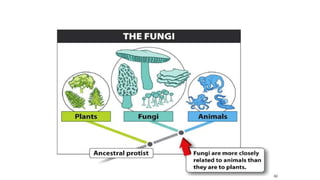
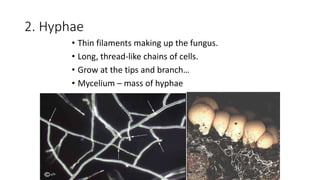
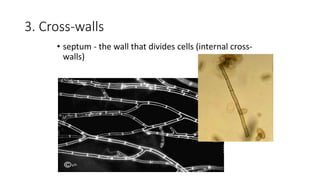
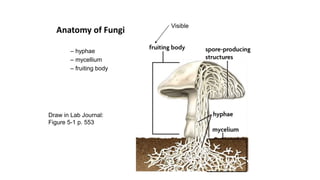
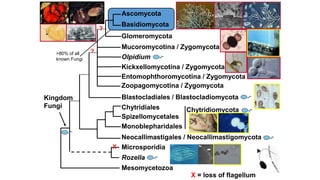
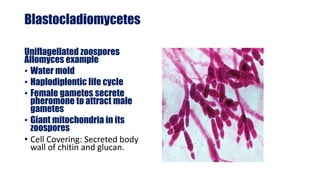
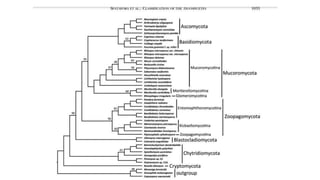
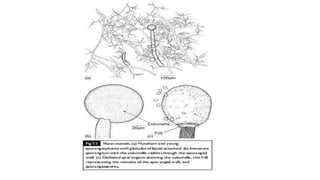
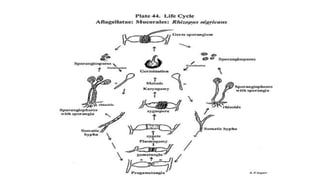
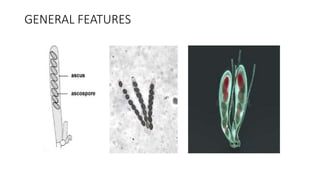
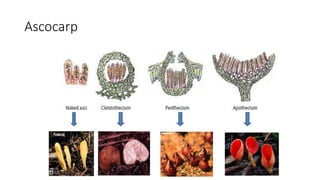
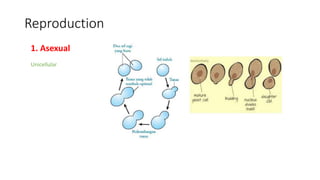
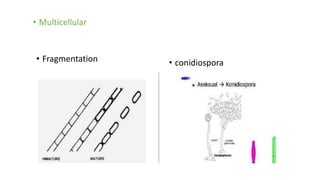
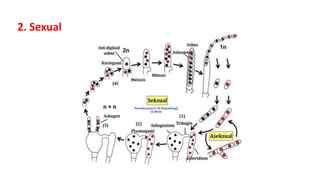
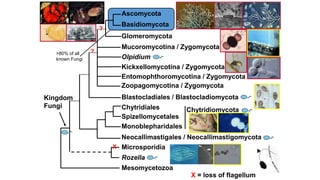
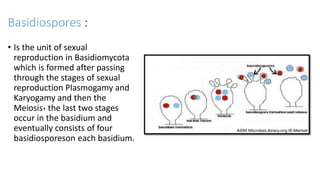
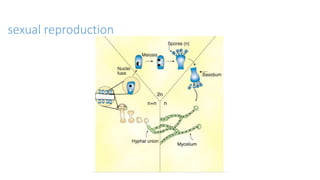
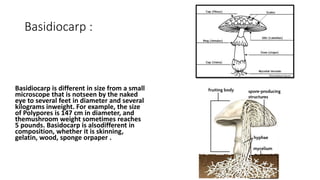
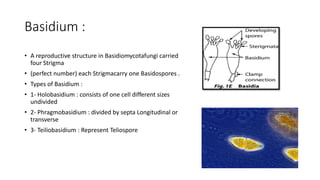
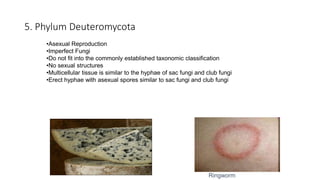
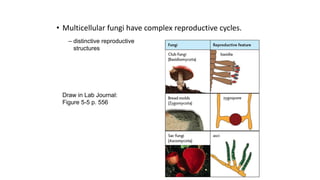
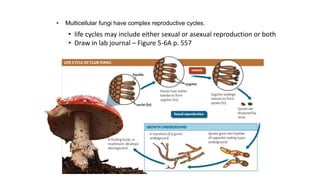
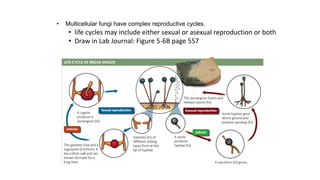
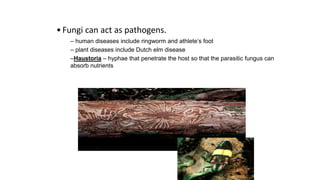
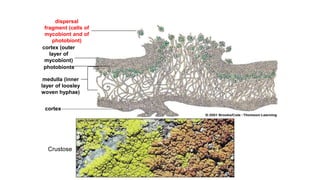
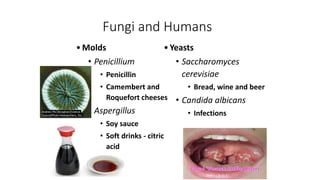
Fungi are a diverse group of organisms that include mushrooms, molds, yeasts, and microfungi. They obtain nutrients by absorbing them from surrounding materials and are important decomposers in ecosystems. Fungi can reproduce both sexually through structures like basidia and ascocarps, and asexually through spores, budding, or fragmentation. They play key roles as decomposers, pathogens, and mutualists with other organisms like plants and algae in lichens.