3A-TYL~1.PPT
•Download as PPT, PDF•
0 likes•2 views
Virginia Bioinformatics Workshop
Report
Share
Report
Share
Recommended
Recommended
Course: Bioinformatics for Biomedical Research (2014).
Session: 1.3- Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensembl, Biomart and IGV.
Statistics and Bioinformatisc Unit (UEB) & High Technology Unit (UAT) from Vall d'Hebron Research Institute (www.vhir.org), Barcelona.Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensem...

Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensem...VHIR Vall d’Hebron Institut de Recerca
More Related Content
Similar to 3A-TYL~1.PPT
Course: Bioinformatics for Biomedical Research (2014).
Session: 1.3- Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensembl, Biomart and IGV.
Statistics and Bioinformatisc Unit (UEB) & High Technology Unit (UAT) from Vall d'Hebron Research Institute (www.vhir.org), Barcelona.Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensem...

Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensem...VHIR Vall d’Hebron Institut de Recerca
Similar to 3A-TYL~1.PPT (20)
Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensem...

Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensem...
A Genome Sequence Analysis System Built With Hypertable

A Genome Sequence Analysis System Built With Hypertable
2015 bio it visualizing genomic variants and annotations is vital for accur...

2015 bio it visualizing genomic variants and annotations is vital for accur...
RNA-seq for DE analysis: the biology behind observed changes - part 6

RNA-seq for DE analysis: the biology behind observed changes - part 6
Using VarSeq to Improve Variant Analysis Research Workflows

Using VarSeq to Improve Variant Analysis Research Workflows
Using VarSeq to Improve Variant Analysis Research Workflows

Using VarSeq to Improve Variant Analysis Research Workflows
Recently uploaded
Recently uploaded (20)
development of diagnostic enzyme assay to detect leuser virus

development of diagnostic enzyme assay to detect leuser virus
Use of mutants in understanding seedling development.pptx

Use of mutants in understanding seedling development.pptx
Genome Projects : Human, Rice,Wheat,E coli and Arabidopsis.

Genome Projects : Human, Rice,Wheat,E coli and Arabidopsis.
Cyathodium bryophyte: morphology, anatomy, reproduction etc.

Cyathodium bryophyte: morphology, anatomy, reproduction etc.
Selaginella: features, morphology ,anatomy and reproduction.

Selaginella: features, morphology ,anatomy and reproduction.
TransientOffsetin14CAftertheCarringtonEventRecordedbyPolarTreeRings

TransientOffsetin14CAftertheCarringtonEventRecordedbyPolarTreeRings
Module for Grade 9 for Asynchronous/Distance learning

Module for Grade 9 for Asynchronous/Distance learning
Pteris : features, anatomy, morphology and lifecycle

Pteris : features, anatomy, morphology and lifecycle
Bhiwandi Bhiwandi ❤CALL GIRL 7870993772 ❤CALL GIRLS ESCORT SERVICE In Bhiwan...

Bhiwandi Bhiwandi ❤CALL GIRL 7870993772 ❤CALL GIRLS ESCORT SERVICE In Bhiwan...
FAIRSpectra - Enabling the FAIRification of Analytical Science

FAIRSpectra - Enabling the FAIRification of Analytical Science
Understanding Partial Differential Equations: Types and Solution Methods

Understanding Partial Differential Equations: Types and Solution Methods
Cot curve, melting temperature, unique and repetitive DNA

Cot curve, melting temperature, unique and repetitive DNA
Porella : features, morphology, anatomy, reproduction etc.

Porella : features, morphology, anatomy, reproduction etc.
Site specific recombination and transposition.........pdf

Site specific recombination and transposition.........pdf
3A-TYL~1.PPT
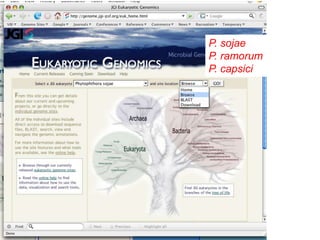
- 1. P. sojae P. ramorum P. capsici
- 2. Detailed manual in use of JGI site
- 3. Tutorial in use of JGI site - includes guide to structural and functional annotation (not enabled in P. sojae and P. ramorum databases but info is generally useful)
- 4. Genome Browser Can type or paste text in here Then type return or click “Refresh” Change orientation of sequence Display 3 frame translation (must be zoomed in) Display DNA sequence (must be zoomed in) Sequence similarity to P. ramorum Blue = coding Pink = non-coding EVERYTHING IS CLICKABLE!! Click this to access the DNA sequence Tracks give different versions of gene models as well as various sets of Blast hits Clicking on a track will expand it (see next page) Current “best” gene models
- 5. Expanded track Red models have been manually annotated Clicking on open models will open the protein page
- 6. Clicking on model will open the nucleic acid sequence Clicking on green bar will open the protein sequence introns Clicking names will open gene in its database Clicking gray bar will open sequence alignement red blocks are insertions in target Clicking red dot will set sequence as source Protein page EVERYTHING IS CLICKABLE!! Clickable All clickable Blue indicates EST support for UTR
- 7. Shift click or command-click to choose lists of sequences to get (in fasta format) or do Clustal alignments on Shift click or command-click to choose lists of genomes and/or databases for the list of alignments. Then click “Apply Filter” number of alignments to display Protein page OPTIONS
- 8. Sequence Display FROM PROTEIN PAGE Padding controls how much flanking sequence is displayed Introns (should start with GTxxG) Blue indicates EST support for UTR
- 9. Advanced Search (much easier than Simple Search) Shift click or command-click to search mutilple databses simulataneously It’s also an easy way to switch databases Do NOT type return or enter after your entry - you MUST click “Search” Accepts and recognizes almost any kind of query Output of search
- 10. Links: P -> protein page T -> transcript page G -> genome browser Check to download sequences
- 11. Transcript page Contains more info on annotation Used mostly for manual annotation
- 12. Gene Ontology Click to go to protein page
- 13. KEGG Click to go to protein page
