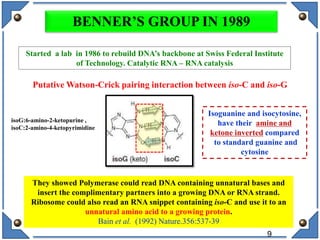
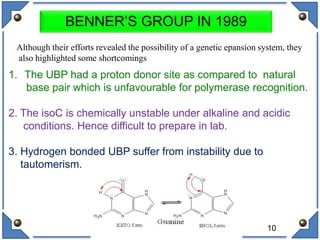
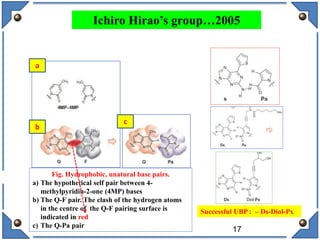
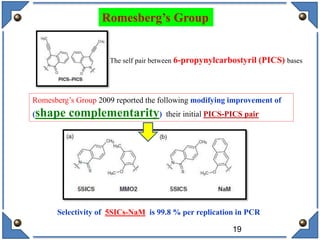
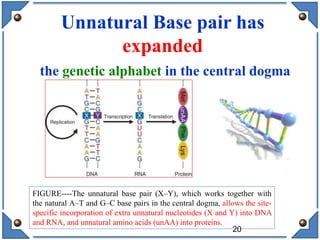
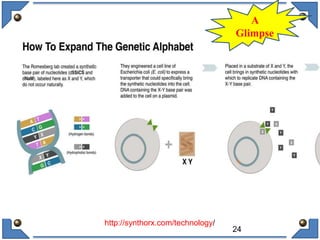
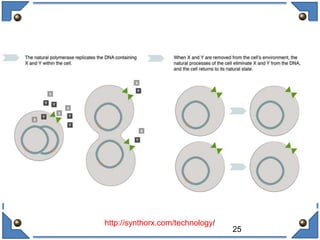
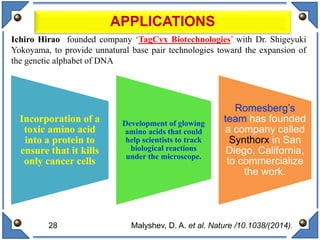
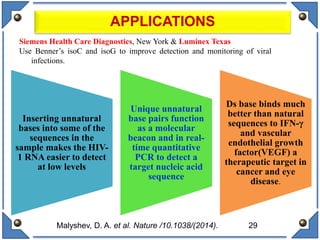
The document discusses the development and implications of unnatural base pairs (UBPs) in DNA, which could expand the genetic code beyond the traditional four bases (A, T, C, G). It highlights key research contributions from various scientists in synthetic biology, including significant breakthroughs in creating semi-synthetic organisms capable of utilizing these expanded genetic alphabets for biotechnology applications. The potential applications of UBPs range from improved detection methods in medical diagnostics to novel genetic technologies, substantiating their importance in future scientific advancements.