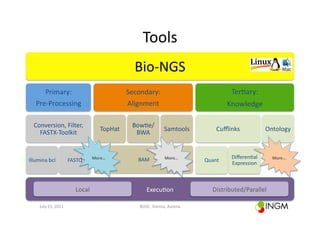
This document describes Bio-NGS, a BioRuby plugin for conducting programmable workflows for Next Generation Sequencing (NGS) data. Bio-NGS provides a software development framework, application, and project environment for analyzing NGS data. It integrates third-party bioinformatics tools as wrappers or bindings and allows for modular, reusable plugins. The document outlines features of the Bio-NGS application, software development framework, and project environment.








![Wrapper
module Bio
module Ngs
module Cufflinks
class Compare
include Bio::Command::Wrapper
set_program Bio::Ngs::Utils.binary("cufflinks/cuffcompare")
use_aliases
add_option "outprefix", :type => :string, :aliases => '-o', :default =>
"Comparison"
add_option "gtf_combine_file", :type => :string, :aliases => '-i'
add_option "gtf_reference", :type => :string, :aliases => '-r'
irb(main):001:0> require:type => :boolean, :aliases => '-R'
add_option "only_overlap", ‘bio-ngs’
add_option "discard_transfrags", :type => :boolean, :aliases => '-M’
irb(main):001:1> cuffcompare = Bio::Ngs::Cufflinks::Compare.new
irb(main):001:2> cuffcompare.params = {….}
irb(main):001:3> cuffcompare.run(:arguments=>[…])
end
end
=> #<Bio::Ngs::Cufflinks::Compare:0x0000000c1630f8 @program="/
end
end usr/local/lib/ruby/gems/1.9.1/gems/bio-ngs-0.2.1/lib/bio/ngs/
ext/bin/linux/cufflinks/cuffcompare", @options={}, @params={}>
July
15,
2011
BOSC,
Vienna,
Austria](https://image.slidesharecdn.com/raoul-bosc2011-biongs-110802102003-phpapp02/85/D03-NextGen-Bio-NGS-10-320.jpg)
![Tasks
No
binary
found
with
this
name:
setupBclToQseq.py
biongs
convert:qseq:fastq:samples_by_lane
SAMPLES
LANE
project
No
binary
found
with
this
name:
fastq_quality_boxplot_graph.sh
OUTPUT
-‐-‐-‐-‐-‐-‐-‐
No
binary
found
with
this
name:
blastn
biongs
project:new
[NAME]
No
binary
found
with
this
name:
blastx
history
biongs
project:update
[TYPE]
WARNING:
no
program
is
associated
with
BCLQSEQ
task,
does
-‐-‐-‐-‐-‐-‐-‐
not
make
sense
to
create
a
thor
task.
biongs
history:8
#
Task
convert:illumina:de:isoform
quality
WARNING:
no
program
is
associated
with
BLASTN
task,
does
not
PARAMETERS:
/Users/bonnalraoul/Desktop/
make
sense
to
create
a
thor
task.
RRep16giugno/DE_lane1-‐2-‐3-‐4-‐6-‐8/DE_lane1-‐2-‐3-‐4-‐6-‐8/ -‐-‐-‐-‐-‐-‐-‐
isoform_exp.diff
/Users/bonnalraoul/Desktop/ biongs
quality:boxplot
FASTQ_QUALITY_STATS
WARNING:
no
program
is
associated
with
BLASTX
task,
does
not
RRep16giugno/COMPARE_lane1-‐2-‐3-‐4-‐6-‐8/COMPA...
biongs
quality:fastq_stats
FASTQ
make
sense
to
create
a
thor
task.
biongs
quality:illumina_b_profile_raw
FASTQ
-‐-‐read-‐length=N
bwa
homology
biongs
quality:illumina_b_profile_svg
FASTQ
-‐-‐read-‐length=N
-‐-‐-‐
-‐-‐-‐-‐-‐-‐-‐-‐
biongs
quality:reads
FASTQ
biongs
bwa:aln:long
[FASTQ]
-‐-‐file-‐out=FILE_OUT
-‐-‐prefix=PREFIX
biongs
homology:convert:blast2text
[XML
FILE]
-‐-‐file-‐ biongs
quality:reads_coverage
FASTQ_QUALITY_STATS
biongs
bwa:aln:short
[FASTQ]
-‐-‐file-‐out=FILE_OUT
-‐-‐ out=FILE_OUT
biongs
quality:scacerplot
EXPR1
EXPR2
OUTPUT
prefix=PREFIX
biongs
homology:convert:go2json
biongs
quality:trim
FASTQ
biongs
bwa:index:long
[FASTA]
biongs
bwa:index:short
[FASTA]
biongs
homology:db:export
[TABLE]
-‐-‐fileout=FILEOUT
rna
biongs
bwa:sam:paired
-‐-‐fastq=one
two
three
-‐-‐file-‐ biongs
homology:db:init
out=FILE_OUT
-‐-‐prefix=PREFIX
-‐-‐sai=one
two
three
-‐-‐-‐
biongs
bwa:sam:single
[SAI]
-‐-‐fastq=FASTQ
-‐-‐file-‐ biongs
homology:download:all
biongs
rna:compare
GTF_REF
OUTPUTDIR
out=FILE_OUT
-‐-‐prefix=PREFIX
biongs
homology:download:goannota(on
GTFS_QUANTIFICATION
biongs
homology:download:uniprot
biongs
rna:idx2fasta
INDEX
FASTA
convert
biongs
homology:load:blast
[FILE]
biongs
rna:mapquant
DIST
INDEX
OUTPUTDIR
FASTQS
-‐-‐-‐-‐-‐-‐-‐
biongs
homology:load:goa
biongs
rna:quant
GTF
OUTPUTDIR
BAM
biongs
convert:bam:extract_genes
BAM
GENES
-‐-‐ensembl-‐ biongs
homology:report:blast
biongs
rna:tophat
DIST
INDEX
OUTPUTDIR
FASTQS
release=N
-‐o,
-‐-‐output=OUTPUT
biongs
convert:bam:merge
-‐i,
-‐-‐input-‐bams=one
two
three
ontology
sff
biongs
convert:bam:sort
BAM
[PREFIX]
-‐-‐-‐-‐-‐-‐-‐-‐
-‐-‐-‐
biongs
convert:bcl:qseq:convert
RUN
OUTPUT
[JOBS]
biongs
ontology:db:export
[TABLE]
-‐-‐fileout=FILEOUT
biongs
sff:extract
[FILE]
biongs
convert:illumina:de:gene
DIFF
GTF
biongs
ontology:db:init
biongs
convert:illumina:de:isoform
DIFF
GTF
biongs
ontology:download:all
biongs
convert:illumina:de:rename_qs
DIFF_FILE
NAMES
biongs
ontology:download:go
biongs
convert:illumina:fastq:trim_b
FASTQ
biongs
ontology:download:goslim
biongs
convert:illumina:humanize:build_compare_kb
GTF
biongs
ontology:load:genego
[FILE]
biongs
convert:illumina:humanize:isoform_exp
GTF
ISOFORM
biongs
ontology:load:go
[FILE]
biongs
convert:qseq:fastq:by_file
FIRST
OUTPUT
biongs
ontology:report:go
biongs
convert:qseq:fastq:by_lane
LANE
OUTPUT
biongs
convert:qseq:fastq:by_lane_index
LANE
INDEX
OUTPUT
July
15,
2011
BOSC,
Vienna,
Austria](https://image.slidesharecdn.com/raoul-bosc2011-biongs-110802102003-phpapp02/85/D03-NextGen-Bio-NGS-11-320.jpg)
![N o
B i n a r y
Tasks
Task
disabled
No
binary
found
with
this
name:
setupBclToQseq.py
biongs
convert:qseq:fastq:samples_by_lane
SAMPLES
LANE
project
Keep
OUTPUT
No
binary
found
with
this
name:
fastq_quality_boxplot_graph.sh
No
binary
found
with
this
name:
blastn
-‐-‐-‐-‐-‐-‐-‐
biongs
project:new
[NAME]
everything
No
binary
found
with
this
name:
blastx
history
biongs
project:update
[TYPE]
WARNING:
no
program
is
associated
with
BCLQSEQ
task,
does
-‐-‐-‐-‐-‐-‐-‐
not
make
sense
to
create
a
thor
task.
biongs
history:8
#
Task
convert:illumina:de:isoform
organized
quality
WARNING:
no
program
is
associated
with
BLASTN
task,
does
not
PARAMETERS:
/Users/bonnalraoul/Desktop/
make
sense
to
create
a
thor
task.
RRep16giugno/DE_lane1-‐2-‐3-‐4-‐6-‐8/DE_lane1-‐2-‐3-‐4-‐6-‐8/ -‐-‐-‐-‐-‐-‐-‐
isoform_exp.diff
/Users/bonnalraoul/Desktop/ biongs
quality:boxplot
FASTQ_QUALITY_STATS
WARNING:
no
program
is
associated
with
BLASTX
task,
does
not
RRep16giugno/COMPARE_lane1-‐2-‐3-‐4-‐6-‐8/COMPA...
biongs
quality:fastq_stats
FASTQ
make
sense
to
create
a
thor
task.
biongs
quality:illumina_b_profile_raw
FASTQ
-‐-‐read-‐length=N
bwa
homology
biongs
quality:illumina_b_profile_svg
FASTQ
-‐-‐read-‐length=N
-‐-‐-‐
-‐-‐-‐-‐-‐-‐-‐-‐
biongs
quality:reads
FASTQ
biongs
bwa:aln:long
[FASTQ]
-‐-‐file-‐out=FILE_OUT
-‐-‐prefix=PREFIX
biongs
homology:convert:blast2text
[XML
FILE]
-‐-‐file-‐ biongs
quality:reads_coverage
FASTQ_QUALITY_STATS
biongs
bwa:aln:short
[FASTQ]
-‐-‐file-‐out=FILE_OUT
-‐-‐ out=FILE_OUT
biongs
quality:scacerplot
EXPR1
EXPR2
OUTPUT
prefix=PREFIX
biongs
bwa:index:long
[FASTA]
biongs
homology:convert:go2json
Repor(ng
biongs
quality:trim
FASTQ
biongs
bwa:index:short
[FASTA]
biongs
bwa:sam:paired
-‐-‐fastq=one
two
three
-‐-‐file-‐
Recall
an
biongs
homology:db:export
[TABLE]
-‐-‐fileout=FILEOUT
rna
biongs
homology:db:init
-‐-‐-‐
old
out=FILE_OUT
-‐-‐prefix=PREFIX
-‐-‐sai=one
two
three
biongs
bwa:sam:single
[SAI]
-‐-‐fastq=FASTQ
-‐-‐file-‐
out=FILE_OUT
-‐-‐prefix=PREFIX
biongs
homology:download:all
biongs
homology:download:goannota(on
biongs
rna:compare
GTF_REF
OUTPUTDIR
GTFS_QUANTIFICATION
convert
analysis
biongs
homology:download:uniprot
biongs
homology:load:blast
[FILE]
biongs
rna:idx2fasta
INDEX
FASTA
biongs
rna:mapquant
DIST
INDEX
OUTPUTDIR
FASTQS
-‐-‐-‐-‐-‐-‐-‐
biongs
homology:load:goa
biongs
rna:quant
GTF
OUTPUTDIR
BAM
biongs
convert:bam:extract_genes
BAM
GENES
-‐-‐ensembl-‐ biongs
homology:report:blast
biongs
rna:tophat
DIST
INDEX
OUTPUTDIR
FASTQS
release=N
-‐o,
-‐-‐output=OUTPUT
biongs
convert:bam:merge
-‐i,
-‐-‐input-‐bams=one
two
three
ontology
sff
biongs
convert:bam:sort
BAM
[PREFIX]
-‐-‐-‐-‐-‐-‐-‐-‐
-‐-‐-‐
biongs
convert:bcl:qseq:convert
RUN
OUTPUT
[JOBS]
biongs
ontology:db:export
[TABLE]
-‐-‐fileout=FILEOUT
biongs
sff:extract
[FILE]
biongs
convert:illumina:de:gene
DIFF
GTF
biongs
ontology:db:init
biongs
convert:illumina:de:isoform
DIFF
GTF
biongs
ontology:download:all
biongs
convert:illumina:de:rename_qs
DIFF_FILE
NAMES
biongs
ontology:download:go
biongs
convert:illumina:fastq:trim_b
FASTQ
biongs
ontology:download:goslim
biongs
convert:illumina:humanize:build_compare_kb
GTF
biongs
ontology:load:genego
[FILE]
biongs
convert:illumina:humanize:isoform_exp
GTF
ISOFORM
biongs
ontology:load:go
[FILE]
biongs
convert:qseq:fastq:by_file
FIRST
OUTPUT
biongs
convert:qseq:fastq:by_lane
LANE
OUTPUT
biongs
ontology:report:go
Basic
Advanced
biongs
convert:qseq:fastq:by_lane_index
LANE
INDEX
OUTPUT
July
15,
2011
BOSC,
Vienna,
Austria](https://image.slidesharecdn.com/raoul-bosc2011-biongs-110802102003-phpapp02/85/D03-NextGen-Bio-NGS-12-320.jpg)
![Tasks
class Rna < Thor
desc "mapquant DIST INDEX OUTPUTDIR FASTQS", "map and quantify"
method_option :paired, :type => :boolean, :default => false, :desc => 'Are reads paired? If you chose
this option pass just the basename
of the file without forward/reverse
and .fastq'
def mapquant(dist, index, outputdir, fastqs)
#tophat
invoke :tophat, [dist, index, outputdir, fastqs], :paired=>options.paired
#cufflinks quantification on gtf
invoke :quant, ["#{index}.gtf", File.join(outputdir,"quantification"), File.join(outputdir,"accepted_hits_sort.bam")]
end
…
end
July
15,
2011
BOSC,
Vienna,
Austria](https://image.slidesharecdn.com/raoul-bosc2011-biongs-110802102003-phpapp02/85/D03-NextGen-Bio-NGS-13-320.jpg)
![Tasks
class Rna < Thor
# you'll end up with 3 accept file, regular, sorted, sorted-indexed
desc "tophat DIST INDEX OUTPUTDIR FASTQS", "run tophat as from command line, default 6 processors and then create a
sorted bam indexed."
method_option :paired, :type => :boolean, :default => false, :desc => 'Are reads paired? If you chose this option pass
just the…’
Bio::Ngs::Tophat.new.thor_task(self, :tophat) do |wrapper, task, dist, index, outputdir, fastqs|
wrapper.params = task.options #merge passed options to the wrapper.
wrapper.params = {"mate-inner-dist"=>dist, "output-dir"=>outputdir, "num-threads"=>6, "solexa1.3-quals"=>true}
fastq_files = task.options[:paired] ? ["#{fastqs}_forward.fastq","#{fastqs}_reverse.fastq"] : ["#{fastqs}"]
wrapper.run :arguments=>[index, fastq_files ].flatten, :separator=>"="
class Rna < Thor
accepted_hits_bam_fn = File.join(outputdir, "accepted_hits.bam")
desc "mapquant DIST INDEX OUTPUTDIR FASTQS", "map and quantify"
method_option :paired, "convert:bam:sort", :default => false, :desc => 'Are reads paired? If you chose
task.invoke :type => :boolean, [accepted_hits_bam_fn] # call the sorting procedure.
end this option pass just the basename
end of the file without forward/reverse
and .fastq'
def mapquant(dist, index, outputdir, fastqs)
#tophat
invoke :tophat, [dist, index, outputdir, fastqs], :paired=>options.paired
#cufflinks quantification on gtf
invoke :quant, ["#{index}.gtf", File.join(outputdir,"quantification"), File.join(outputdir,"accepted_hits_sort.bam")]
end
…
end
July
15,
2011
BOSC,
Vienna,
Austria](https://image.slidesharecdn.com/raoul-bosc2011-biongs-110802102003-phpapp02/85/D03-NextGen-Bio-NGS-14-320.jpg)
![Tasks
class Rna < Thor
# you'll end up with 3 accept file, regular, sorted, sorted-indexed
desc "tophat DIST INDEX OUTPUTDIR FASTQS", "run tophat as from command line, default 6 processors and then create a
sorted bam indexed."
method_option :paired, :type => :boolean, :default => false, :desc => 'Are reads paired? If you chose this option pass
just the…’
Bio::Ngs::Tophat.new.thor_task(self, :tophat) do |wrapper, task, dist, index, outputdir, fastqs|
wrapper.params = task.options #merge passed options to the wrapper.
wrapper.params = {"mate-inner-dist"=>dist, "output-dir"=>outputdir, "num-threads"=>6, "solexa1.3-quals"=>true}
fastq_files = task.options[:paired] ? ["#{fastqs}_forward.fastq","#{fastqs}_reverse.fastq"] : ["#{fastqs}"]
wrapper.run :arguments=>[index, fastq_files ].flatten, :separator=>"="
class Rna < Thor
accepted_hits_bam_fn = File.join(outputdir, "accepted_hits.bam")
desc "mapquant DIST INDEX OUTPUTDIR FASTQS", "map and quantify"
method_option :paired, "convert:bam:sort", :default => false, :desc => 'Are reads paired? If you chose
task.invoke :type => :boolean, [accepted_hits_bam_fn] # call the sorting procedure.
end this option pass just the basename
end of the file without forward/reverse
and .fastq'
def mapquant(dist, index, outputdir, fastqs)
#tophat
invoke :tophat, [dist, index, outputdir, fastqs], :paired=>options.paired
#cufflinks quantification on gtf
invoke :quant, ["#{index}.gtf", File.join(outputdir,"quantification"), File.join(outputdir,"accepted_hits_sort.bam")]
end
…
end
class Rna < Thor
desc "quant GTF OUTPUTDIR BAM ", "Genes and transcripts quantification"
Bio::Ngs::Cufflinks::Quantification.new.thor_task(self, :quant) do |wrapper, task, gtf, outputdir, bam|
wrapper.params = task.options
wrapper.params = {"num-threads" => 6, "output-dir" => outputdir, "GTF" => gtf }
wrapper.run :arguments=>[bam], :separator => "="
end
end
July
15,
2011
BOSC,
Vienna,
Austria](https://image.slidesharecdn.com/raoul-bosc2011-biongs-110802102003-phpapp02/85/D03-NextGen-Bio-NGS-15-320.jpg)