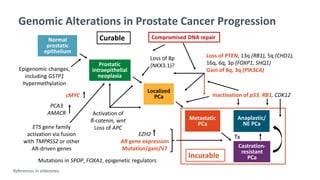
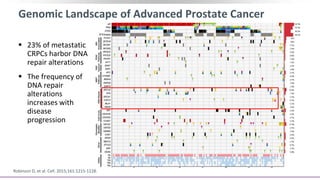
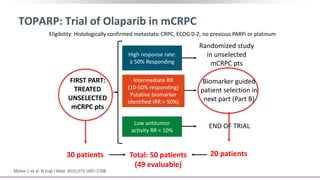
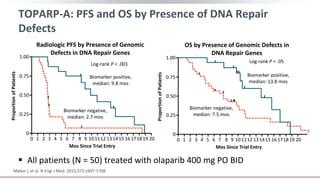
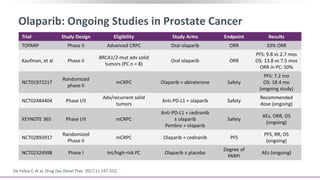
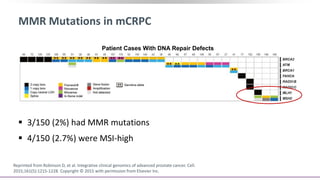
The document discusses the molecular and genetic understanding of castration-resistant prostate cancer (CRPC), emphasizing the role of genomic alterations and DNA repair defects. It highlights the heritability of prostate cancer, the impact of germline mutations in DNA repair genes, and the therapeutic implications of exploiting these defects for treatment, such as the use of PARP inhibitors. Additionally, it presents findings from studies on the prevalence of DNA repair alterations in prostate cancer and the efficacy of treatments targeting these genetic vulnerabilities.















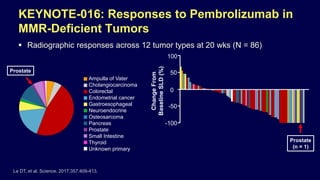
![Pembrolizumab for MSI-H or MMR-Deficient
Cancers
In May 2017, the PD-1 inhibitor pembrolizumab received
accelerated approval from FDA for:
– MSI-H or MMR-deficient CRC that has progressed following
treatment with a fluoropyrimidine, oxaliplatin, and irinotecan
– For ALL unresectable or metastatic MSI-H or MMR-deficient
SOLID TUMORS (pediatric and adult) that have progressed on
prior treatment and with no satisfactory alternative treatment
options
Dosage and administration (MSI-H cancers): 200 mg IV every 3
wks
Pembrolizumab [package insert]. 2017.](https://image.slidesharecdn.com/geneticsinprostatecancer-190204134738/85/Genetics-in-prostate-cancer-28-320.jpg)