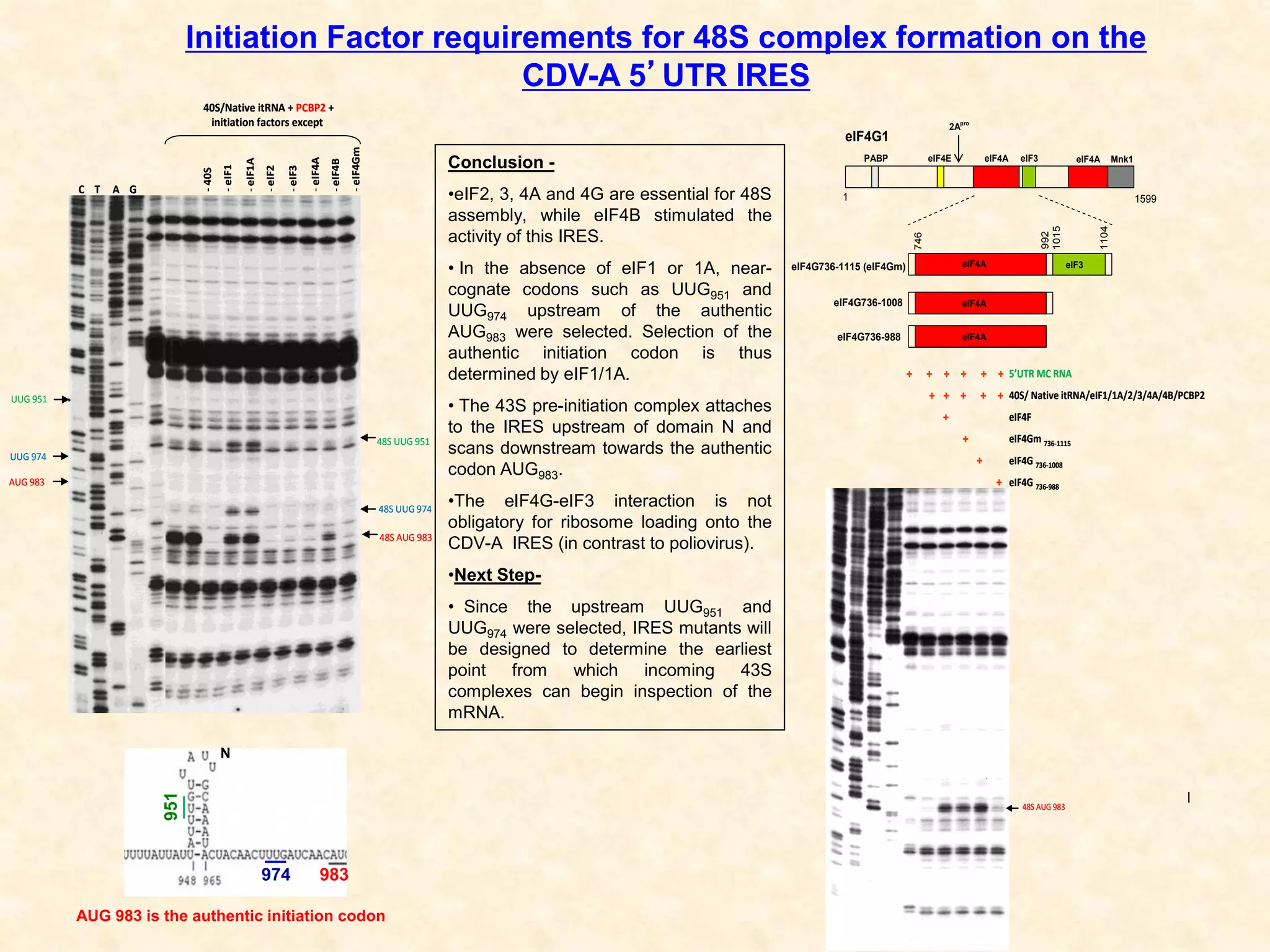
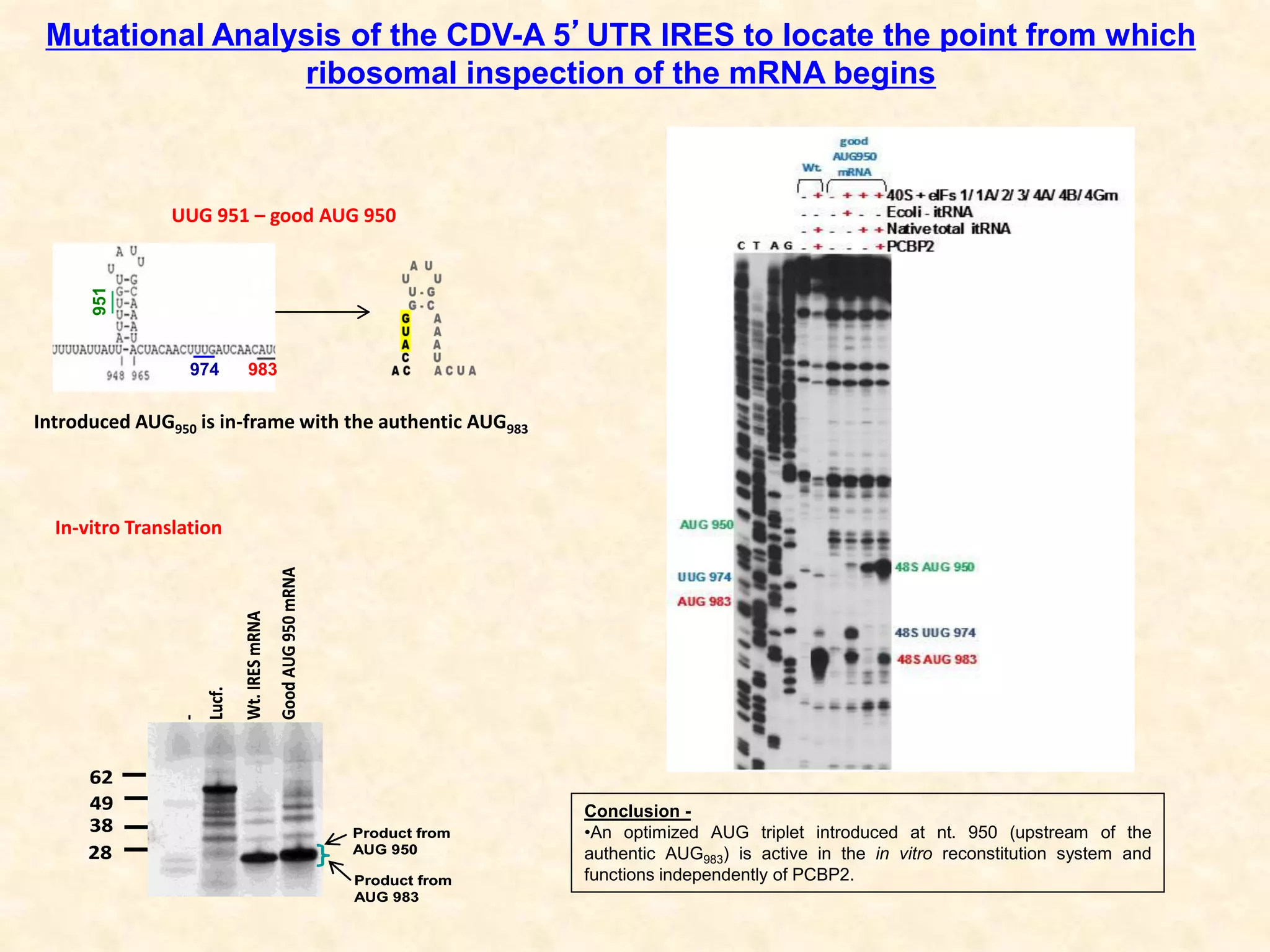
1) Researchers studied the internal ribosomal entry site (IRES) in the 5' untranslated region of the canine dicistrovirus (CDV-A) genome.
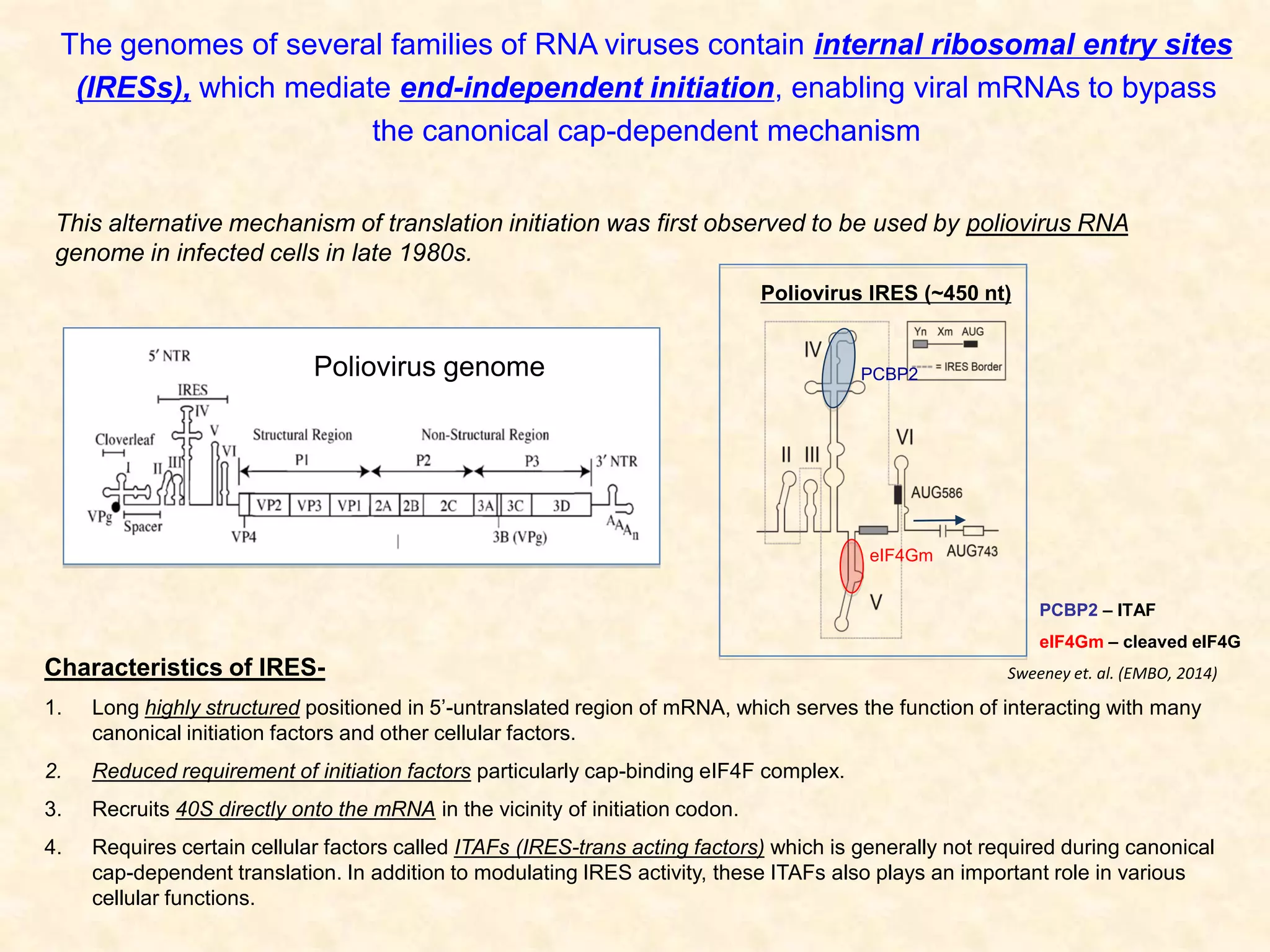
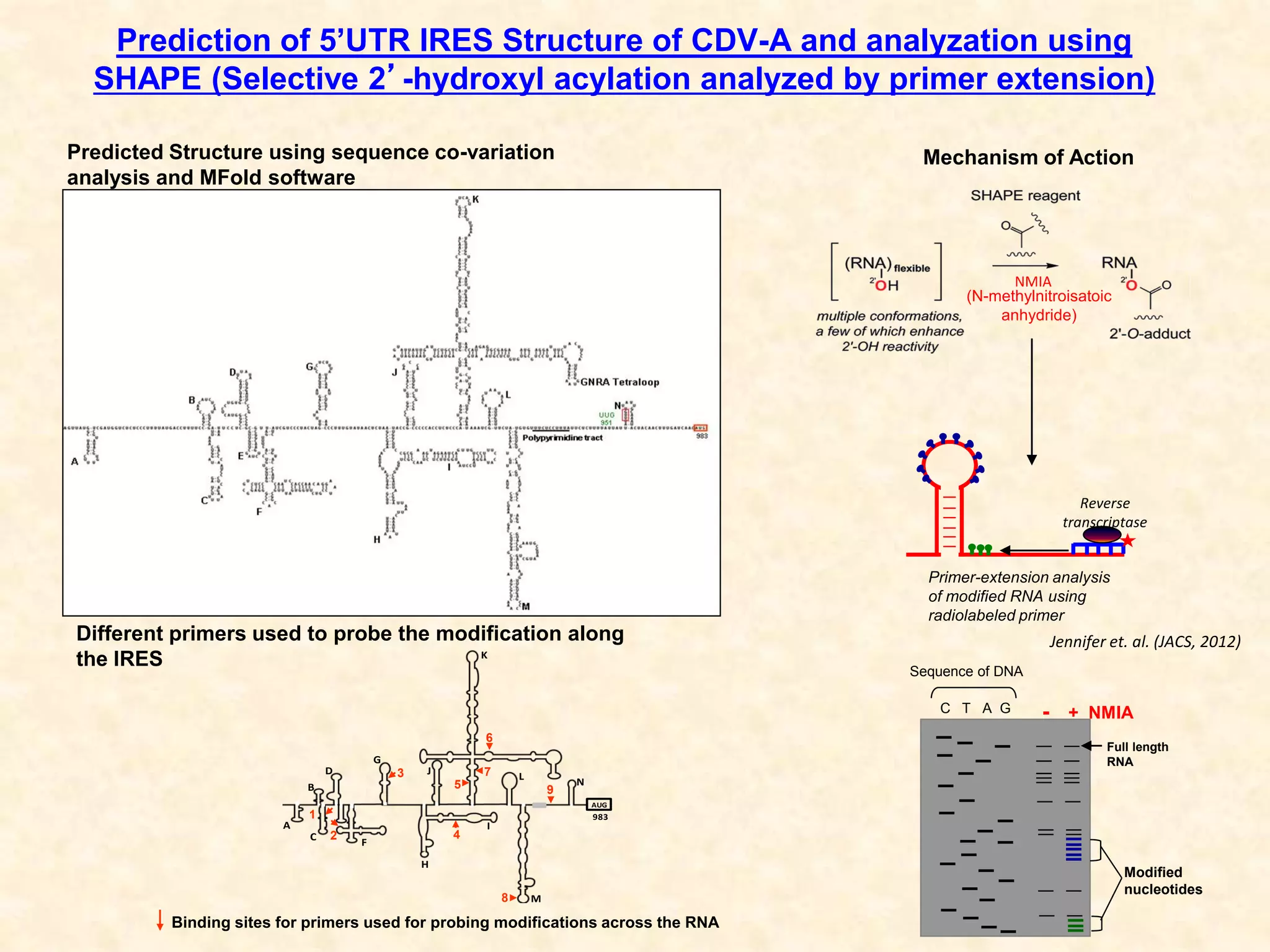
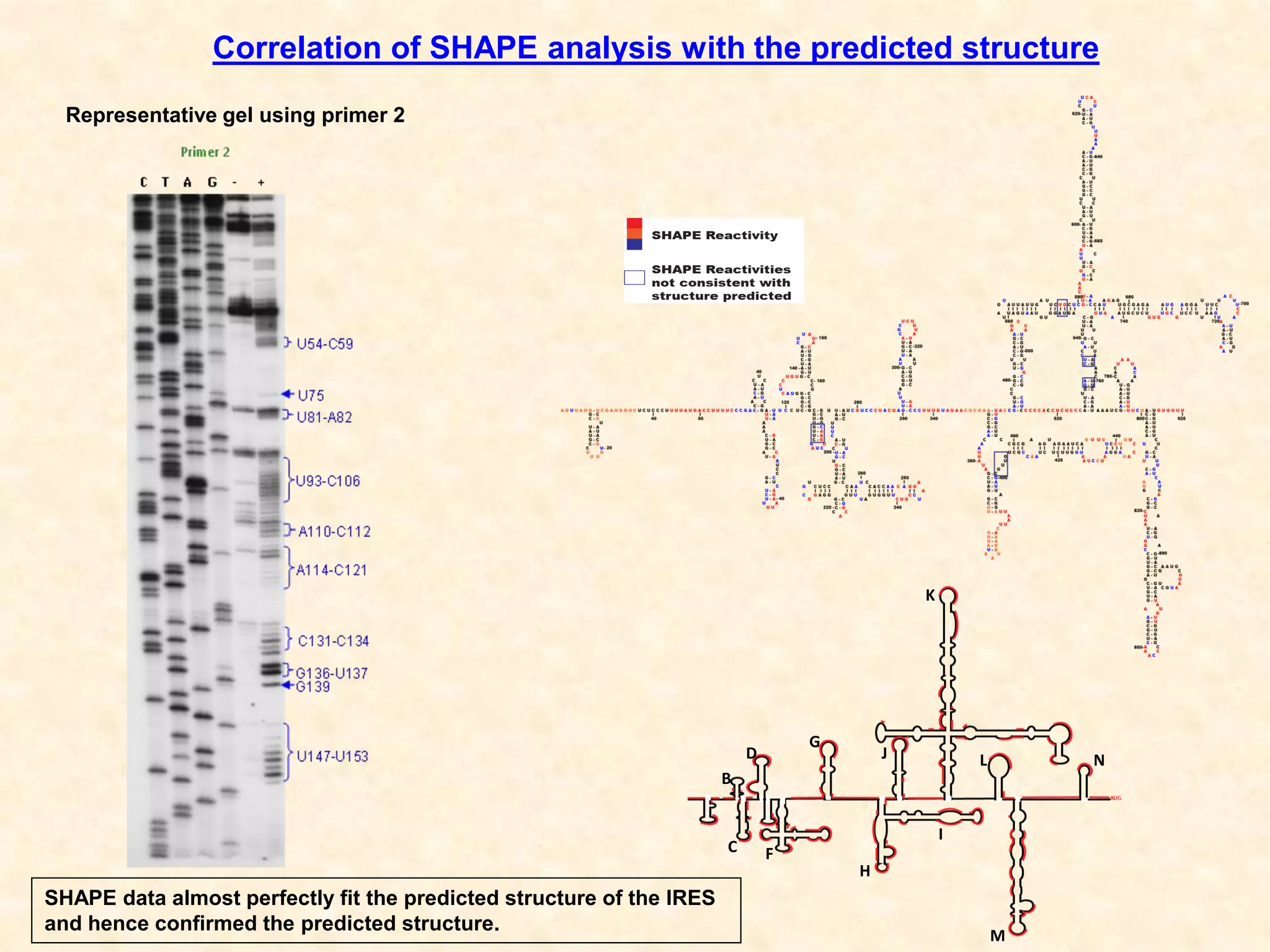
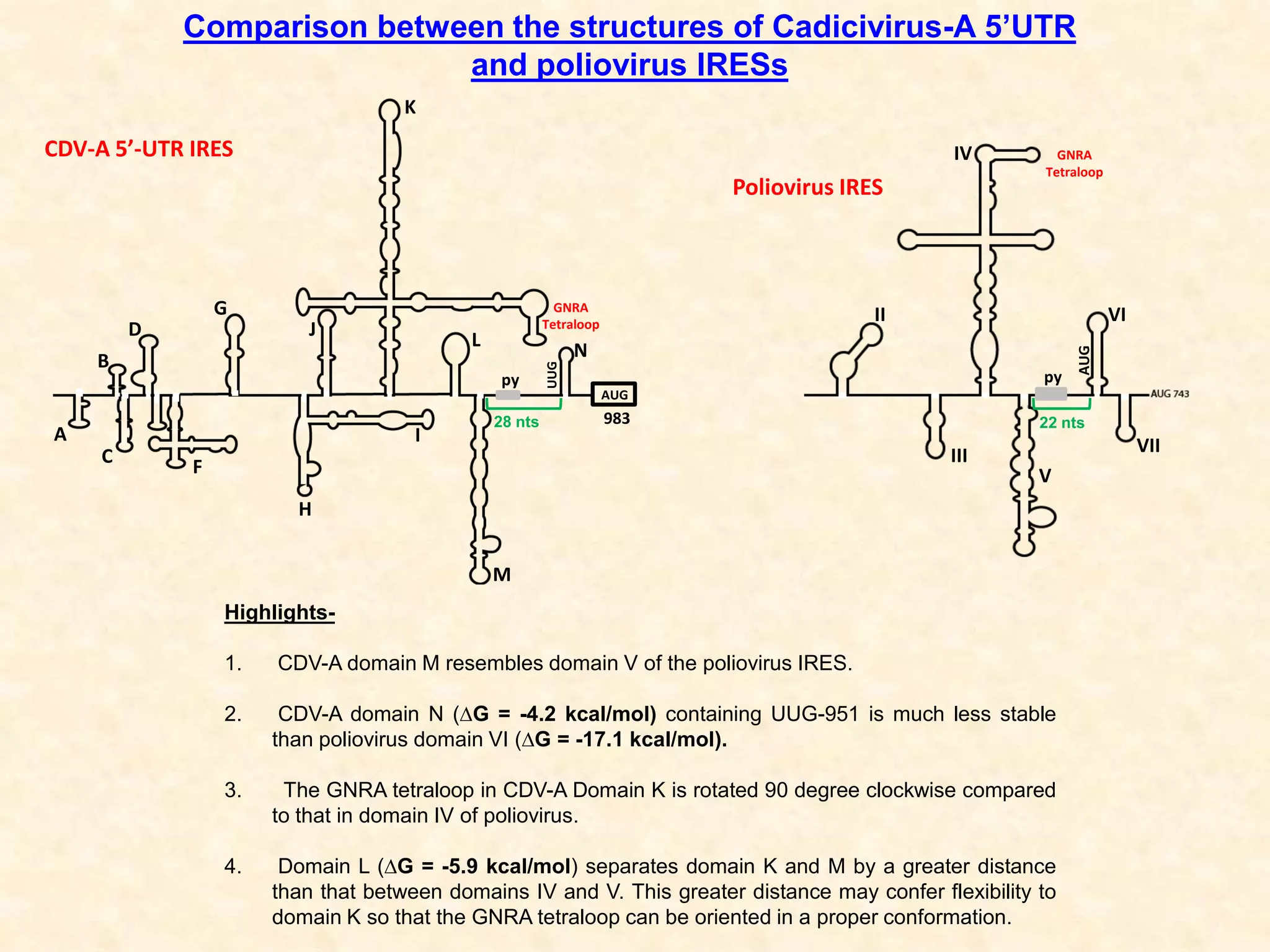
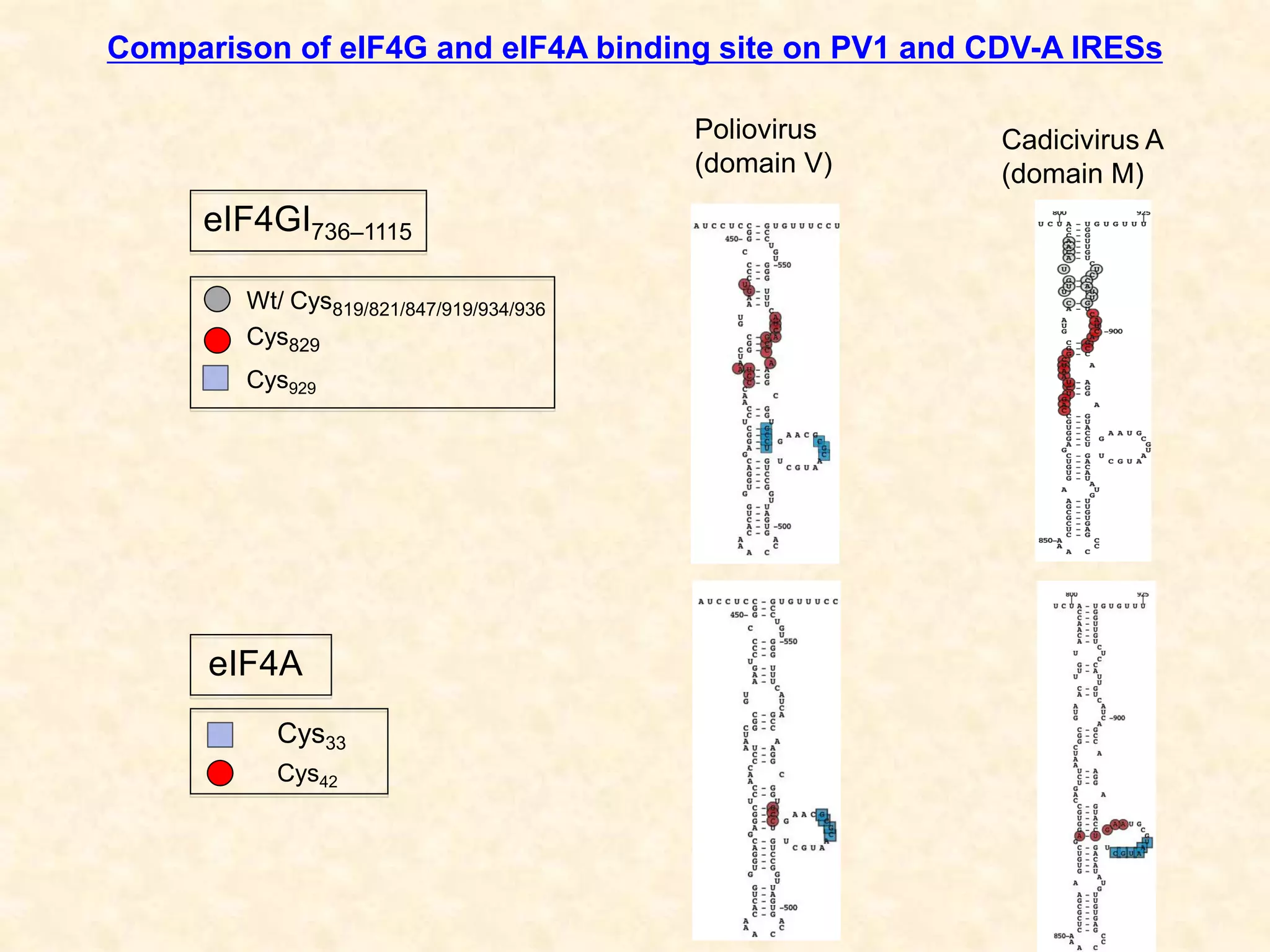
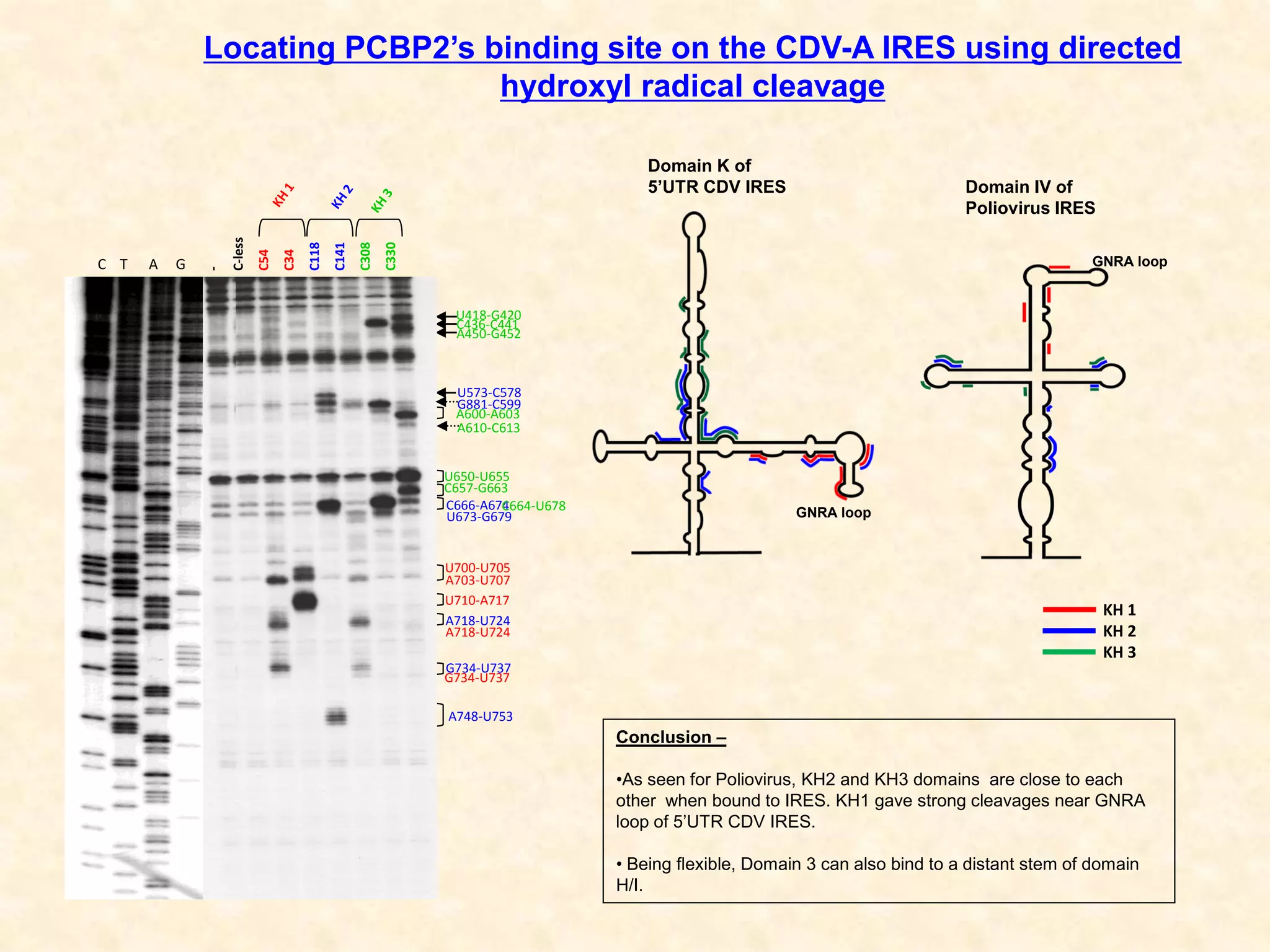
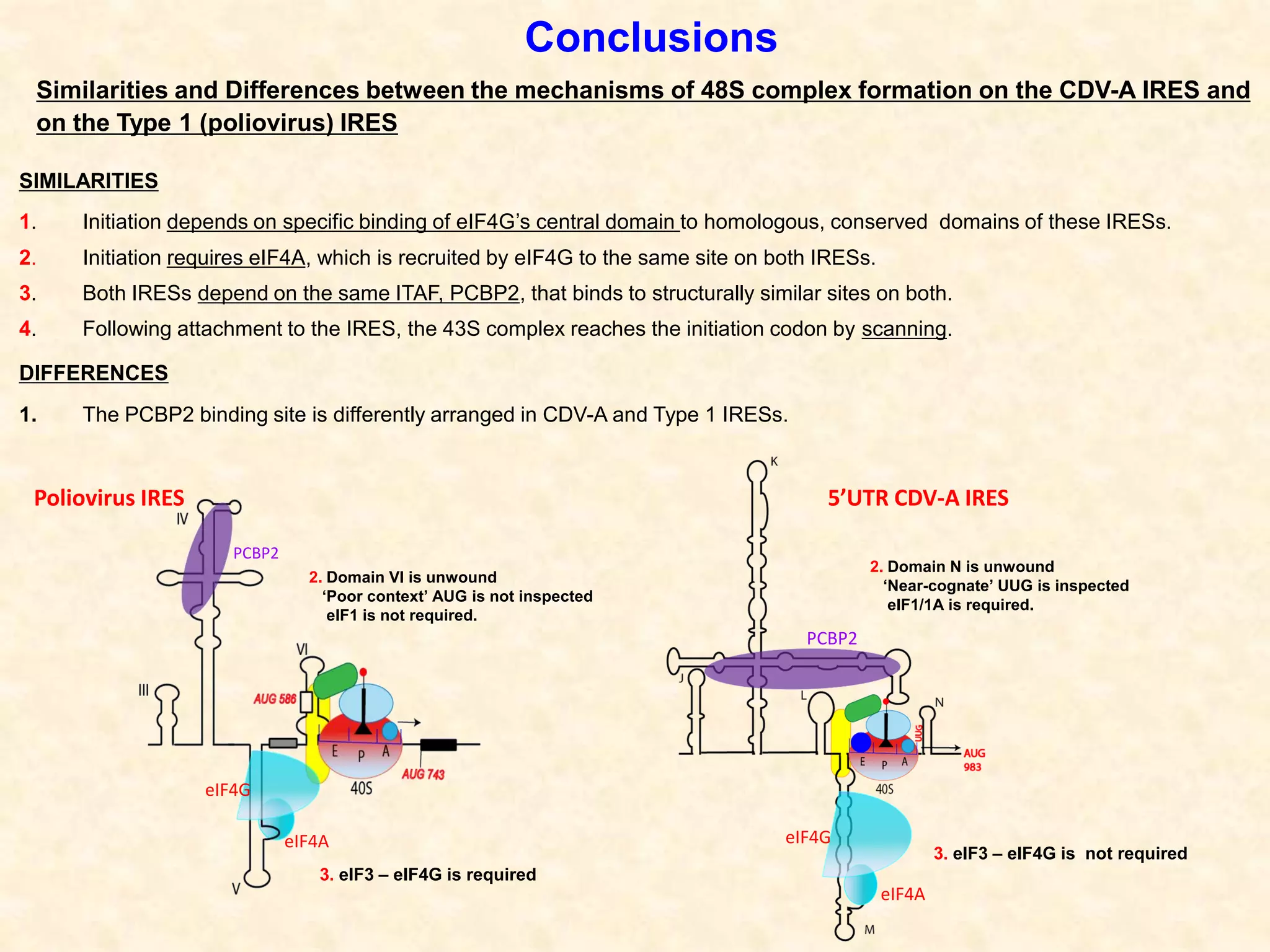
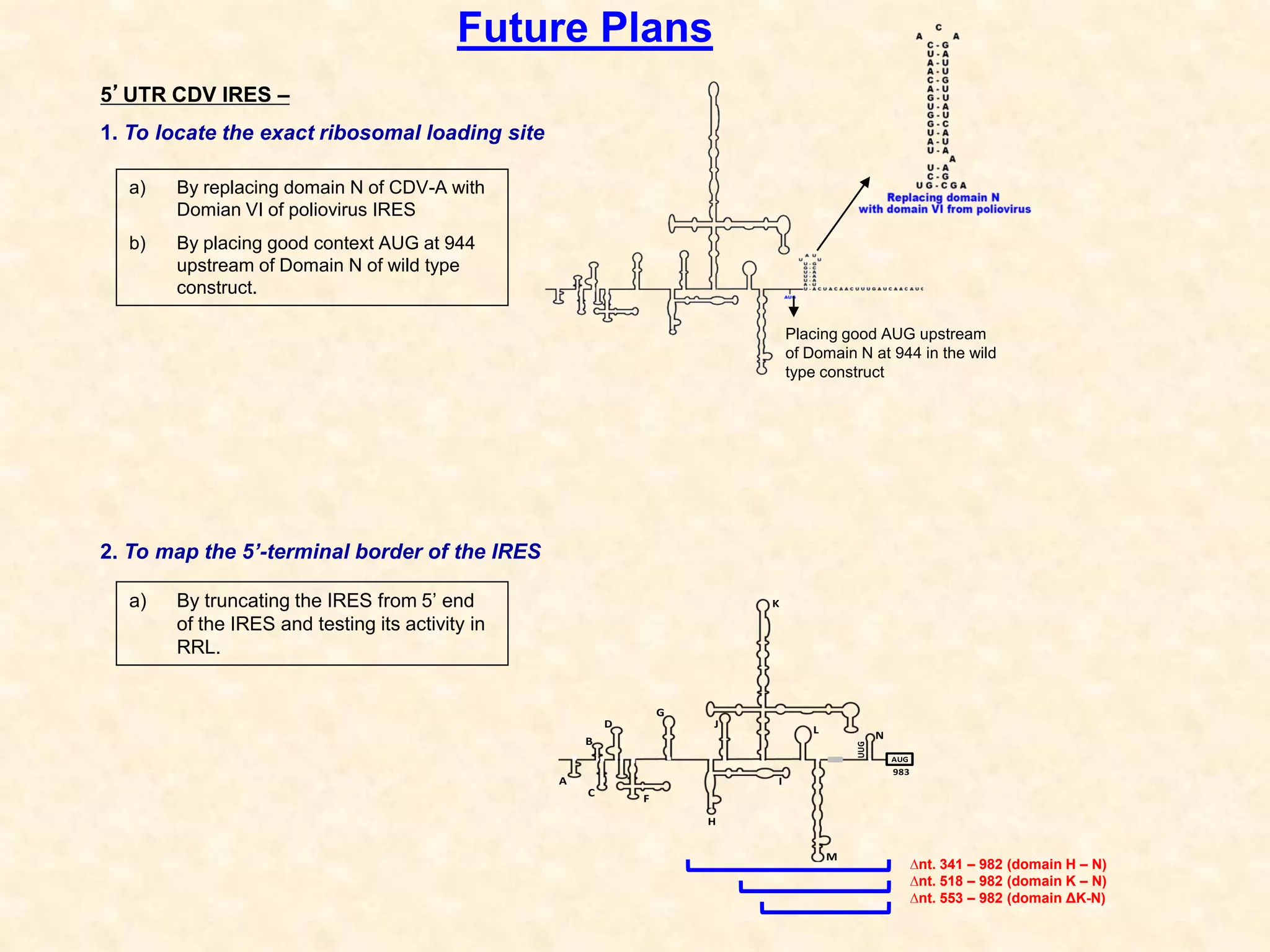
2) Using computational prediction and SHAPE analysis, they determined the secondary structure of the CDV-A 5'UTR IRES, which resembles the poliovirus IRES structure.
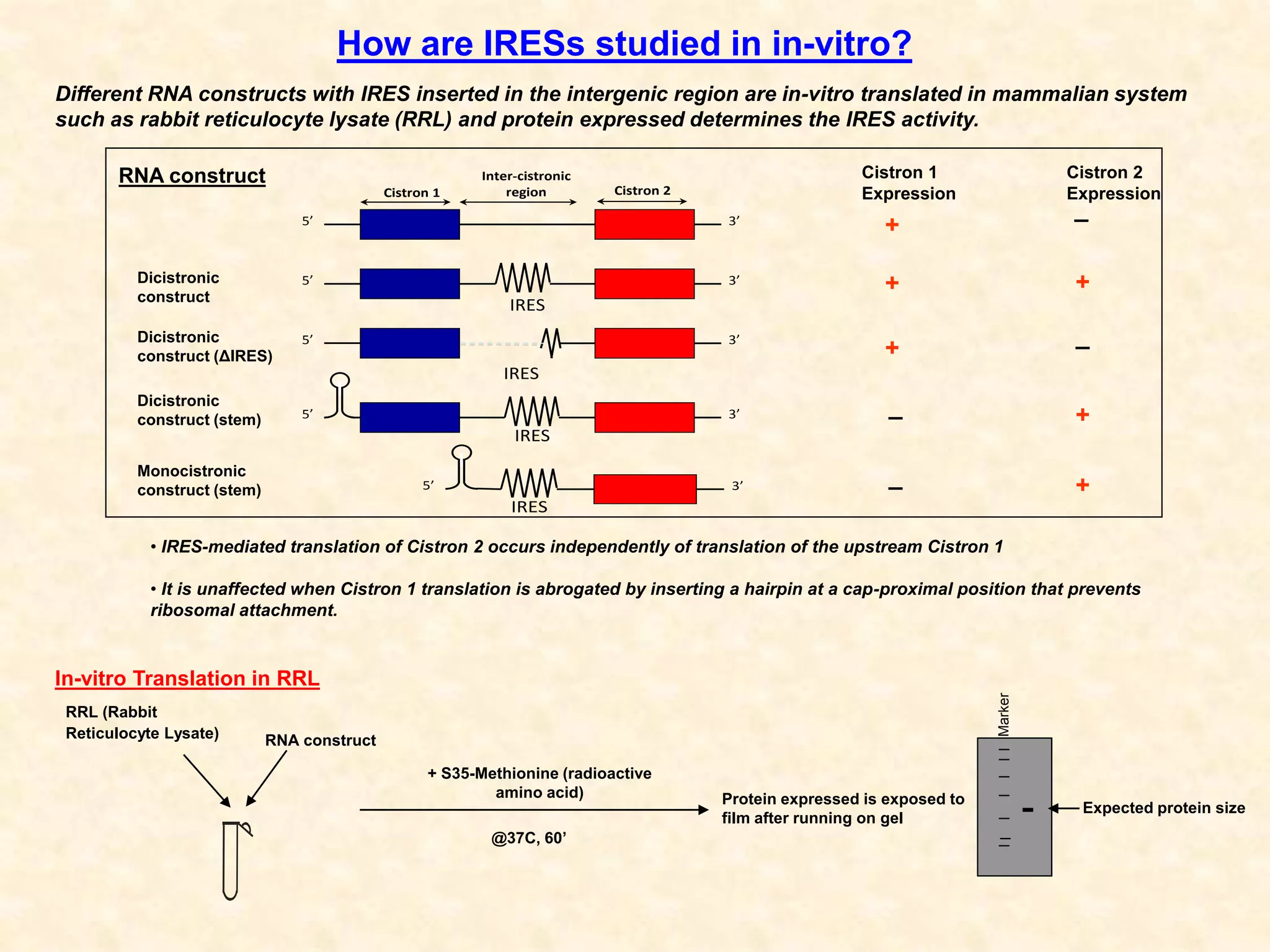
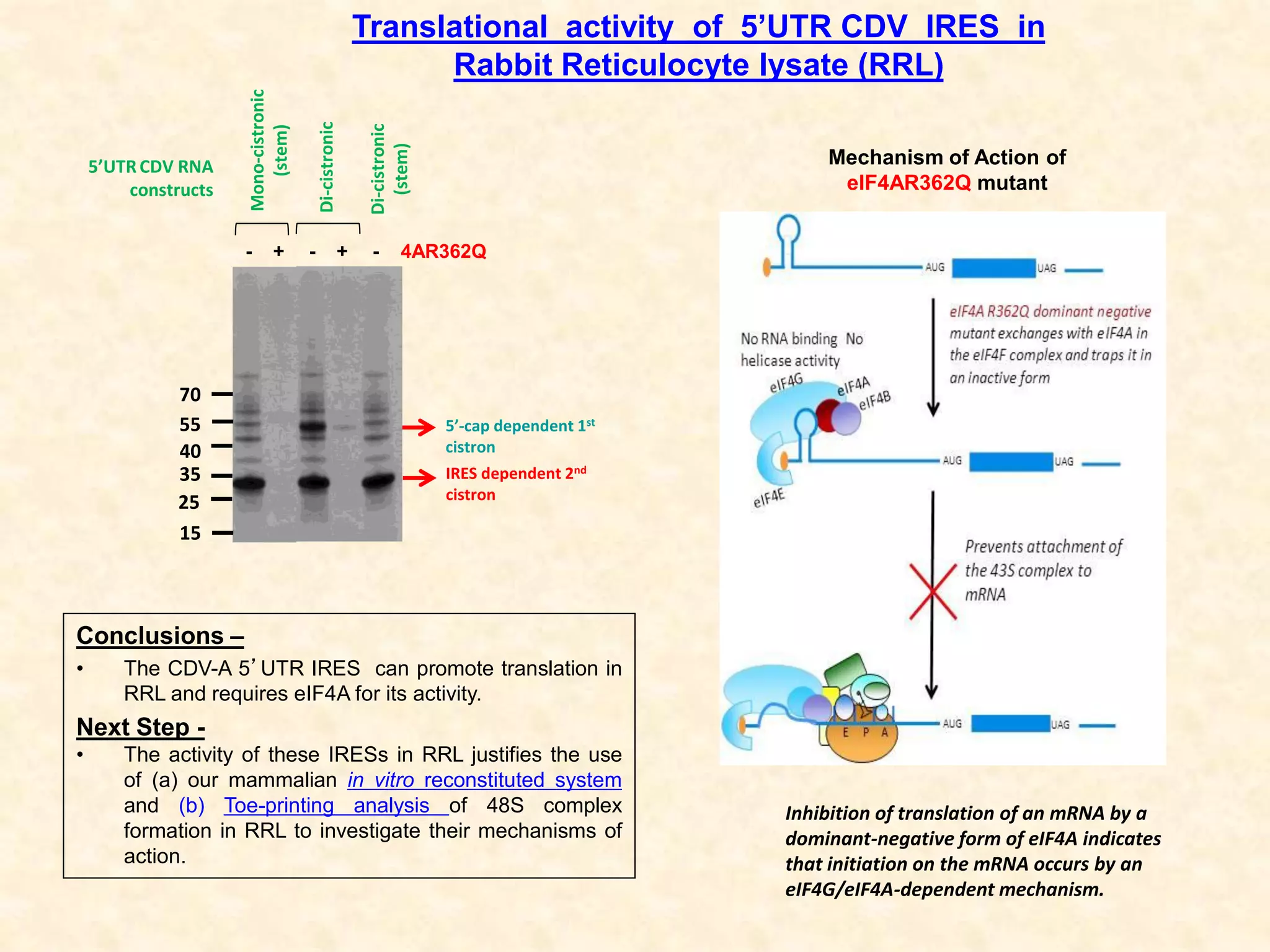
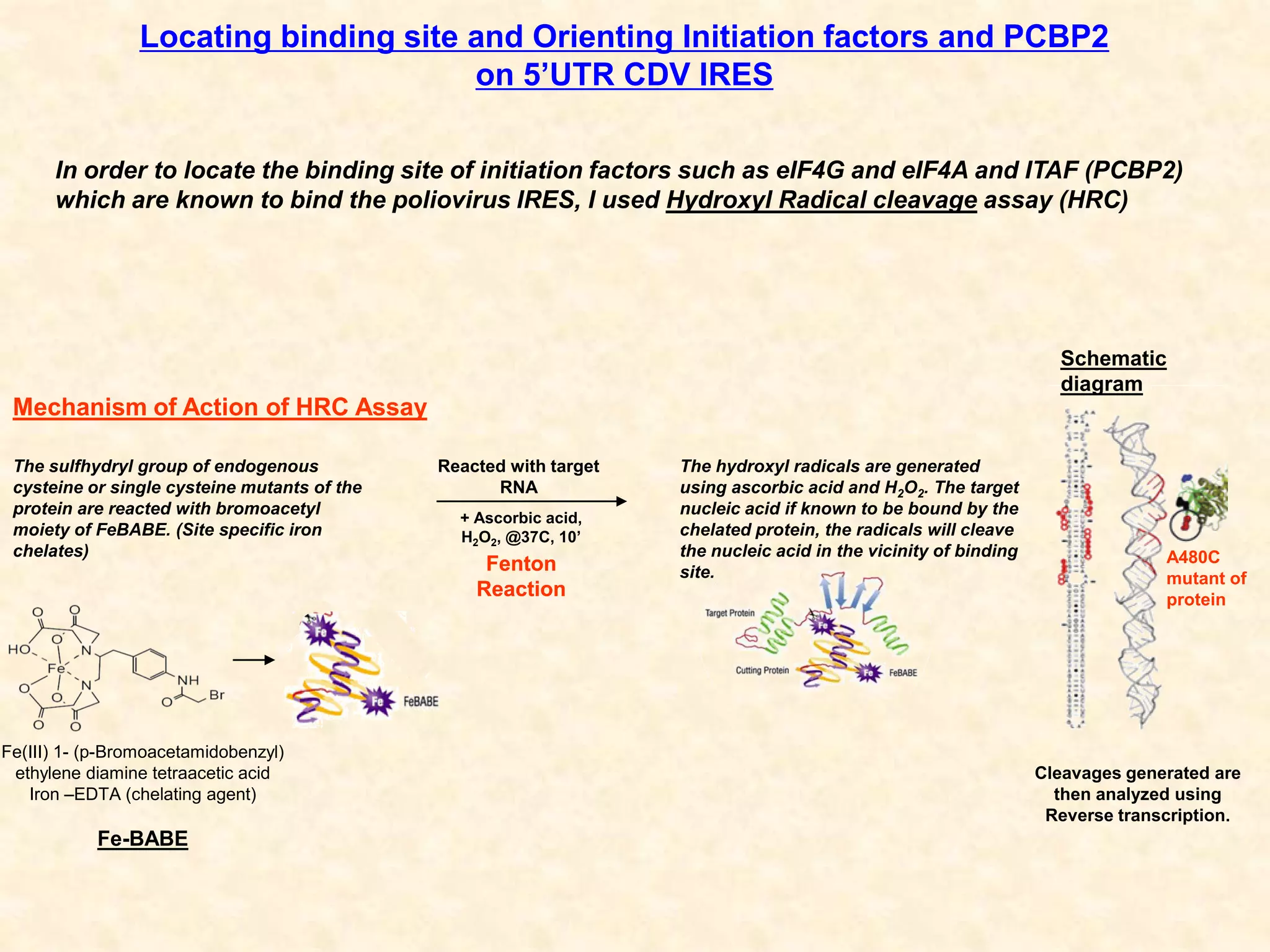
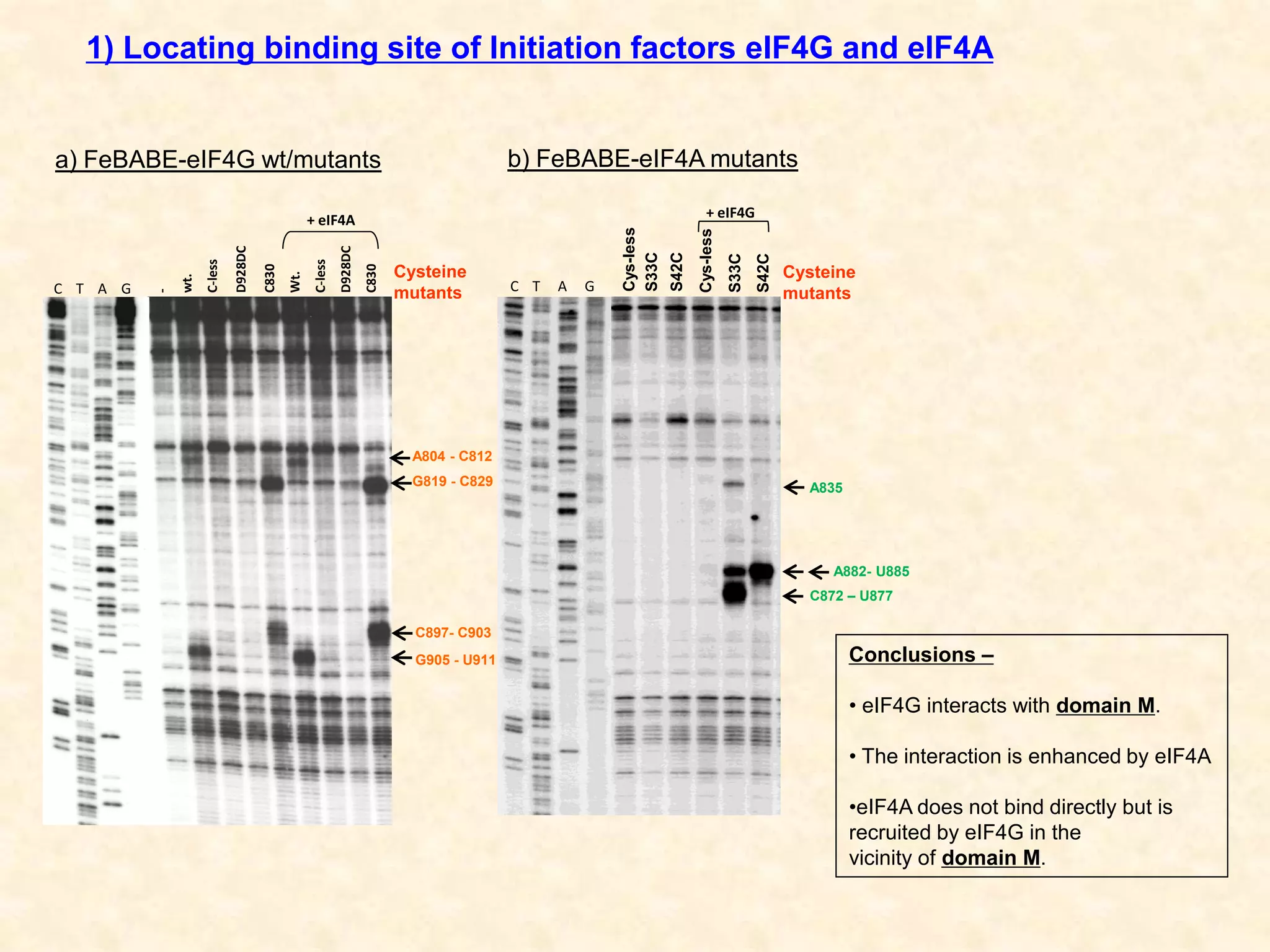
3) In vitro translation assays in rabbit reticulocyte lysate showed that the CDV-A 5'UTR IRES can direct cap-independent translation, and requires the initiation factor eIF4A.