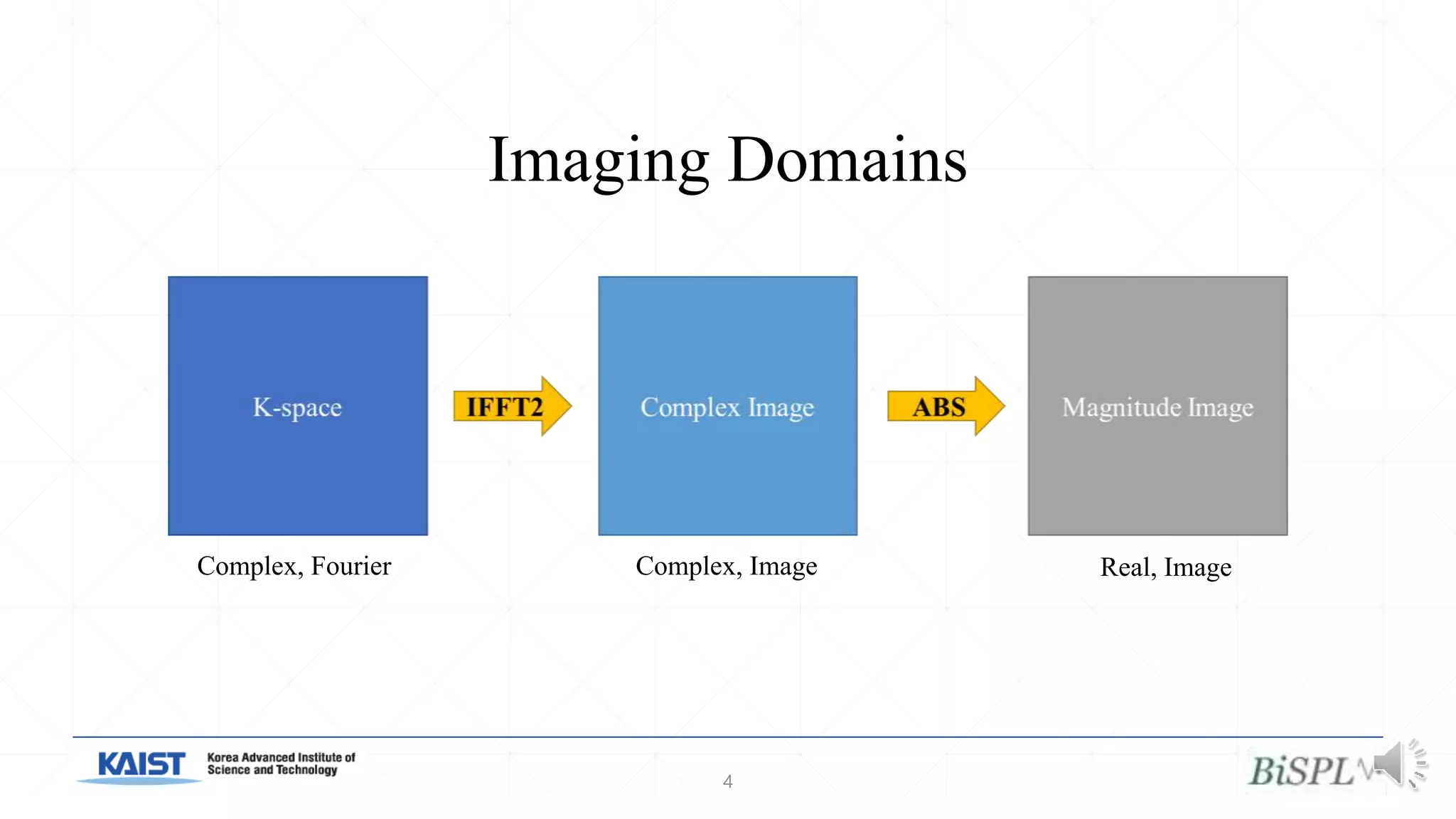
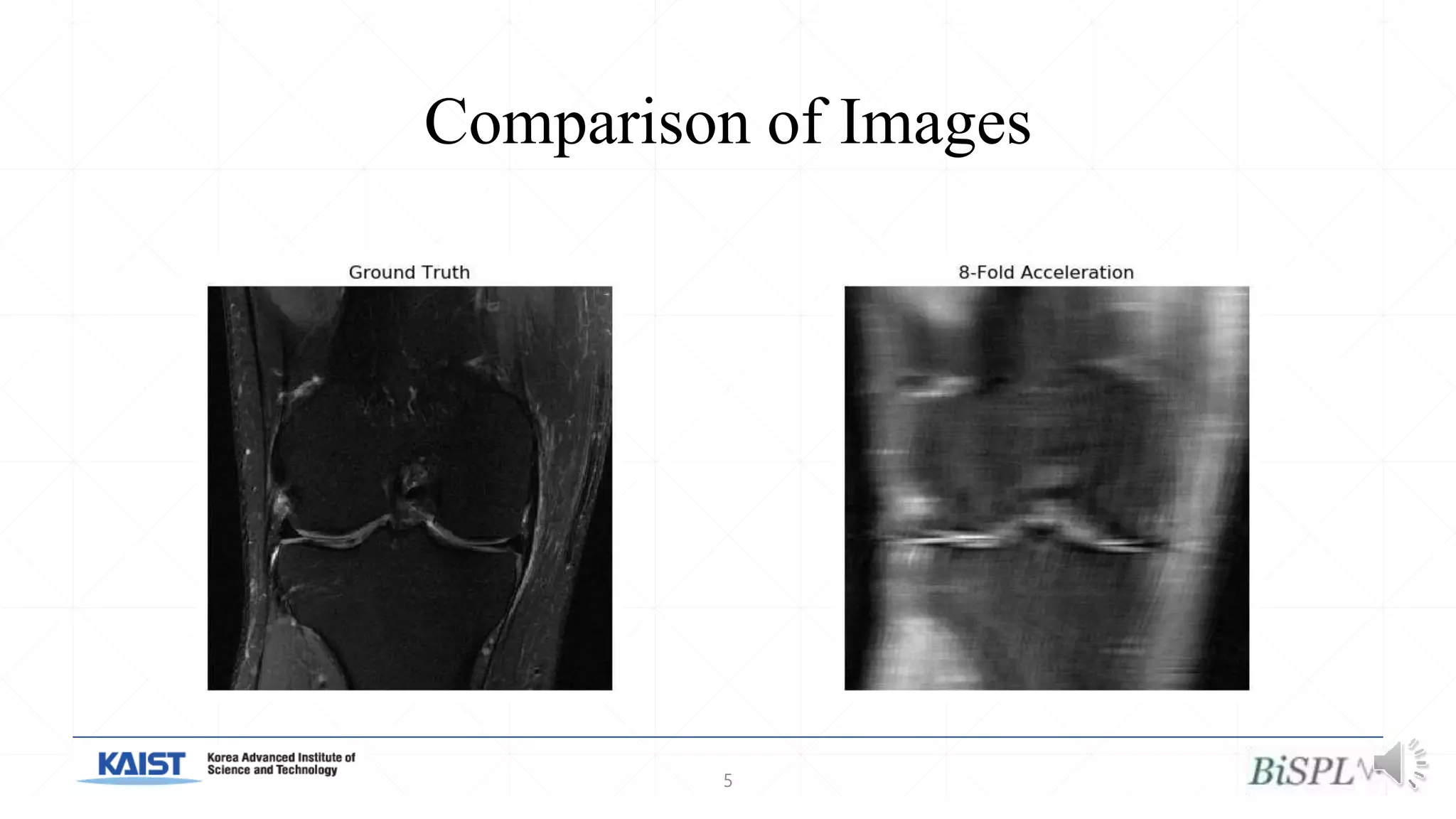
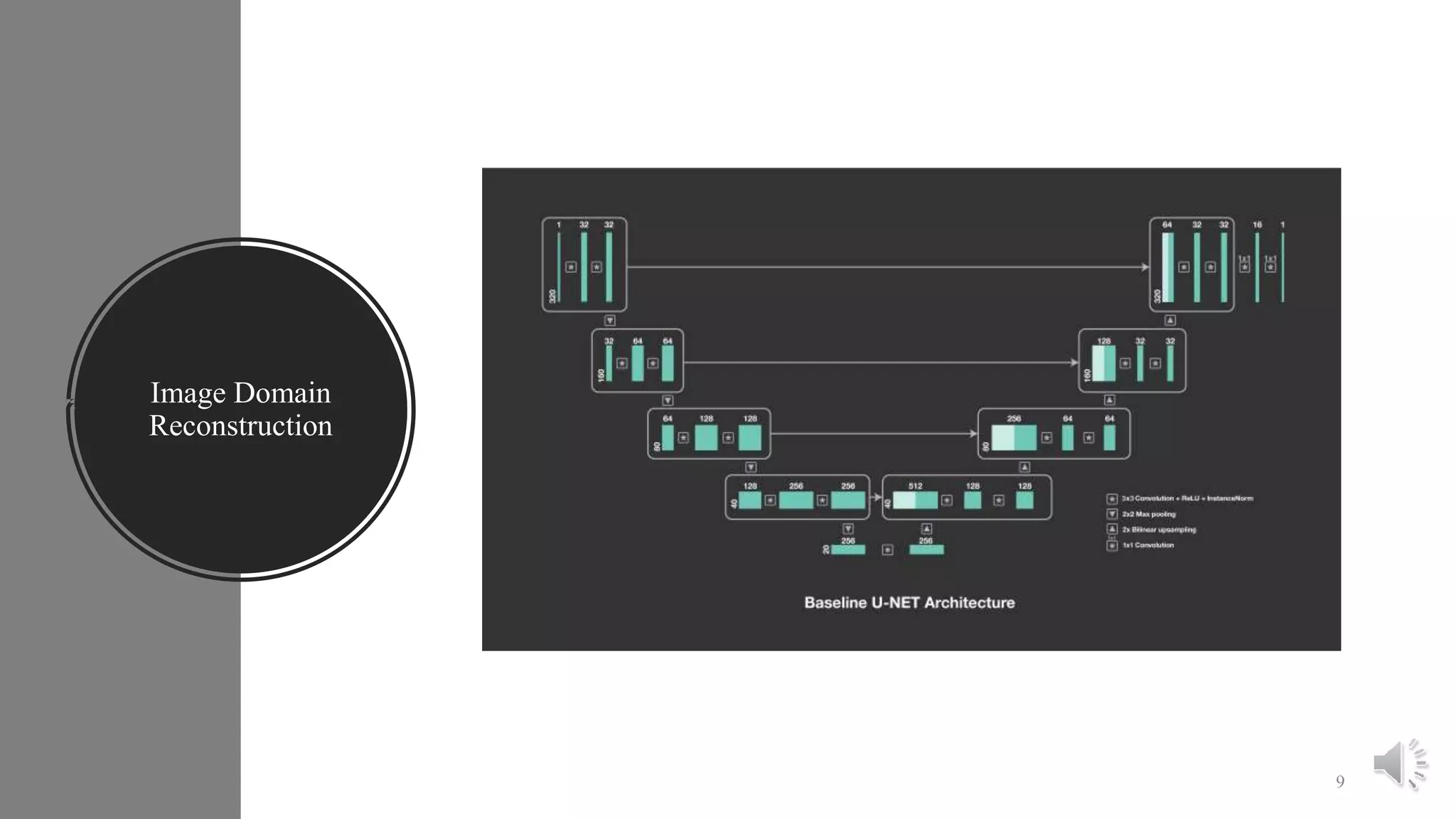
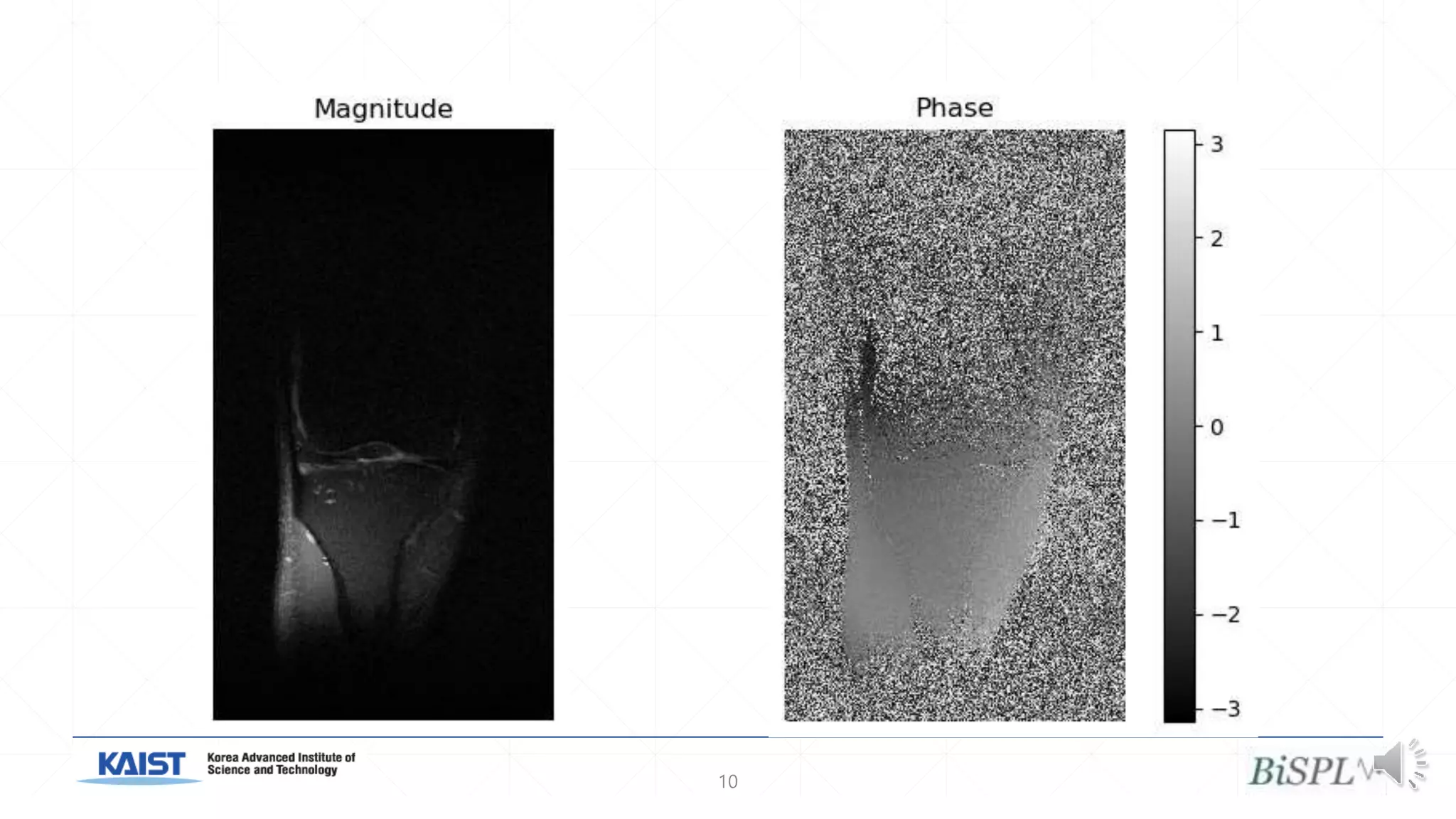
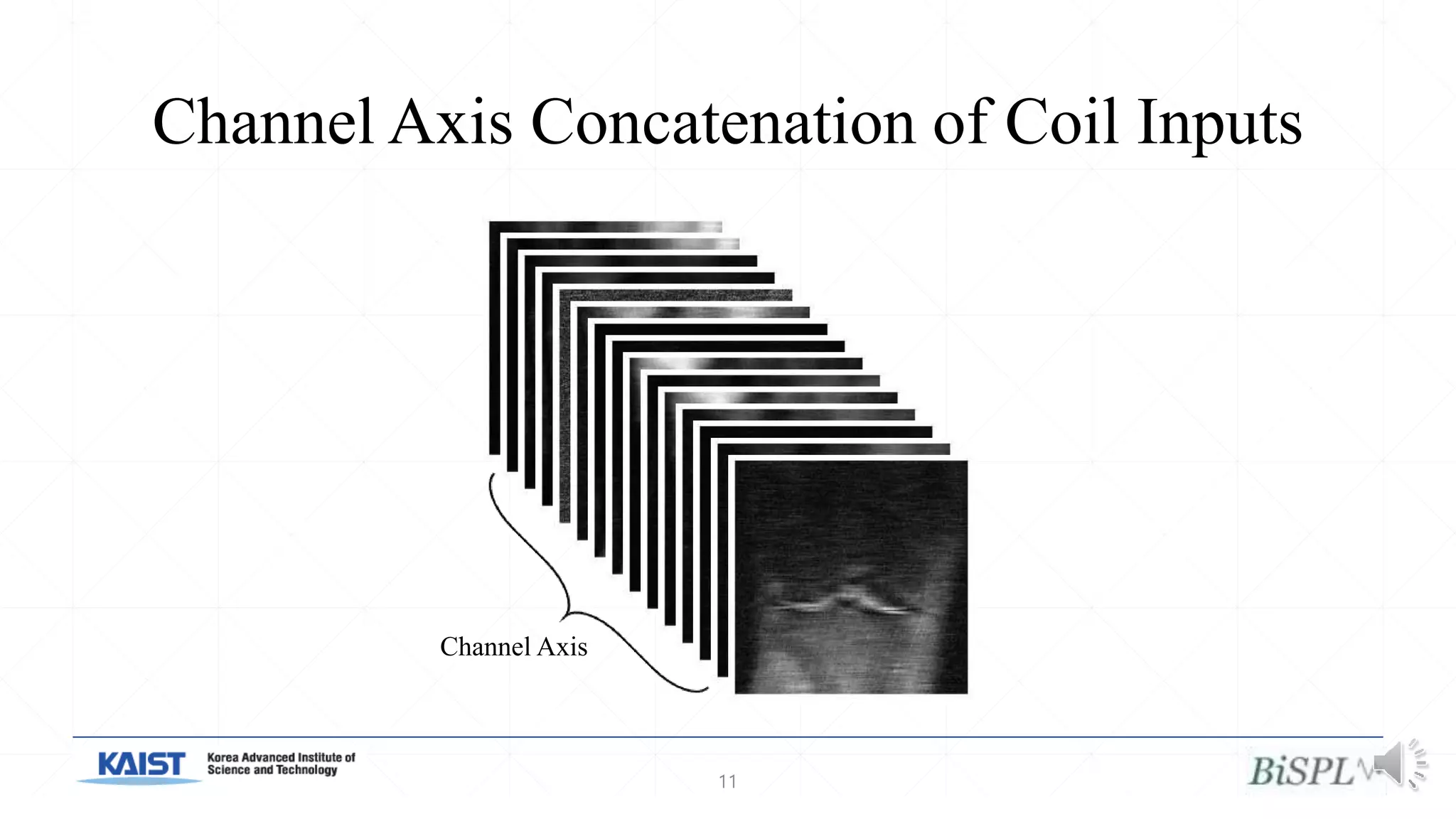
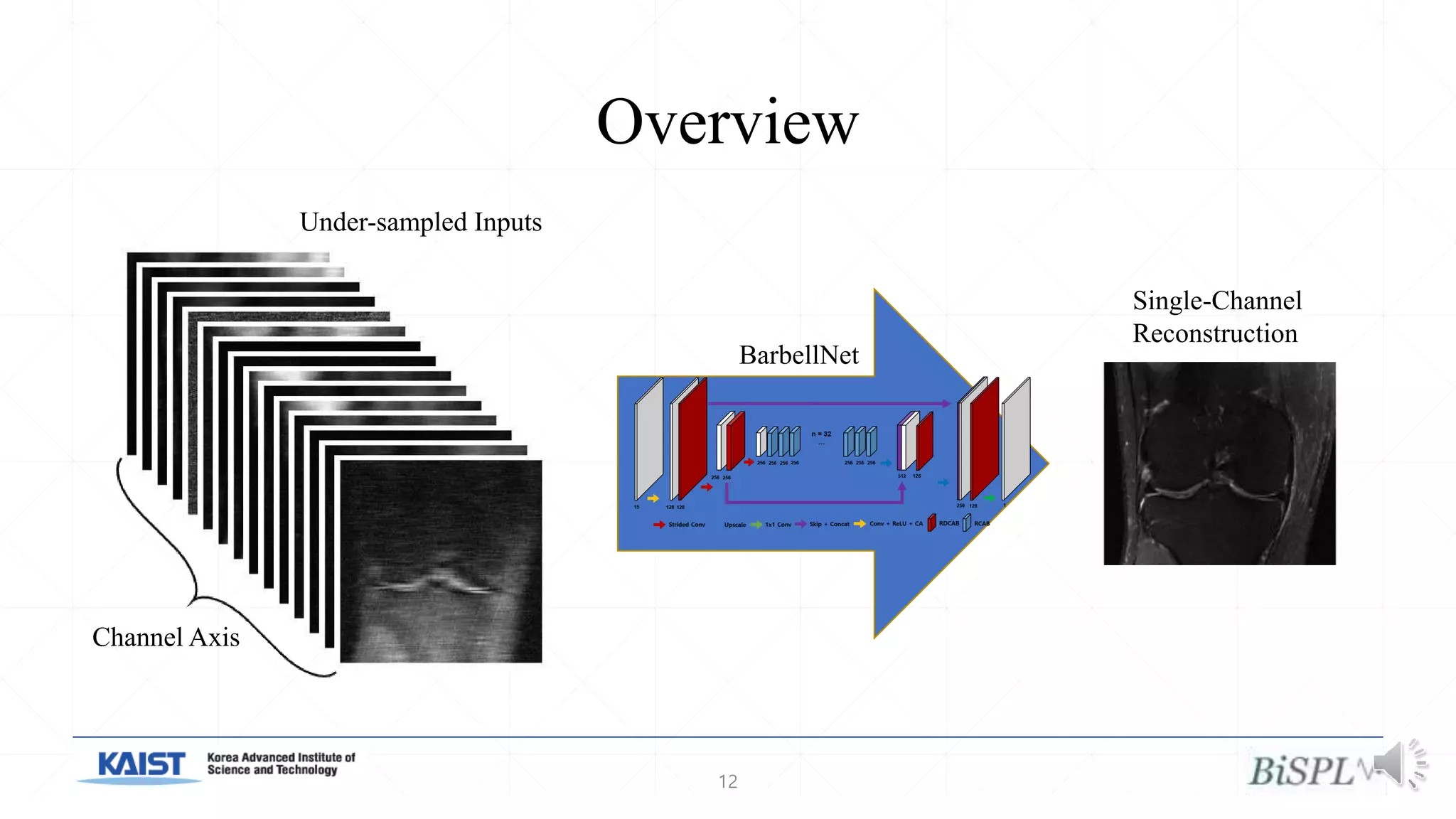
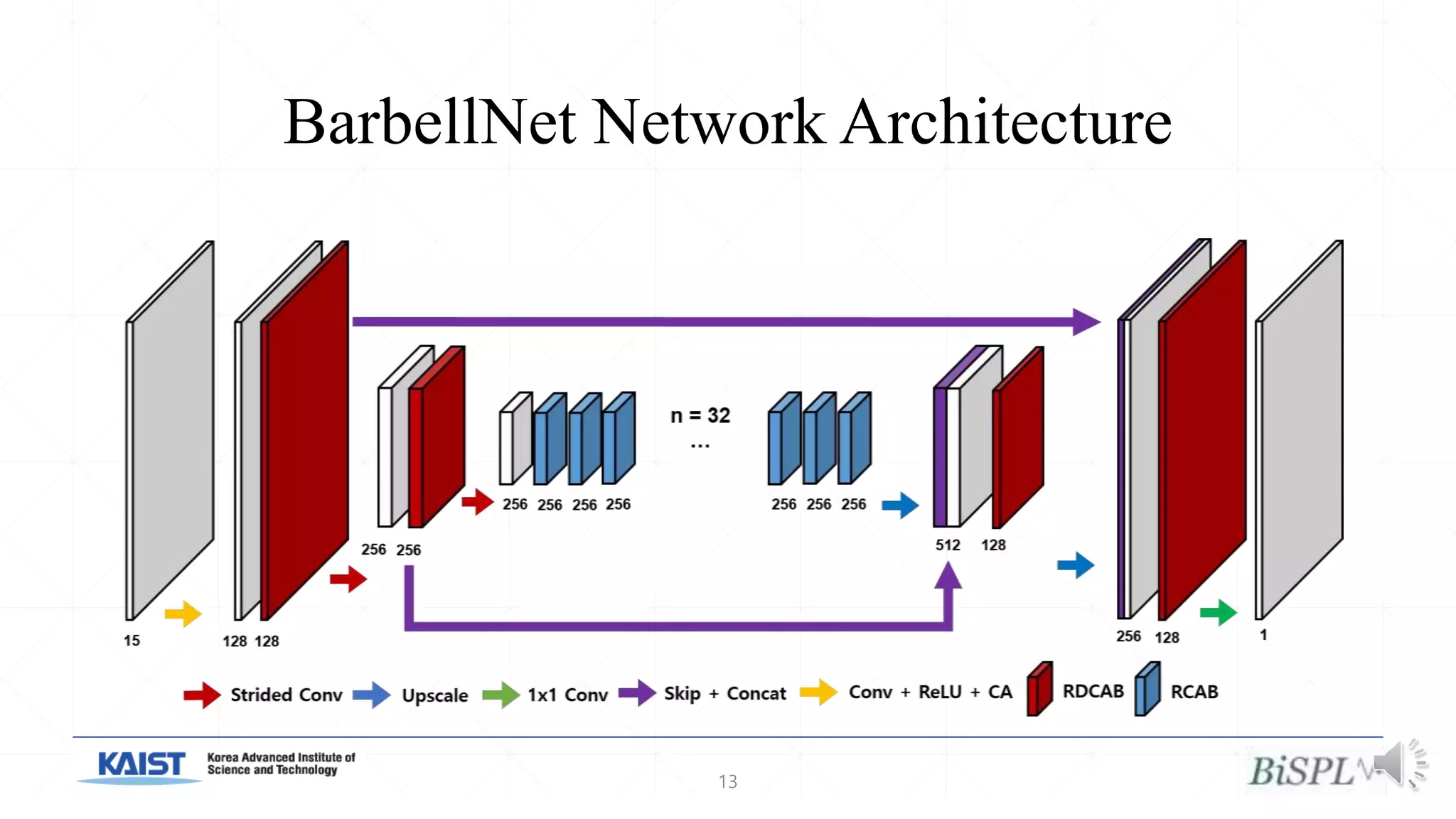
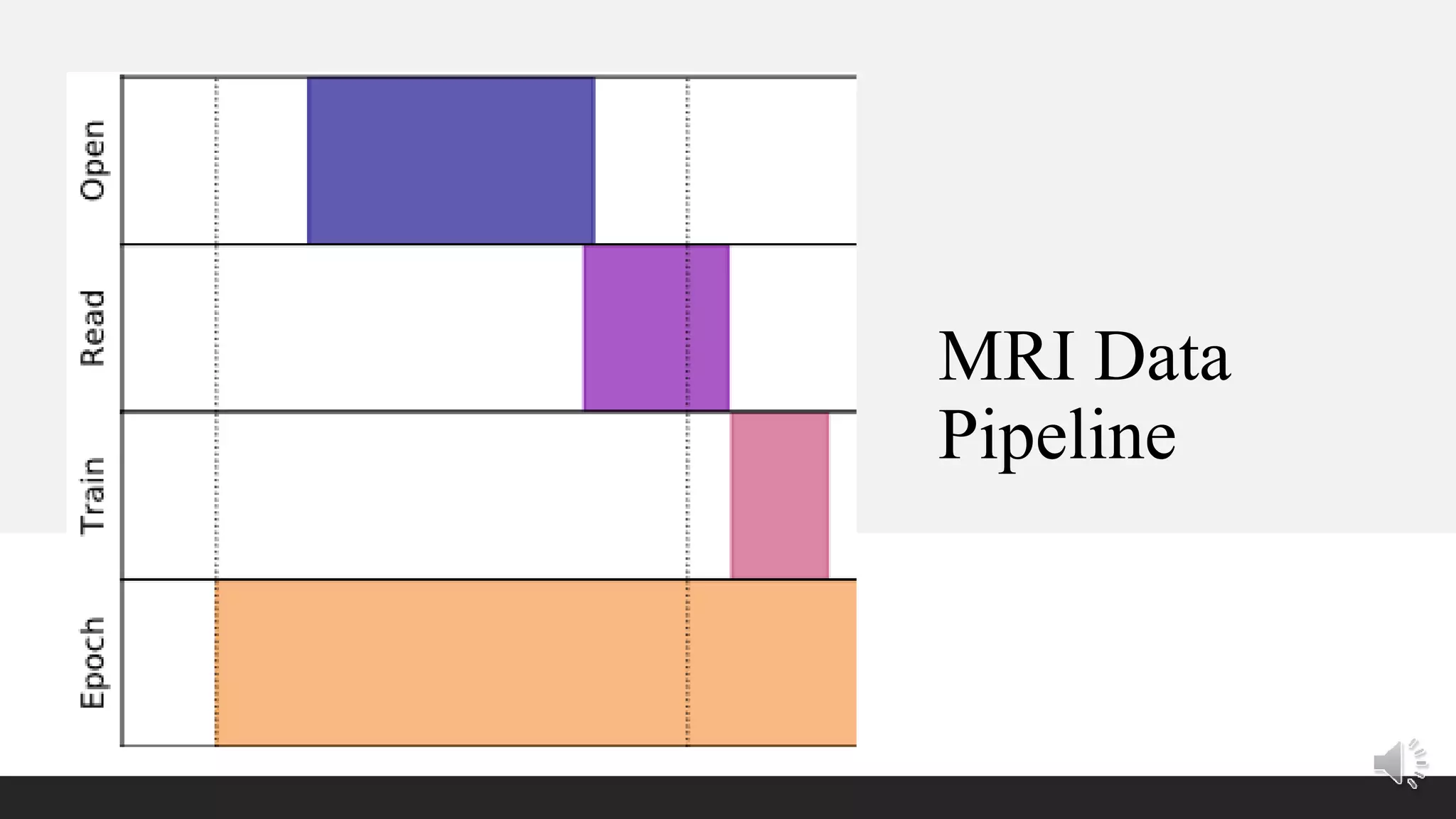
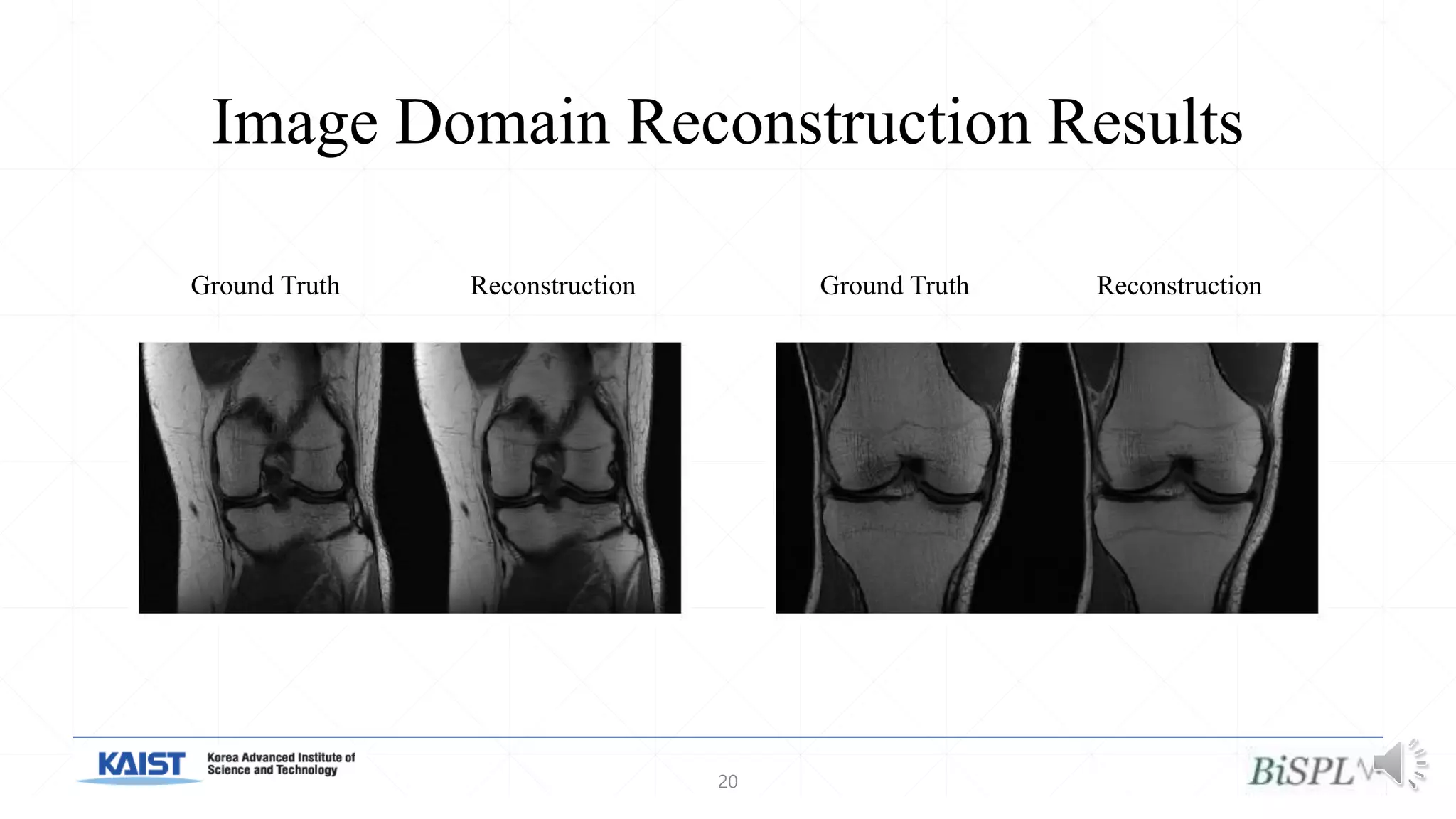
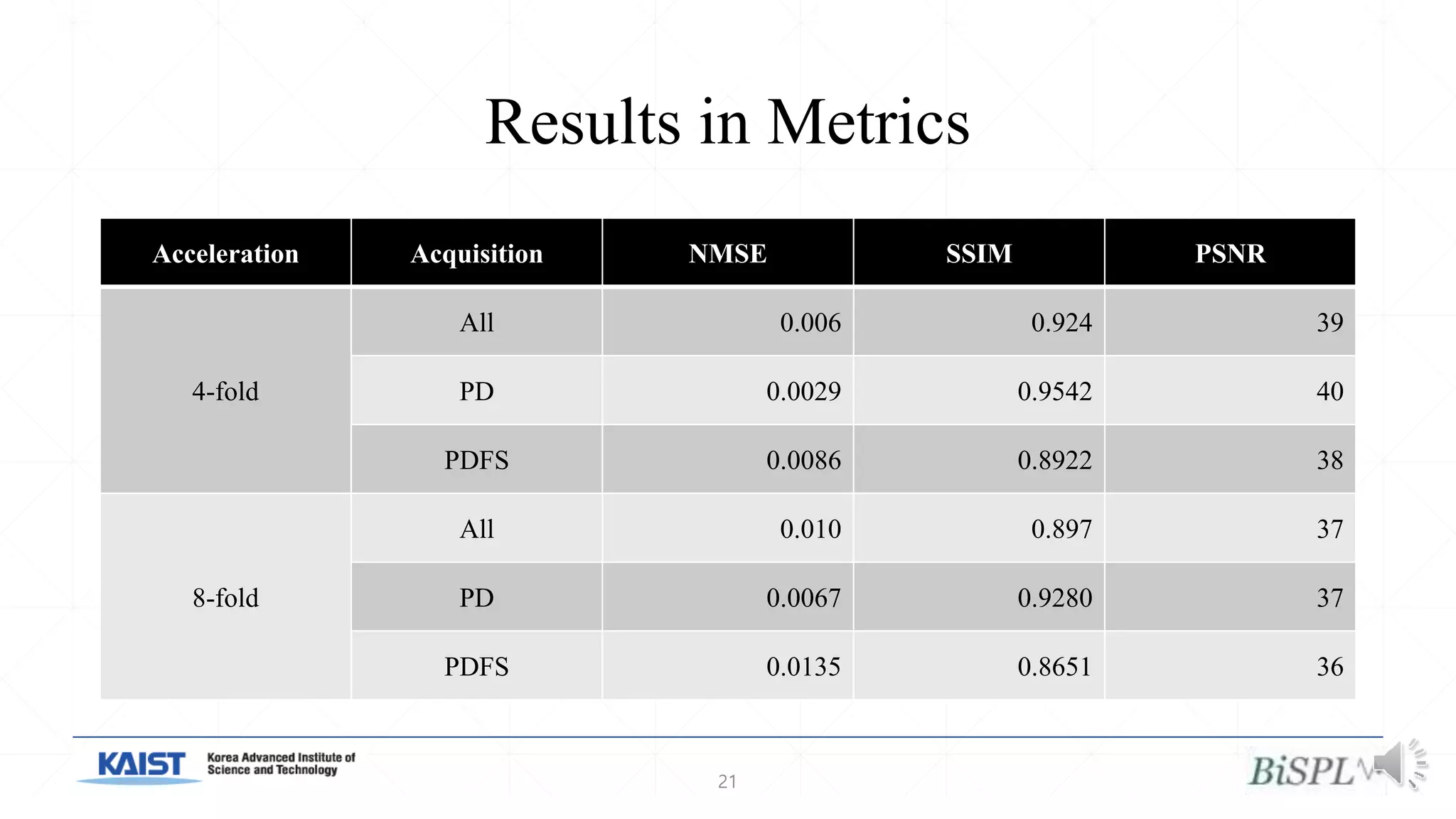
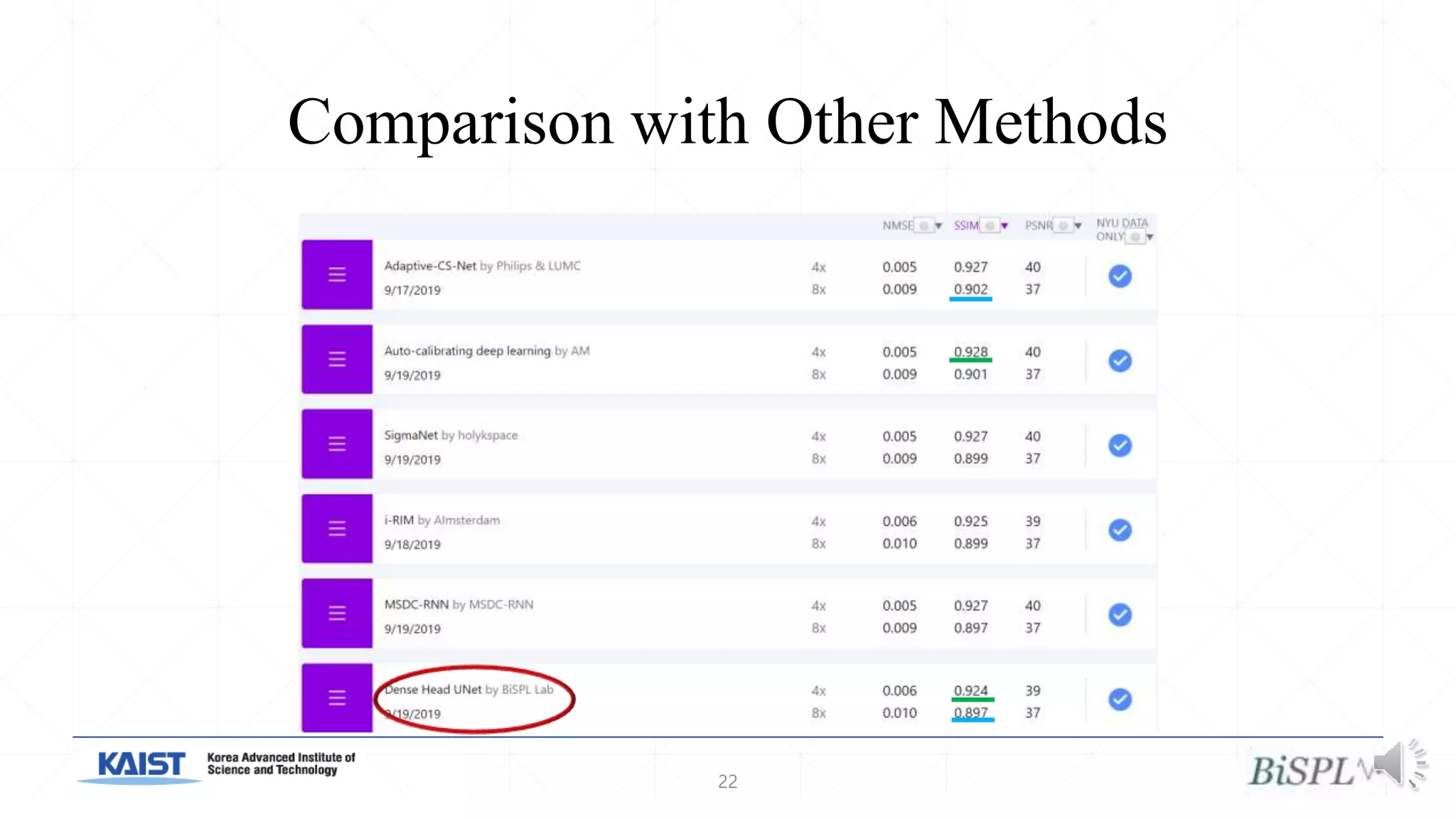
The document presents a method for accelerated parallel MRI using deep learning and discusses the challenges of using k-space data, which comprises complex values. Solutions include concatenating coil inputs, using magnitude images as input, and implementing a specific network architecture for reconstruction. Practical implementation tips and results from image domain reconstruction experiments are also provided, showcasing various performance metrics.