Kshama Dey OIST
•Download as PPTX, PDF•
0 likes•6 views
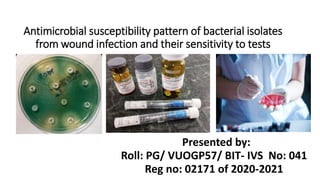
Antimicrobial susceptibility pattern of bacterial isolates from wound infection and their sensitivity to tests
Report
Share
Report
Share
Recommended
Bacteriological profile and antibiogram of aerobic burn wound isolates in a t...

Bacteriological profile and antibiogram of aerobic burn wound isolates in a t...Dr Muktikesh Dash, MD, PGDFM
Recommended
Bacteriological profile and antibiogram of aerobic burn wound isolates in a t...

Bacteriological profile and antibiogram of aerobic burn wound isolates in a t...Dr Muktikesh Dash, MD, PGDFM
A total number of 74 coagulase negative Staphylococci were isolated from orthopaedic patients in Ahmadu Bello University Teaching Hospital, Zaria, Nigeria. They were further characterized into various Staphylococci species using API STAPH identification kit: Staph xylosus (31.1%), Staph lentus (10.8%), Staph hominis (10.8%), Staph cohnii cohnii (5.4%), Staph epidermidis (4.1%) others were Staph cohnii ureal., Staph hyicus, Staph lugdunensis (2.7% each) Staph caprae , Staph capitis, Staph haemolyticus, Staph scuiri, Staph chromogenes and Staph warneri (1.4% each). Microcossus spp was 8.2% while 13.5% isolates were undetermined. Kirby Baurer disk method was used for the antibiotics susceptibility test, the result showed gentamicin and ciprofloxacin to be most active (96.6%), followed by vancomycin (93.1) and pefloxacin (87.9). The isolates were resistant to ampicillin (96.6), amoxicillin clavulanic acid (65.5%), clindamycin 41.4%). The aim of this study is to classify the coagulase negative Staphylococci isolates into species and to determine their antibiotic susceptibilityANTIBIOTICS SUSCEPTIBILITY PATTERN OF COAGULASE NEGATIVE STAPHYLOCOCCI ISOLAT...

ANTIBIOTICS SUSCEPTIBILITY PATTERN OF COAGULASE NEGATIVE STAPHYLOCOCCI ISOLAT...International Educational Applied Scientific Research Journal (IEASRJ)
More Related Content
Similar to Kshama Dey OIST
A total number of 74 coagulase negative Staphylococci were isolated from orthopaedic patients in Ahmadu Bello University Teaching Hospital, Zaria, Nigeria. They were further characterized into various Staphylococci species using API STAPH identification kit: Staph xylosus (31.1%), Staph lentus (10.8%), Staph hominis (10.8%), Staph cohnii cohnii (5.4%), Staph epidermidis (4.1%) others were Staph cohnii ureal., Staph hyicus, Staph lugdunensis (2.7% each) Staph caprae , Staph capitis, Staph haemolyticus, Staph scuiri, Staph chromogenes and Staph warneri (1.4% each). Microcossus spp was 8.2% while 13.5% isolates were undetermined. Kirby Baurer disk method was used for the antibiotics susceptibility test, the result showed gentamicin and ciprofloxacin to be most active (96.6%), followed by vancomycin (93.1) and pefloxacin (87.9). The isolates were resistant to ampicillin (96.6), amoxicillin clavulanic acid (65.5%), clindamycin 41.4%). The aim of this study is to classify the coagulase negative Staphylococci isolates into species and to determine their antibiotic susceptibilityANTIBIOTICS SUSCEPTIBILITY PATTERN OF COAGULASE NEGATIVE STAPHYLOCOCCI ISOLAT...

ANTIBIOTICS SUSCEPTIBILITY PATTERN OF COAGULASE NEGATIVE STAPHYLOCOCCI ISOLAT...International Educational Applied Scientific Research Journal (IEASRJ)
Similar to Kshama Dey OIST (20)
Alan Lesniewicz Memorial Lecture at UIC - July 2015

Alan Lesniewicz Memorial Lecture at UIC - July 2015
Prevalence and antibiotic susceptibility pattern of staphylococcus aureus in ...

Prevalence and antibiotic susceptibility pattern of staphylococcus aureus in ...
Bacteriological and Mycological Profile of Chronic Suppurative Otitis Media I...

Bacteriological and Mycological Profile of Chronic Suppurative Otitis Media I...
Emergence of tegicycline resistance in multidrug resistant (mdr) organisms i...

Emergence of tegicycline resistance in multidrug resistant (mdr) organisms i...
Isolation, Characterization, and Antibiotics Resistance Profile of Staphyloco...

Isolation, Characterization, and Antibiotics Resistance Profile of Staphyloco...
Antimicrobial Susceptibility Profile of Escherichia Coli Isolates from Urine ...

Antimicrobial Susceptibility Profile of Escherichia Coli Isolates from Urine ...
ANTIBIOTICS SUSCEPTIBILITY PATTERN OF COAGULASE NEGATIVE STAPHYLOCOCCI ISOLAT...

ANTIBIOTICS SUSCEPTIBILITY PATTERN OF COAGULASE NEGATIVE STAPHYLOCOCCI ISOLAT...
Multidrug Resistance Pattern of Staphylococcus Aureus Isolates in Maiduguri M...

Multidrug Resistance Pattern of Staphylococcus Aureus Isolates in Maiduguri M...
Multidrug Resistance Pattern of Staphylococcus Aureus Isolates in Maiduguri ...

Multidrug Resistance Pattern of Staphylococcus Aureus Isolates in Maiduguri ...
Detection of Integrons in Multidrug Resistant Wound Isolates

Detection of Integrons in Multidrug Resistant Wound Isolates
Trends in Antibiotic Resistance of Vibrio Cholerae Isolates in Kenya (2006 - ...

Trends in Antibiotic Resistance of Vibrio Cholerae Isolates in Kenya (2006 - ...
IOSR Journal of Pharmacy (IOSRPHR), www.iosrphr.org, call for paper, research...

IOSR Journal of Pharmacy (IOSRPHR), www.iosrphr.org, call for paper, research...
Occurrence and antibiotic susceptibility of Streptococcus pyogenes isolated f...

Occurrence and antibiotic susceptibility of Streptococcus pyogenes isolated f...
International Journal of Pharmaceutical Science Invention (IJPSI)

International Journal of Pharmaceutical Science Invention (IJPSI)
More from Arnim Banerjee
More from Arnim Banerjee (14)
Recently uploaded
Recently uploaded (20)
SAMASTIPUR CALL GIRL 7857803690 LOW PRICE ESCORT SERVICE

SAMASTIPUR CALL GIRL 7857803690 LOW PRICE ESCORT SERVICE
9999266834 Call Girls In Noida Sector 22 (Delhi) Call Girl Service

9999266834 Call Girls In Noida Sector 22 (Delhi) Call Girl Service
Hire 💕 9907093804 Hooghly Call Girls Service Call Girls Agency

Hire 💕 9907093804 Hooghly Call Girls Service Call Girls Agency
GUIDELINES ON SIMILAR BIOLOGICS Regulatory Requirements for Marketing Authori...

GUIDELINES ON SIMILAR BIOLOGICS Regulatory Requirements for Marketing Authori...
TEST BANK For Radiologic Science for Technologists, 12th Edition by Stewart C...

TEST BANK For Radiologic Science for Technologists, 12th Edition by Stewart C...
Call Girls Alandi Call Me 7737669865 Budget Friendly No Advance Booking

Call Girls Alandi Call Me 7737669865 Budget Friendly No Advance Booking
Feature-aligned N-BEATS with Sinkhorn divergence (ICLR '24)

Feature-aligned N-BEATS with Sinkhorn divergence (ICLR '24)
Formation of low mass protostars and their circumstellar disks

Formation of low mass protostars and their circumstellar disks
Chemical Tests; flame test, positive and negative ions test Edexcel Internati...

Chemical Tests; flame test, positive and negative ions test Edexcel Internati...
Connaught Place, Delhi Call girls :8448380779 Model Escorts | 100% verified

Connaught Place, Delhi Call girls :8448380779 Model Escorts | 100% verified
Discovery of an Accretion Streamer and a Slow Wide-angle Outflow around FUOri...

Discovery of an Accretion Streamer and a Slow Wide-angle Outflow around FUOri...
FAIRSpectra - Enabling the FAIRification of Spectroscopy and Spectrometry

FAIRSpectra - Enabling the FAIRification of Spectroscopy and Spectrometry
Vip profile Call Girls In Lonavala 9748763073 For Genuine Sex Service At Just...

Vip profile Call Girls In Lonavala 9748763073 For Genuine Sex Service At Just...
Kshama Dey OIST
- 1. Antimicrobial susceptibility pattern of bacterial isolates from wound infection and their sensitivity to tests Presented by: Roll: PG/ VUOGP57/ BIT- IVS No: 041 Reg no: 02171 of 2020-2021
- 2. Introduction The skin serves as a defence barrier against pathogen invasion. As a result a wound is formed when the normal anatomical structures is disrupted by a accidents or by surgical operation, chemical, physical, mechanical or thermal events, resulting in a change in skin function. Skin is exposed to injuries scratches, thus it is more susceptible to colonization by pathogen. Wound infection can be caused by a single pathogen (mono-microbial) or by multiple pathogen (poly- microbial). Any wound is at some risk of becoming infected. Infected wounds are likely to be more painful, hypersensitive, and odorous resulting in amplified uneasiness and discomfort for the patient. Wound infection has a significant impact on patient morbidity and mortality. Wound infection can be caused by a variety of pathogens including Bacteria, Viruses, and Fungal parasites. Gram positive cocci like Staphylococcus aureus , Streptococcus sp, are the most Common bacteria found in infected wounds. Gram negative bacteria such as Pseudomonas aeruginosa, Klebsiella pneumoniae, Proteus sp, Escherichia coli, and Enterobacter sp are the most Common bacteria found in infected wounds. Due to the widespread proliferation of antibiotic resistance bacteria, wound care has become more difficult. So, appropriate antibiotics selected by antimicrobial sensitivity testing have great importance.
- 3. Aim of the study • Isolate and identify the bacteria responsible for wound infection. • The determination of Antibiotic Susceptibility Pattern of Bacteria.
- 4. Classification of wounds Wounds can be classified in several ways like depending on the healing time,depth of wound, level of contamination etc. Classification of Wounds Healing time Depth of wound Based on contamination of wound Acute Chronic Partial Thickness Clean Wound Superficial Full Thickness infected wound Colonized wound Contaminated wound
- 5. Materials and methods Sample inoculated in growth medium Incubated at 37°c for overnight. Plates were observed for colonies appearance. Gram staining Biochemical tests i) For Gram positive cocci (GPC) Catalase and coagulase tests were done routinely ii) For Gram negative bacilli (GNB) , common biochemical tests done are abbreviated as ICUT. A) Indole test B) Citrate test C) Urease test D) Triple sugar iron test ( TSI)
- 6. Antibiotic susceptibility test For AST, were prepared lawn culture with test organisms On Mueller Hinton agar plate. The susceptibility of bacterial isolates against different antibiotics were tested by the modified kirby Bauer disc diffusion method.Amoxyclav, Amikacin, Cefepime, Ceftazidime, Ceftriaxone,Ciprofloxin, CoTrimoxazole, Doxycycline, Gentamicin, Levofloxacin, Linezolid, Piperacillin/Tazobactam, Vancomycin were used in this study for Antibiotic Susceptibility tests. Antibiotic discs were placed on agar plate by sterile Forceps . Measuring the clear of zone by ruler and classified as sensitive or resistant according to the standard CLSI table.
- 7. Results A total numbers of 70 wound samples received at Microbiology Department. Out of 70 samples, 50(71.42%) samples showed positive Bacterial growth and 20( 28.58%) samples showed no growth. There were 37 Females and 33 Males, age ranged from 15 to 80 year. Among 50 positive samples of wound infection, 42( 84%) were Gram negative bacilli and 8(16%) were Gram positive cocci and their 27 were Females and 23 were Males. Age: There were greater incidence of wound infection in the 46-55 year age group. Sex: According to the overall results analysis, Females were more infected than Males. Gram negative bacilli (GNB) were the most prevalent bacteria isolated from the wound swabs. Pseudomonas aeruginosa isolated higher percentage (28%) followed by Klebsiella pneumoniae (24%), Escherichia coli (20%), others Gram-negative organisms (12%). Staphylococcus aureus (16%) was the only Gram positive organism isolated in this study. Pseudomonas aeruginosa, 28% Klebsiella pneumoniae, 24% Escherichia coli., 20% Staphylococcus aureus, 16% Other organisms, 12% Age Group Total No. of Positive Swab No. of Positive Swab Female Male 15-25 8 5 3 26-35 8 5 3 36-45 9 5 4 46-55 15 6 9 56-65 3 1 2 >65 7 5 2 Total 50 27 23
- 8. Antibiotic susceptibility pattern of Pseudomonas aeruginosa Gram negative pseudomonas aeruginosa was the predominant organism responsible for wound infection. This organism was highly sensitive to Piperacillin-Tazobactum (85.71%) followed by Gentamicin (78.57%), Ceftazidime (64.28%), Cefepime (57.14%). Where as it was totally resistant to Amoxyclav & Levofloxacin. 0 10 20 30 40 50 60 70 80 90 100 AMC PIT LE CAZ CPM GEN Sensitive Resistant No. of Organism (%) RXN No. & (%) of Sensitivity and Resistant to Various Antibiotics AMC PIT LE CAZ CPM GEN 14(28%) S - 12 (85.71) - 9 (64.28) 8 (57.14) 11 (78.57) R 14 (100) 2 (14.29) 14 (100) 5 (35.72) 6 (42.86) 3 (21.43)
- 9. Antibiotic Susceptibility Pattern of Klebsiella pneumoniae, Escherichia coli and other gram negative organisms. Antibiotic Susceptibility Pattern of Staphylococcus aureus. No. of Organism (%) RXN No (%) of Sensitivity and Resistant to Various Antibiotics AMC VA GEN LZ CIP DO 8(16%) S 2 (25) 7 (87.5) 6 (75) 7 (87.5) 3 (37.5) 6 (75) R 6 (75) 1 (12.5) 2 (25) 1 (12.5) 5 (62.5) 2 (25) No. of Organism (%) RXN No (%) of Sensitivity and Resistant to Various Antibiotics AMC PIT LE AK COT CTR 12(24%) S 4 (33.33) 10 (83.34) 9 (75) 11 (91.66) 8 (66.66) 2 (16.66) R 8 (66.67) 2 (16.66) 3 (25) 1 (8.34) 4 (33.34) 10 (83.34) 10(20%) S 7 (70) 10 (100) 3 (30) 8 (80) 6 (60) 1 (10) R 3 (30) - 7 (70) 2 (20) 4 (40) 9 (90) 6(12%) S - 4 (66.66) 3 (50) 5 (83.33) 4 (66.66) - R 6 (100) 2 (33.34) 3 (50) 1 (16.67) 2 (33.34) 6 (100)
- 10. Discussion The knowledge of bacteriology of an infection and Laboratory susceptibility testing of microorganisms could make drug selection in antimicrobial chemotherapy more rational. Wound infection is one of the most common and serious complication among hospital acquired infection. Wound infection can increase the length of hospital stay, delay of recovery and account for mortality rate up to 70%-80%. In the present study, it was demonstrated that there was high prevalence (71.42%) of pathogenic bacteria found in Wounds. The results of percentage prevalence of the bacteria isolates from this study also indicated that the rate of isolation of Gram negative bacteria was more than Gram positive bacteria.
- 11. Name of the Scientist What was found in previous study Which was found in the presence study Messele A, Estifanos T et. al in 2018 Found that, 68.4% pathogenic bacteria in Wounds. It was found that, 71.42% pathogenic bacteria in Wounds. Priscilla R et.al in 2019 and Pondei k et.al in 2013 Found that, 86.20% & 85.05% Gram negative bacilli. It was found that, 84% GNB and Pseudomonas aeruginosa was the most prevalent gram negative organisms. Baba et.al in 2016 and Maharjan N et.al in 2020 Found that, 54.54% & 60.6% GPC in Wounds. Also reported that Staphylococcus aureus was the predominant bacterial species. It was found that, 16% GPC &Staphylococcus aureus was the only Gram positive organism. Maharjan N et.al in 2020 Found that, Piperacillin/Tazobactam(94.1%) was the most effective antibiotic for Pseudomonas aeruginosa. Pseudomonas aeruginosa was sensitive to piperacillin/Tazobactam (85.71%) & Gentamicin (78.57%). Baba et.al in 2016 Reported that, Staphylococcus aureus was sensitive to Rifampicin (96%) &Ciprofloxin (65%). It was found that, Staphylococcus aureus was highly sensitive to Vancomycin, Linezolid, Gentamicin. Mama M et al.2014 Found that, Ciprofloxin & Doxycycline were the most effective drugs for Klebsiella Pneumoniae. Klebsiella pneumoniaewas sensitive to Amikacin, Piperacillin/Tazobactam, Levofloxacin. Tinajanaserafimovskaet.alin 2020 Reported that, Escherichia coli was 88% sensitive to Clindamycin &Ciprofloxin. Escherichia coli was totally susceptible to Piperacillin/Tazobactam, 80% to Amikacin. Mahamedet.al Found that, Enterobacter sp was multiple drugs resistant. It was found that, Enterobacter sp was 100% resistant to Amikacin & Ceftriaxone.
- 12. Conclusion It can be concluded that in the present study Gram negative pseudomonas aeruginosa was the main bacteria responsible for wound infection, followed by Klebsiella pneumoniae, Escherichia coli, Staphylococcus aureus and other Gram negative bacteria. The results of the study is helpful for identifying the common causes of wound infection in the locality. It will be helpful to Select the appropriate antibiotic in appreciate dosage to control the infection. The accumulated results on different pathogens and their sensitivity will guide the physician in choosing empirical treatment of serious patients before the individual’s laboratory results are analysed. Helps the local pattern of antibiotic susceptibility.
- 13. THANK YOU
