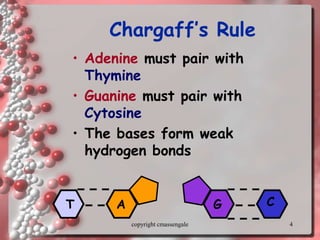
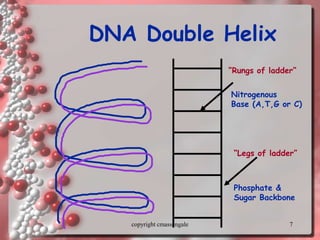
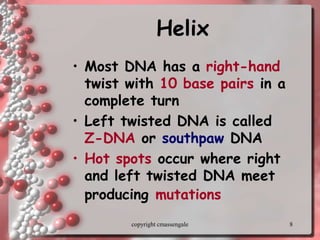
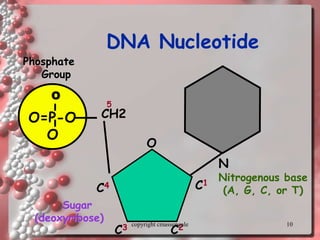
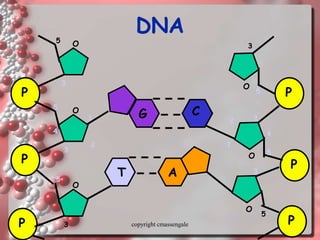
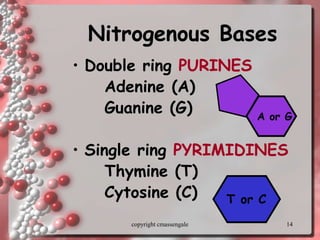
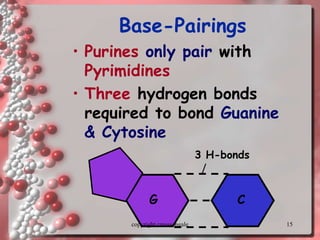
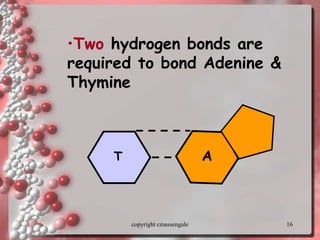
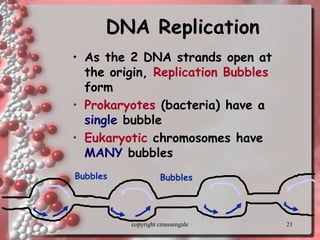
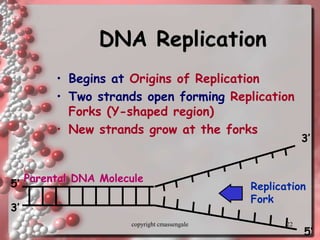
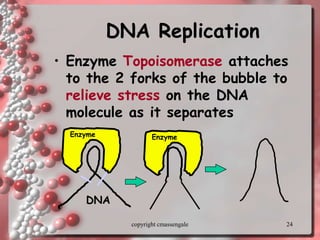
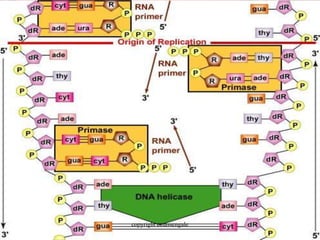
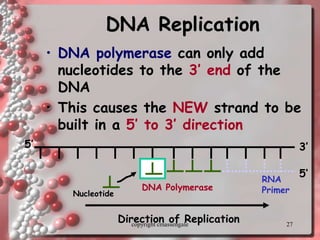
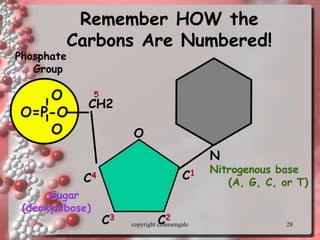
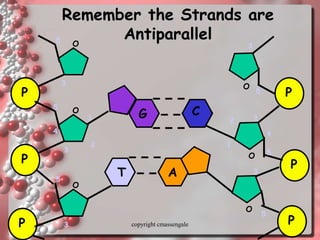
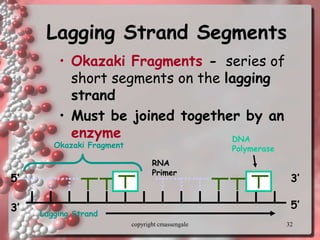
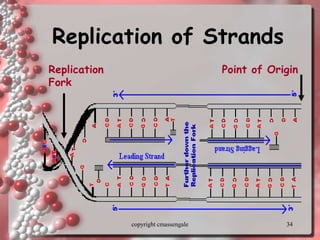
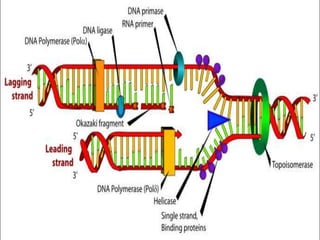
Rosalind Franklin discovered the DNA structure through x-ray crystallography. Watson and Crick then built the first DNA model using Franklin's research. DNA is made of two antiparallel strands coiled into a double helix. Each strand is a backbone of sugar and phosphate groups with nitrogenous bases projecting inward in base pairs - adenine pairs with thymine and cytosine pairs with guanine. DNA replication copies the DNA before cell division, using enzymes like helicase to unwind the strands and DNA polymerase to add complementary nucleotides to each new strand.