RNA editing as a drug target in tryp. development of a high throughput sceening assay to identify inhibitors of rel 1 sulsa 2012
•
0 likes•56 views
Aspects of development of an HT assay for identifying inhibitors of the Tryp target protein REL 1: Potential microbial therapeutics by 'hit to lead'
Report
Share
Report
Share
Download to read offline
Recommended
Lab talk 201109 radioligand assay for validating in slico predicted rel1 anta...

Lab talk 201109 radioligand assay for validating in slico predicted rel1 anta...Laurence Dawkins-Hall
preliminary quantification of radio mimetic anti REL 1 hits by titration: Scoping likely IC50Paloma Pérez-Enfermedades raras de la piel

Los días 20 y 21 de octubre de 2016, la Fundacion Ramón Areces organizó un simposio internacional para analizar las 'Enfermedades raras de la piel: de la clínica al gen y viceversa'. El doctor Fernando Larcher Laguzzi, del CIEMAT-Universidad Carlos III de Madrid-IIS Fundación Jiménez Díaz, ejerció de coordinador.
Rna editing as a drug target identification of inhibitors of rel 1 bsp 210

Low through put radiomimetic assay to identify putative inhibitors of the Tryp enzyme REL 1 'in vitro' predicted by 'in silico' modelling
RNA editing as a drug target in tryp assay dev woodshole 2011

Aspects of HT assay development for identifying putative inhibitors of the Tryp RNA editing enzyme REL 1 by compound library screens
Recommended
Lab talk 201109 radioligand assay for validating in slico predicted rel1 anta...

Lab talk 201109 radioligand assay for validating in slico predicted rel1 anta...Laurence Dawkins-Hall
preliminary quantification of radio mimetic anti REL 1 hits by titration: Scoping likely IC50Paloma Pérez-Enfermedades raras de la piel

Los días 20 y 21 de octubre de 2016, la Fundacion Ramón Areces organizó un simposio internacional para analizar las 'Enfermedades raras de la piel: de la clínica al gen y viceversa'. El doctor Fernando Larcher Laguzzi, del CIEMAT-Universidad Carlos III de Madrid-IIS Fundación Jiménez Díaz, ejerció de coordinador.
Rna editing as a drug target identification of inhibitors of rel 1 bsp 210

Low through put radiomimetic assay to identify putative inhibitors of the Tryp enzyme REL 1 'in vitro' predicted by 'in silico' modelling
RNA editing as a drug target in tryp assay dev woodshole 2011

Aspects of HT assay development for identifying putative inhibitors of the Tryp RNA editing enzyme REL 1 by compound library screens
Polysaccharide based nanoparticles for encapsualtion and release of antineopl...

Polysaccharide based nanoparticles for encapsualtion and release of antineopl...Tomsk Polytechnic University
nanoparticles, chitosan, iron nanoparticles, doxorubicin, temozolomide , 5-fluorouracil, controlled releaseDocking & Designing Small Molecules within Rosetta Code Framework

This is the presentation I gave at my PhD defense.
Rna editing as a drug target identification of inhibitors of rel 1 bsp 2010

Low through put radiomimetic assay to identify putative inhibitors of the Tryp enzyme REL 1 'in vitro' predicted by 'in silico' modelling
Accessing genetically tagged heterocycle libraries via a chemoresistant DNA s...

Presented at the Global Medicinal Chemistry and GPCR Summit. To find out more, visit:
www.global-engage.com
Andreas Brunschweiger, an Independent Group Leader at TU Dortmund, discusses the limitations of DNA-encoded compound libraries (DELs) and getting around these.
Expression of Genetically Engineered Chitinase Gene of Pyrococcus furiosus

ABSTRACT: Wild-type Pyrococcus furiosus is most likely unable to grow on chitin in the natural biotope due to a nucleotide insertion which separates the chitinase gene into two ORFs, whereas a genetically engineered strain with the deleted nucleotide is able to grow on chitin. In the latest studies, the recombinant enzyme activity against the crystal chitins was examined. But there are still some conflictions. In our study, to shed a light on whether the construct composed of a catalytic domain and a chitin binding domain show any activity against crystalline chitin, the construct was created in the pET 28b (+) expression vector and expressed in Escherichia coli. The chitinase with an approximately 55 kDa molecular weight was determined. The activity of the enzyme was measured spectrophotometrically. Despite the presence of enzyme activity against the colloidal chitin, no significant activity against the crystal chitin has been measured.
Comparative analysis of docking of RGS9 gene

Comparative Analysis of Docking of Regulator of G-protein Signaling 9 (RGS9)-[Homo sapiens] and its Mutated Structure
Using Calorimetric Data to Drive Accuracy in Computer-Aided Drug Design

The slides are here, and the audio is on YouTube: https://youtu.be/400T1R7yduc
This talk begins by discussing the state of the art in computer-aided drug design (CADD), the need for improved force fields to increase accuracy, and the use of host-guest binding data to drive improvements in force fields. This talk was presented at the 2018 CalCon/ICCT meeting, August, 2018, Lake Tahoe, CA.
Simulating JAK-STAT Signal Transduction Pathway

Poster Presentation: Vishakha Sharma and Adriana Compagnoni. Grace Hopper Celebration of Women in Computing (GHC 2013), Minneapolis, MN, October 2-5, 2013.
DNA sequencing- Maxam- Gilbert sequencing

published a DNA sequencing method in 1977 based on chemical modification of DNA and subsequent cleavage at specific bases. Also known as chemical sequencing, this method allowed purified samples of double-stranded DNA to be used without further cloning.
Maxam-Gilbert sequencing requires radioactive labeling at one 5' end of the DNA and purification of the DNA fragment to be sequenced. Chemical treatment then generates breaks at a small proportion of one or two of the four nucleotide bases in each of four reactions (G, A+G, C, C+T). The concentration of the modifying chemicals is controlled to introduce on average one modification per DNA molecule. Thus a series of labeled fragments is generated, from the radiolabeled end to the first "cut" site in each molecule. The fragments in the four reactions are electrophoresed side by side in denaturing acrylamide gels for size separation. To visualize the fragments, the gel is exposed to X-ray film for autoradiography, yielding a series of dark bands each corresponding to a radiolabeled DNA fragment, from which the sequence may be inferred.
Diversity and distribution of nicotinic acetylcholine receptors in the locus ...

Diversity and distribution of nicotinic acetylcholine
receptors in the
locus ceruleus
neurons
Kani chemical biology

This course required us to present an article which prof gave us randomly. And my article is a review paper related to TLR signaling! I upload here just hope that it can be useful for someone who is interested in this approach for studyding TLR signaling dynamics based on Synthetic ligands!
Many thanks for your look at my presentation and leave some comments if I got mistakes inside!
Graffinity: Novel affinity ligands for downstream processing of proteins

Graffinity is a potentially revolutionary technology for the discovery of novel media for the purification of protein biologics. Proof of concept is sound. One example is a ligand that works as well as & better than protein A. We are open to suggestions or proposals for collaboration.
CONTACT fm@novalix-pharma.com
Optimization of experimental protocols for cellular lysis

In this project, we have compared existing sample preparation methods for proteomics studies against newly developed FASP method and our in-house developed SDS-TCA protocol. For our
preliminary studies, we have chosen a very well characterized soil microbe Pseudomonas putida.
Approach for limited cell ChIP-Seq on a semiconductor-based sequencing platform

Approach for limited cell ChIP-Seq on a semiconductor-based sequencing platformThermo Fisher Scientific
Dendritic cell (DC) lineages coordinate immune system activity
through functional specialization.
• Irf4, a transcription factor(TF), is required for CD11b+ DC
lineage development from bone marrow stem cells and has
been implicated in multiple inflammatory diseases, eg. asthma.
• The epigenetic consequences of immune specialization in
CD11b+ DCs and relation to inflammatory diseases remain
largely unexplored partly due to the difficulty of using highly
purified, and typically, limited populations of cells in ChIP-seq
(chromatin immunoprecipitation then sequencing) assays.
• A robust, multiplexed ChIP-seq protocol – using an input
control, TF (CTCF) and histone modification marks (H3K9me3-
methylation, H3K27ac-acetylation) - was developed using
limited amounts of K562 cells, for the Ion ProtonTM system.
• Peak-calling analysis was performed using using MACS2.
• Significant data correlations were observed with ENCODE.
• The Ion ProtonTM results are based on chromatin derived from
1 million(M) cells, making it viable for generating data from a
limited number of primary cells. This is in contrast to the 10M
cells recommended by ENCODE.
• The developed methodology was used to compare Irf4 genomic
binding sites generated from flow-sorted populations of 1, 3, 5,
and 20M CD11b+ lineage murine DCs.
• Comparable Irf4 ChIP-seq results were obtained from 5M
versus 20M cells, indicating that as low as 5M flow-sorted cells
can be used to acquire high quality(FDR: 10-19) data.
• We identified genomic Irf4 binding sites proximal to genes,
whose activity is consistent with CD11b+ DC lineage activity
and/or known to contribute to inflammatory disease.
• We examined Irf4 functional regulation of the identified gene
targets via RNA-seq analysis with CD11b+ DCs and a related
lineage, CD103+ DCs. Integrating expression analysis with
ChIP-seq indicates a unique CD11b+ DC gene expression
program concordant with Irf4 loci association in comparison to
CD103+ DC (data not shown).RNA editing as a drug target in tryp assay dev woods_hole2_0411 acs v2

Aspects of HT assay development for identifying putative inhibitors of the Tryp RNA editing enzyme REL 1 by compound library screens
RNA editing as a drug target in tryp. development of a high throughput fluore...

RNA editing as a drug target in tryp. development of a high throughput fluore...Laurence Dawkins-Hall
Procedural steps for developing a FRET based assay to screen for inhibitors of the Tryp enzyme REL 1: Potential microbial therapeuticsMore Related Content
What's hot
Polysaccharide based nanoparticles for encapsualtion and release of antineopl...

Polysaccharide based nanoparticles for encapsualtion and release of antineopl...Tomsk Polytechnic University
nanoparticles, chitosan, iron nanoparticles, doxorubicin, temozolomide , 5-fluorouracil, controlled releaseDocking & Designing Small Molecules within Rosetta Code Framework

This is the presentation I gave at my PhD defense.
Rna editing as a drug target identification of inhibitors of rel 1 bsp 2010

Low through put radiomimetic assay to identify putative inhibitors of the Tryp enzyme REL 1 'in vitro' predicted by 'in silico' modelling
Accessing genetically tagged heterocycle libraries via a chemoresistant DNA s...

Presented at the Global Medicinal Chemistry and GPCR Summit. To find out more, visit:
www.global-engage.com
Andreas Brunschweiger, an Independent Group Leader at TU Dortmund, discusses the limitations of DNA-encoded compound libraries (DELs) and getting around these.
Expression of Genetically Engineered Chitinase Gene of Pyrococcus furiosus

ABSTRACT: Wild-type Pyrococcus furiosus is most likely unable to grow on chitin in the natural biotope due to a nucleotide insertion which separates the chitinase gene into two ORFs, whereas a genetically engineered strain with the deleted nucleotide is able to grow on chitin. In the latest studies, the recombinant enzyme activity against the crystal chitins was examined. But there are still some conflictions. In our study, to shed a light on whether the construct composed of a catalytic domain and a chitin binding domain show any activity against crystalline chitin, the construct was created in the pET 28b (+) expression vector and expressed in Escherichia coli. The chitinase with an approximately 55 kDa molecular weight was determined. The activity of the enzyme was measured spectrophotometrically. Despite the presence of enzyme activity against the colloidal chitin, no significant activity against the crystal chitin has been measured.
Comparative analysis of docking of RGS9 gene

Comparative Analysis of Docking of Regulator of G-protein Signaling 9 (RGS9)-[Homo sapiens] and its Mutated Structure
Using Calorimetric Data to Drive Accuracy in Computer-Aided Drug Design

The slides are here, and the audio is on YouTube: https://youtu.be/400T1R7yduc
This talk begins by discussing the state of the art in computer-aided drug design (CADD), the need for improved force fields to increase accuracy, and the use of host-guest binding data to drive improvements in force fields. This talk was presented at the 2018 CalCon/ICCT meeting, August, 2018, Lake Tahoe, CA.
Simulating JAK-STAT Signal Transduction Pathway

Poster Presentation: Vishakha Sharma and Adriana Compagnoni. Grace Hopper Celebration of Women in Computing (GHC 2013), Minneapolis, MN, October 2-5, 2013.
DNA sequencing- Maxam- Gilbert sequencing

published a DNA sequencing method in 1977 based on chemical modification of DNA and subsequent cleavage at specific bases. Also known as chemical sequencing, this method allowed purified samples of double-stranded DNA to be used without further cloning.
Maxam-Gilbert sequencing requires radioactive labeling at one 5' end of the DNA and purification of the DNA fragment to be sequenced. Chemical treatment then generates breaks at a small proportion of one or two of the four nucleotide bases in each of four reactions (G, A+G, C, C+T). The concentration of the modifying chemicals is controlled to introduce on average one modification per DNA molecule. Thus a series of labeled fragments is generated, from the radiolabeled end to the first "cut" site in each molecule. The fragments in the four reactions are electrophoresed side by side in denaturing acrylamide gels for size separation. To visualize the fragments, the gel is exposed to X-ray film for autoradiography, yielding a series of dark bands each corresponding to a radiolabeled DNA fragment, from which the sequence may be inferred.
Diversity and distribution of nicotinic acetylcholine receptors in the locus ...

Diversity and distribution of nicotinic acetylcholine
receptors in the
locus ceruleus
neurons
Kani chemical biology

This course required us to present an article which prof gave us randomly. And my article is a review paper related to TLR signaling! I upload here just hope that it can be useful for someone who is interested in this approach for studyding TLR signaling dynamics based on Synthetic ligands!
Many thanks for your look at my presentation and leave some comments if I got mistakes inside!
Graffinity: Novel affinity ligands for downstream processing of proteins

Graffinity is a potentially revolutionary technology for the discovery of novel media for the purification of protein biologics. Proof of concept is sound. One example is a ligand that works as well as & better than protein A. We are open to suggestions or proposals for collaboration.
CONTACT fm@novalix-pharma.com
Optimization of experimental protocols for cellular lysis

In this project, we have compared existing sample preparation methods for proteomics studies against newly developed FASP method and our in-house developed SDS-TCA protocol. For our
preliminary studies, we have chosen a very well characterized soil microbe Pseudomonas putida.
Approach for limited cell ChIP-Seq on a semiconductor-based sequencing platform

Approach for limited cell ChIP-Seq on a semiconductor-based sequencing platformThermo Fisher Scientific
Dendritic cell (DC) lineages coordinate immune system activity
through functional specialization.
• Irf4, a transcription factor(TF), is required for CD11b+ DC
lineage development from bone marrow stem cells and has
been implicated in multiple inflammatory diseases, eg. asthma.
• The epigenetic consequences of immune specialization in
CD11b+ DCs and relation to inflammatory diseases remain
largely unexplored partly due to the difficulty of using highly
purified, and typically, limited populations of cells in ChIP-seq
(chromatin immunoprecipitation then sequencing) assays.
• A robust, multiplexed ChIP-seq protocol – using an input
control, TF (CTCF) and histone modification marks (H3K9me3-
methylation, H3K27ac-acetylation) - was developed using
limited amounts of K562 cells, for the Ion ProtonTM system.
• Peak-calling analysis was performed using using MACS2.
• Significant data correlations were observed with ENCODE.
• The Ion ProtonTM results are based on chromatin derived from
1 million(M) cells, making it viable for generating data from a
limited number of primary cells. This is in contrast to the 10M
cells recommended by ENCODE.
• The developed methodology was used to compare Irf4 genomic
binding sites generated from flow-sorted populations of 1, 3, 5,
and 20M CD11b+ lineage murine DCs.
• Comparable Irf4 ChIP-seq results were obtained from 5M
versus 20M cells, indicating that as low as 5M flow-sorted cells
can be used to acquire high quality(FDR: 10-19) data.
• We identified genomic Irf4 binding sites proximal to genes,
whose activity is consistent with CD11b+ DC lineage activity
and/or known to contribute to inflammatory disease.
• We examined Irf4 functional regulation of the identified gene
targets via RNA-seq analysis with CD11b+ DCs and a related
lineage, CD103+ DCs. Integrating expression analysis with
ChIP-seq indicates a unique CD11b+ DC gene expression
program concordant with Irf4 loci association in comparison to
CD103+ DC (data not shown).What's hot (20)
Polysaccharide based nanoparticles for encapsualtion and release of antineopl...

Polysaccharide based nanoparticles for encapsualtion and release of antineopl...
Docking & Designing Small Molecules within Rosetta Code Framework

Docking & Designing Small Molecules within Rosetta Code Framework
Rna editing as a drug target identification of inhibitors of rel 1 bsp 2010

Rna editing as a drug target identification of inhibitors of rel 1 bsp 2010
Accessing genetically tagged heterocycle libraries via a chemoresistant DNA s...

Accessing genetically tagged heterocycle libraries via a chemoresistant DNA s...
Expression of Genetically Engineered Chitinase Gene of Pyrococcus furiosus

Expression of Genetically Engineered Chitinase Gene of Pyrococcus furiosus
Using Calorimetric Data to Drive Accuracy in Computer-Aided Drug Design

Using Calorimetric Data to Drive Accuracy in Computer-Aided Drug Design
Diversity and distribution of nicotinic acetylcholine receptors in the locus ...

Diversity and distribution of nicotinic acetylcholine receptors in the locus ...
Graffinity: Novel affinity ligands for downstream processing of proteins

Graffinity: Novel affinity ligands for downstream processing of proteins
Optimization of experimental protocols for cellular lysis

Optimization of experimental protocols for cellular lysis
Approach for limited cell ChIP-Seq on a semiconductor-based sequencing platform

Approach for limited cell ChIP-Seq on a semiconductor-based sequencing platform
Similar to RNA editing as a drug target in tryp. development of a high throughput sceening assay to identify inhibitors of rel 1 sulsa 2012
RNA editing as a drug target in tryp assay dev woods_hole2_0411 acs v2

Aspects of HT assay development for identifying putative inhibitors of the Tryp RNA editing enzyme REL 1 by compound library screens
RNA editing as a drug target in tryp. development of a high throughput fluore...

RNA editing as a drug target in tryp. development of a high throughput fluore...Laurence Dawkins-Hall
Procedural steps for developing a FRET based assay to screen for inhibitors of the Tryp enzyme REL 1: Potential microbial therapeuticsBacterial Endotoxin Test

Bacterial Endotoxin Test in sterile pharmaceutical
Read More - http://www.pharmaguideline.com/2011/04/bacterial-endotoxin-test-bet-validation.html
Lab talk 020410 inducing rel 1 to set up ht fret assay_progress

Outlining procedural pipeline for expressing and purifying active rREL 1 for incorporation into high throughput fluorescent (FRET) screening assay for identifying antagonistic hits of REL 1
Rapid and real-time diagnosis of Laem-Singh virus using a portable Real-Amp T...

The most prominent of growth-retarded in black tiger shrimp (Penaeus monodon) is associated with an infection of Laem-Singh virus (LSNV) that has been involved the reduction of annual production volume. To more accurately, rapidly and easily determine its effect on shrimp industries for further control disease, a portable Real-Amp Turbidimeter with real-time detection method was evaluated for LSNV detection.The device exploited the turbidity that produced by-product of magnesium pyrophosphate and displayed turbidity values in real-time on the provided software screen. The results of quantitative can be accomplished by reaction time measuring. The sensitivity of a turbidity detection was comparable to that of the nested RT-PCR, while it was 1000-time more sensitive than RT-PCR. With the use of a portable Real-Amp Turbidimeter. The application of RT-LAMP using a portable Real-Amp Turbidimeter offers a simple and rapid detection platform for LSNV detection in the field assessment. So, This 's awesome technic not only for detection of LSNV but also for many pathogen.
RNA editing as a drug target development of a high through put fluorescent as...

RNA editing as a drug target development of a high through put fluorescent as...Laurence Dawkins-Hall
Procedural steps for developing a FRET based assay to screen for inhibitors of the Tryp enzyme REL 1: Potential microbial therapeuticsCharacteristics of Loop Mediated Isothermal Amplification Technique

Characteristics of LAMP Technique & Comparison with in Vitro Amplification Techniques
Real time PCR practical training 

this power point cover the most important point of how to use and analyse Real time PCR result
SmartScreen Technology for Building a Better Assay

A lipid derived nanoparticle the recreates the cellular membrane in solution based assays. See increased enzymatic activity, identify more relevant biological substrates, find novel hits from the compound library.
Fragment screening library workshop (IQPC 2008)

I also ran a workshop on selection of compounds for fragment screening just before the 2008 IQPC compound library conference and these are the slides I used.
Optimal RNAlater® incubation and removal conditions prior to isolation of tot...

RNA is highly sensitive to degradation. Handling methods and prolonged storage of cells can greatly affect the quality of the RNA that can be later isolated. Contamination with RNases is the most significant problem, especially as they are so ubiquitous in the environment. They can degrade RNA to the point where results of downstream analyses become meaningless.
Submerging cells in RNAlater, an RNA stabilization reagent, helps to stabilize the RNA within the cells and prevent degradation, supporting accurate downstream gene expression analyses. However, to avoid any interference from any RNAlater components in isolation and analyses, cells must be pelleted and the reagent must be removed. The separation of cells from excess RNAlater via centrifugation is impeded due to the higher density of the reagent compared to standard culture medium. This means it requires higher centrifugal forces, which might damage cells due to increased shearing forces, leading to reduced RNA yield. The aim of this study was to establish the optimal conditions for the recovery of cells from RNAlater after RNA stabilization for maximum RNA yield and integrity.
SuperScript IV Reverse Transcriptase for RNA Analysis | ESHG 2015 Poster PS14...

SuperScript IV Reverse Transcriptase for RNA Analysis | ESHG 2015 Poster PS14...Thermo Fisher Scientific
Survey and interview studies conducted over a three year period revealed that researchers are not satisfied with their current reverse transcriptase and are performing reactions with increasingly difficult samples, such as poorly purified RNA and unpurified RNA (direct RT) that both contain inhibitors. To meet this performance gap, the Thermo Fisher Life Sciences Solutions group produced a new reverse transcriptase, SuperScript® IV, and experiments we performed show that it is the most robust reverse transcriptase compared to other enzymes. SuperScript® IV characterization was performed in the context of “real world” situations where users do not have perfect RNA samples. In the presence of a variety of inhibitors, we demonstrate that SuperScript® IV possesses superior performance in a variety of inhibitors, such as alcohols, salts, detergents, phenol, heparin, hematin, bile salts, and formalin typically found in sample preparation reagents, cell lines, blood, feces, and FFPE samples. This enzyme can even detect RNA targets in unpurified RNA samples (directly lysed cells) and whole blood without sacrificing sensitivity and yield. The introduction of SuperScript® IV enables researchers to obtain more consistent results independent of sample quality and simplify and speed up workflows by eliminating RNA purification.Similar to RNA editing as a drug target in tryp. development of a high throughput sceening assay to identify inhibitors of rel 1 sulsa 2012 (20)
RNA editing as a drug target in tryp assay dev woods_hole2_0411 acs v2

RNA editing as a drug target in tryp assay dev woods_hole2_0411 acs v2
RNA editing as a drug target in tryp. development of a high throughput fluore...

RNA editing as a drug target in tryp. development of a high throughput fluore...
Lab talk 020410 inducing rel 1 to set up ht fret assay_progress

Lab talk 020410 inducing rel 1 to set up ht fret assay_progress
Rapid and real-time diagnosis of Laem-Singh virus using a portable Real-Amp T...

Rapid and real-time diagnosis of Laem-Singh virus using a portable Real-Amp T...
RNA editing as a drug target development of a high through put fluorescent as...

RNA editing as a drug target development of a high through put fluorescent as...
Characteristics of Loop Mediated Isothermal Amplification Technique

Characteristics of Loop Mediated Isothermal Amplification Technique
SmartScreen Technology for Building a Better Assay

SmartScreen Technology for Building a Better Assay
Optimal RNAlater® incubation and removal conditions prior to isolation of tot...

Optimal RNAlater® incubation and removal conditions prior to isolation of tot...
SuperScript IV Reverse Transcriptase for RNA Analysis | ESHG 2015 Poster PS14...

SuperScript IV Reverse Transcriptase for RNA Analysis | ESHG 2015 Poster PS14...
housman_mini_4_5 foot PPT poster template 50 percent

housman_mini_4_5 foot PPT poster template 50 percent
More from Laurence Dawkins-Hall
DSRG report 2001

Describing the activities of the DNA Sequencing Research group (DSRG) of the ABRF for the year 2001:
Journal of Bio molecular Techniques 12:27–29 © 2001 ABRF
An evaluation of methods used to sequence pGEM template within core facilitie...

An evaluation of methods used to sequence pGEM template within core facilitie...Laurence Dawkins-Hall
A horizontal study by ABRF designed to investigate & collate cycle sequencing methods for the pGEM template and subsequent data output on automated sequencing platforms (cf. AB 3700 & AB 3100) 06 isafg p77 genome wide expression consequences of a disease resistance qtl ...

06 isafg p77 genome wide expression consequences of a disease resistance qtl ...Laurence Dawkins-Hall
Presentation describing the genetic pathophysiology of Trypanosomiasis resistance in WT cattleGenomic approaches to trypanosome resistance

Elucidating changes in gene expression in Tryp susceptible and resistant cattle during progression of tryp infection using Affymetrix gene expression Micro arrays
Genome responses of trypanosome infected cattle

Genomic gene expression changes resulting from Trypanosomiasis: a horizontal study Examining expression changes elucidated by micro arrays in seminal tissues associated with the pathophysiology of Trypanosomiasis during disease progression
Systemic analysis of data combined from genetic qtl's and gene expression dat...

Systemic analysis of data combined from genetic qtl's and gene expression dat...Laurence Dawkins-Hall
Elucidating changes in gene expression by Micro array genomic sweeps of genetic QTLs linked to Tryp resistance in WT cattle to identify putative candidates underpinning pathophysiology A systematic, data driven approach to the combined analysis of microarray and...

A systematic, data driven approach to the combined analysis of microarray and...Laurence Dawkins-Hall
The use of gene expression data from Micro arrays coupled with WT QTL's linked to Tryp resistance phenotypes in Cattle to elucidate pertinent genetic changes underpinning phenotype in putative candidate genes Lab talk 181209 radioligand assay for validating in silico rel 1 antagonist h...

Lab talk 181209 radioligand assay for validating in silico rel 1 antagonist h...Laurence Dawkins-Hall
Titrating preliminary hits derived from anti REL 1 radio mimetic assay: Precursor to full regression analysis of lead compounds to establish IC50'sLab talk 180909 radioligand assay to validate in silico predicted rel1 inhib...

Lab talk 180909 radioligand assay to validate in silico predicted rel1 inhib...Laurence Dawkins-Hall
implementation of a radio ligand assay to validate antagonists of the Tryp RNA editing enzyme REL 1 to validate in silico predictionsLab talk 060809 radioligand assay to validate in slico predicted rel inhibito...

Lab talk 060809 radioligand assay to validate in slico predicted rel inhibito...Laurence Dawkins-Hall
Radio ligand assay to validate putative antagonists of REL1 predicted by in silico modellingLabtalk 101210 optimising fret assay_den_optimising r_rel 1 production_heat s...

Labtalk 101210 optimising fret assay_den_optimising r_rel 1 production_heat s...Laurence Dawkins-Hall
optimising HT FRET drug screening assay with ref to denaturation regimen; potentiation of soluble REL 1 production by heat shock inclusion Labtalk 151010 optimising fret assay_ligation buffer_[atp]_fluorophores![Labtalk 151010 optimising fret assay_ligation buffer_[atp]_fluorophores](data:image/gif;base64,R0lGODlhAQABAIAAAAAAAP///yH5BAEAAAAALAAAAAABAAEAAAIBRAA7)
![Labtalk 151010 optimising fret assay_ligation buffer_[atp]_fluorophores](data:image/gif;base64,R0lGODlhAQABAIAAAAAAAP///yH5BAEAAAAALAAAAAABAAEAAAIBRAA7)
Optimising parameters associated with a high through put FRET assay for identifying Tryp REL 1 antagonists, viz.
1. Ligation conditions, cf. buffer composition e.g. pH; ligation time etc
2. Mixing and matching fluorophores to maximise S:N and hence assay sensitivity
Lab talk 300710 optimisation of fret assay parameters_calculating km of atp s...

Lab talk 300710 optimisation of fret assay parameters_calculating km of atp s...Laurence Dawkins-Hall
Optimisation of FRET parameters to improve FRET compound screening assay, viz
1. Denaturation regimen
2. Establishing Km of [ATP] substrateLab talk 070510 inducing r_rel1 for establishing fret assay_ic50 studies with...

Lab talk 070510 inducing r_rel1 for establishing fret assay_ic50 studies with...Laurence Dawkins-Hall
1. more work at optimising expression and purification of rREL 1 for incorporation into a high throughput FRET assay for compound library screens to identify REL 1 antagonists
2. Regression analysis of titrants of compounds identified as demonstrating REL 1 inhibition by radiomimetic assay; quantification of efficacy by means of IC50 Lab talk 190210 efficacy studies on radioligand hits_beginnings of fret assay...

Lab talk 190210 efficacy studies on radioligand hits_beginnings of fret assay...Laurence Dawkins-Hall
1. Titration of REL 1 antagonist hits identified by radio mimetic assay to quantify efficacy (IC50)
2. Exposition of HT FRET assay principles for large scale compound library screens to empirically identify REL 1 antagonist hitsLab talk 020710 comparing bac r_rel 1 with e coli rrel 1 for use in fret assay

Comparing activity of baculovirus and E Coli expressed rREL 1 fractions in context of HT FRET assay for screening compound libraries to identify REL1 antagonist hits
Genomic analysis of a hypertensive qtl on rat chr 1

Transcriptomic analysis of rat QTL linked to hypertension by exomic sequencing and micro array gene expression analysis
Abstract bookfinal-3 bsp 2012 glasgow

Abstract Booklet_BSP meeting Strathclyde University, Glasgow, 2012
More from Laurence Dawkins-Hall (18)
An evaluation of methods used to sequence pGEM template within core facilitie...

An evaluation of methods used to sequence pGEM template within core facilitie...
06 isafg p77 genome wide expression consequences of a disease resistance qtl ...

06 isafg p77 genome wide expression consequences of a disease resistance qtl ...
Systemic analysis of data combined from genetic qtl's and gene expression dat...

Systemic analysis of data combined from genetic qtl's and gene expression dat...
A systematic, data driven approach to the combined analysis of microarray and...

A systematic, data driven approach to the combined analysis of microarray and...
Lab talk 181209 radioligand assay for validating in silico rel 1 antagonist h...

Lab talk 181209 radioligand assay for validating in silico rel 1 antagonist h...
Lab talk 180909 radioligand assay to validate in silico predicted rel1 inhib...

Lab talk 180909 radioligand assay to validate in silico predicted rel1 inhib...
Lab talk 060809 radioligand assay to validate in slico predicted rel inhibito...

Lab talk 060809 radioligand assay to validate in slico predicted rel inhibito...
Labtalk 101210 optimising fret assay_den_optimising r_rel 1 production_heat s...

Labtalk 101210 optimising fret assay_den_optimising r_rel 1 production_heat s...
Labtalk 151010 optimising fret assay_ligation buffer_[atp]_fluorophores![Labtalk 151010 optimising fret assay_ligation buffer_[atp]_fluorophores](data:image/gif;base64,R0lGODlhAQABAIAAAAAAAP///yH5BAEAAAAALAAAAAABAAEAAAIBRAA7)
![Labtalk 151010 optimising fret assay_ligation buffer_[atp]_fluorophores](data:image/gif;base64,R0lGODlhAQABAIAAAAAAAP///yH5BAEAAAAALAAAAAABAAEAAAIBRAA7)
Labtalk 151010 optimising fret assay_ligation buffer_[atp]_fluorophores
Lab talk 300710 optimisation of fret assay parameters_calculating km of atp s...

Lab talk 300710 optimisation of fret assay parameters_calculating km of atp s...
Lab talk 070510 inducing r_rel1 for establishing fret assay_ic50 studies with...

Lab talk 070510 inducing r_rel1 for establishing fret assay_ic50 studies with...
Lab talk 190210 efficacy studies on radioligand hits_beginnings of fret assay...

Lab talk 190210 efficacy studies on radioligand hits_beginnings of fret assay...
Lab talk 020710 comparing bac r_rel 1 with e coli rrel 1 for use in fret assay

Lab talk 020710 comparing bac r_rel 1 with e coli rrel 1 for use in fret assay
Genomic analysis of a hypertensive qtl on rat chr 1

Genomic analysis of a hypertensive qtl on rat chr 1
Recently uploaded
(May 29th, 2024) Advancements in Intravital Microscopy- Insights for Preclini...

(May 29th, 2024) Advancements in Intravital Microscopy- Insights for Preclini...Scintica Instrumentation
Intravital microscopy (IVM) is a powerful tool utilized to study cellular behavior over time and space in vivo. Much of our understanding of cell biology has been accomplished using various in vitro and ex vivo methods; however, these studies do not necessarily reflect the natural dynamics of biological processes. Unlike traditional cell culture or fixed tissue imaging, IVM allows for the ultra-fast high-resolution imaging of cellular processes over time and space and were studied in its natural environment. Real-time visualization of biological processes in the context of an intact organism helps maintain physiological relevance and provide insights into the progression of disease, response to treatments or developmental processes.
In this webinar we give an overview of advanced applications of the IVM system in preclinical research. IVIM technology is a provider of all-in-one intravital microscopy systems and solutions optimized for in vivo imaging of live animal models at sub-micron resolution. The system’s unique features and user-friendly software enables researchers to probe fast dynamic biological processes such as immune cell tracking, cell-cell interaction as well as vascularization and tumor metastasis with exceptional detail. This webinar will also give an overview of IVM being utilized in drug development, offering a view into the intricate interaction between drugs/nanoparticles and tissues in vivo and allows for the evaluation of therapeutic intervention in a variety of tissues and organs. This interdisciplinary collaboration continues to drive the advancements of novel therapeutic strategies.
Observation of Io’s Resurfacing via Plume Deposition Using Ground-based Adapt...

Since volcanic activity was first discovered on Io from Voyager images in 1979, changes
on Io’s surface have been monitored from both spacecraft and ground-based telescopes.
Here, we present the highest spatial resolution images of Io ever obtained from a groundbased telescope. These images, acquired by the SHARK-VIS instrument on the Large
Binocular Telescope, show evidence of a major resurfacing event on Io’s trailing hemisphere. When compared to the most recent spacecraft images, the SHARK-VIS images
show that a plume deposit from a powerful eruption at Pillan Patera has covered part
of the long-lived Pele plume deposit. Although this type of resurfacing event may be common on Io, few have been detected due to the rarity of spacecraft visits and the previously low spatial resolution available from Earth-based telescopes. The SHARK-VIS instrument ushers in a new era of high resolution imaging of Io’s surface using adaptive
optics at visible wavelengths.
Nutraceutical market, scope and growth: Herbal drug technology

As consumer awareness of health and wellness rises, the nutraceutical market—which includes goods like functional meals, drinks, and dietary supplements that provide health advantages beyond basic nutrition—is growing significantly. As healthcare expenses rise, the population ages, and people want natural and preventative health solutions more and more, this industry is increasing quickly. Further driving market expansion are product formulation innovations and the use of cutting-edge technology for customized nutrition. With its worldwide reach, the nutraceutical industry is expected to keep growing and provide significant chances for research and investment in a number of categories, including vitamins, minerals, probiotics, and herbal supplements.
extra-chromosomal-inheritance[1].pptx.pdfpdf![extra-chromosomal-inheritance[1].pptx.pdfpdf](data:image/gif;base64,R0lGODlhAQABAIAAAAAAAP///yH5BAEAAAAALAAAAAABAAEAAAIBRAA7)
![extra-chromosomal-inheritance[1].pptx.pdfpdf](data:image/gif;base64,R0lGODlhAQABAIAAAAAAAP///yH5BAEAAAAALAAAAAABAAEAAAIBRAA7)
Slide 1: Title Slide
Extrachromosomal Inheritance
Slide 2: Introduction to Extrachromosomal Inheritance
Definition: Extrachromosomal inheritance refers to the transmission of genetic material that is not found within the nucleus.
Key Components: Involves genes located in mitochondria, chloroplasts, and plasmids.
Slide 3: Mitochondrial Inheritance
Mitochondria: Organelles responsible for energy production.
Mitochondrial DNA (mtDNA): Circular DNA molecule found in mitochondria.
Inheritance Pattern: Maternally inherited, meaning it is passed from mothers to all their offspring.
Diseases: Examples include Leber’s hereditary optic neuropathy (LHON) and mitochondrial myopathy.
Slide 4: Chloroplast Inheritance
Chloroplasts: Organelles responsible for photosynthesis in plants.
Chloroplast DNA (cpDNA): Circular DNA molecule found in chloroplasts.
Inheritance Pattern: Often maternally inherited in most plants, but can vary in some species.
Examples: Variegation in plants, where leaf color patterns are determined by chloroplast DNA.
Slide 5: Plasmid Inheritance
Plasmids: Small, circular DNA molecules found in bacteria and some eukaryotes.
Features: Can carry antibiotic resistance genes and can be transferred between cells through processes like conjugation.
Significance: Important in biotechnology for gene cloning and genetic engineering.
Slide 6: Mechanisms of Extrachromosomal Inheritance
Non-Mendelian Patterns: Do not follow Mendel’s laws of inheritance.
Cytoplasmic Segregation: During cell division, organelles like mitochondria and chloroplasts are randomly distributed to daughter cells.
Heteroplasmy: Presence of more than one type of organellar genome within a cell, leading to variation in expression.
Slide 7: Examples of Extrachromosomal Inheritance
Four O’clock Plant (Mirabilis jalapa): Shows variegated leaves due to different cpDNA in leaf cells.
Petite Mutants in Yeast: Result from mutations in mitochondrial DNA affecting respiration.
Slide 8: Importance of Extrachromosomal Inheritance
Evolution: Provides insight into the evolution of eukaryotic cells.
Medicine: Understanding mitochondrial inheritance helps in diagnosing and treating mitochondrial diseases.
Agriculture: Chloroplast inheritance can be used in plant breeding and genetic modification.
Slide 9: Recent Research and Advances
Gene Editing: Techniques like CRISPR-Cas9 are being used to edit mitochondrial and chloroplast DNA.
Therapies: Development of mitochondrial replacement therapy (MRT) for preventing mitochondrial diseases.
Slide 10: Conclusion
Summary: Extrachromosomal inheritance involves the transmission of genetic material outside the nucleus and plays a crucial role in genetics, medicine, and biotechnology.
Future Directions: Continued research and technological advancements hold promise for new treatments and applications.
Slide 11: Questions and Discussion
Invite Audience: Open the floor for any questions or further discussion on the topic.
原版制作(carleton毕业证书)卡尔顿大学毕业证硕士文凭原版一模一样

原版纸张【微信:741003700 】【(carleton毕业证书)卡尔顿大学毕业证】【微信:741003700 】学位证,留信认证(真实可查,永久存档)offer、雅思、外壳等材料/诚信可靠,可直接看成品样本,帮您解决无法毕业带来的各种难题!外壳,原版制作,诚信可靠,可直接看成品样本。行业标杆!精益求精,诚心合作,真诚制作!多年品质 ,按需精细制作,24小时接单,全套进口原装设备。十五年致力于帮助留学生解决难题,包您满意。
本公司拥有海外各大学样板无数,能完美还原海外各大学 Bachelor Diploma degree, Master Degree Diploma
1:1完美还原海外各大学毕业材料上的工艺:水印,阴影底纹,钢印LOGO烫金烫银,LOGO烫金烫银复合重叠。文字图案浮雕、激光镭射、紫外荧光、温感、复印防伪等防伪工艺。材料咨询办理、认证咨询办理请加学历顾问Q/微741003700
留信网认证的作用:
1:该专业认证可证明留学生真实身份
2:同时对留学生所学专业登记给予评定
3:国家专业人才认证中心颁发入库证书
4:这个认证书并且可以归档倒地方
5:凡事获得留信网入网的信息将会逐步更新到个人身份内,将在公安局网内查询个人身份证信息后,同步读取人才网入库信息
6:个人职称评审加20分
7:个人信誉贷款加10分
8:在国家人才网主办的国家网络招聘大会中纳入资料,供国家高端企业选择人才
Toxic effects of heavy metals : Lead and Arsenic

Heavy metals are naturally occuring metallic chemical elements that have relatively high density, and are toxic at even low concentrations. All toxic metals are termed as heavy metals irrespective of their atomic mass and density, eg. arsenic, lead, mercury, cadmium, thallium, chromium, etc.
Comparing Evolved Extractive Text Summary Scores of Bidirectional Encoder Rep...

Comparing Evolved Extractive Text Summary Scores of Bidirectional Encoder Rep...University of Maribor
Slides from:
11th International Conference on Electrical, Electronics and Computer Engineering (IcETRAN), Niš, 3-6 June 2024
Track: Artificial Intelligence
https://www.etran.rs/2024/en/home-english/Deep Behavioral Phenotyping in Systems Neuroscience for Functional Atlasing a...

Functional Magnetic Resonance Imaging (fMRI) provides means to characterize brain activations in response to behavior. However, cognitive neuroscience has been limited to group-level effects referring to the performance of specific tasks. To obtain the functional profile of elementary cognitive mechanisms, the combination of brain responses to many tasks is required. Yet, to date, both structural atlases and parcellation-based activations do not fully account for cognitive function and still present several limitations. Further, they do not adapt overall to individual characteristics. In this talk, I will give an account of deep-behavioral phenotyping strategies, namely data-driven methods in large task-fMRI datasets, to optimize functional brain-data collection and improve inference of effects-of-interest related to mental processes. Key to this approach is the employment of fast multi-functional paradigms rich on features that can be well parametrized and, consequently, facilitate the creation of psycho-physiological constructs to be modelled with imaging data. Particular emphasis will be given to music stimuli when studying high-order cognitive mechanisms, due to their ecological nature and quality to enable complex behavior compounded by discrete entities. I will also discuss how deep-behavioral phenotyping and individualized models applied to neuroimaging data can better account for the subject-specific organization of domain-general cognitive systems in the human brain. Finally, the accumulation of functional brain signatures brings the possibility to clarify relationships among tasks and create a univocal link between brain systems and mental functions through: (1) the development of ontologies proposing an organization of cognitive processes; and (2) brain-network taxonomies describing functional specialization. To this end, tools to improve commensurability in cognitive science are necessary, such as public repositories, ontology-based platforms and automated meta-analysis tools. I will thus discuss some brain-atlasing resources currently under development, and their applicability in cognitive as well as clinical neuroscience.
Orion Air Quality Monitoring Systems - CWS

Professional air quality monitoring systems provide immediate, on-site data for analysis, compliance, and decision-making.
Monitor common gases, weather parameters, particulates.
Recently uploaded (20)
(May 29th, 2024) Advancements in Intravital Microscopy- Insights for Preclini...

(May 29th, 2024) Advancements in Intravital Microscopy- Insights for Preclini...
Mammalian Pineal Body Structure and Also Functions

Mammalian Pineal Body Structure and Also Functions
Observation of Io’s Resurfacing via Plume Deposition Using Ground-based Adapt...

Observation of Io’s Resurfacing via Plume Deposition Using Ground-based Adapt...
Nutraceutical market, scope and growth: Herbal drug technology

Nutraceutical market, scope and growth: Herbal drug technology
In silico drugs analogue design: novobiocin analogues.pptx

In silico drugs analogue design: novobiocin analogues.pptx
erythropoiesis-I_mechanism& clinical significance.pptx

erythropoiesis-I_mechanism& clinical significance.pptx
Comparing Evolved Extractive Text Summary Scores of Bidirectional Encoder Rep...

Comparing Evolved Extractive Text Summary Scores of Bidirectional Encoder Rep...
Deep Behavioral Phenotyping in Systems Neuroscience for Functional Atlasing a...

Deep Behavioral Phenotyping in Systems Neuroscience for Functional Atlasing a...
BLOOD AND BLOOD COMPONENT- introduction to blood physiology

BLOOD AND BLOOD COMPONENT- introduction to blood physiology
RNA editing as a drug target in tryp. development of a high throughput sceening assay to identify inhibitors of rel 1 sulsa 2012
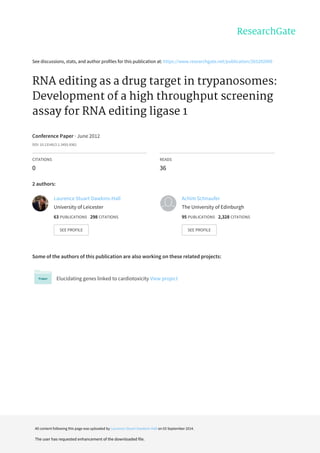
- 2. ,QVWLWXWH RI ,PPXQRORJ ,QIHFWLRQ 5HVHDUFK DQG &HQWUH RI ,PPXQ,QVWLWXWH RI ,PPXQRORJ ,QIHFWLRQ 5HVHDUFK DQG &HQWUH RI ,PPXQ,QVWLWXWH RI ,PPXQRORJ ,QIHFWLRQ 5HVHDUFK DQG &HQWUH RI ,PPXQ,QVWLWXWH RI ,PPXQRORJ ,QIHFWLRQ 5HVHDUFK DQG &HQWUH RI ,PPXQLW ,QIHFWLRQ (YROXWLRQ 8QLYHUVLW RI (GLQEXUJK 8.LW ,QIHFWLRQ (YROXWLRQ 8QLYHUVLW RI (GLQEXUJK 8.LW ,QIHFWLRQ (YROXWLRQ 8QLYHUVLW RI (GLQEXUJK 8.LW ,QIHFWLRQ (YROXWLRQ 8QLYHUVLW RI (GLQEXUJK 8. ! # $ $! # $ $ %% FRET (Fluorescence Resonance Energy Transfer) is achieved when two fluorophores with overlapping spectra are fixed in close proximity, such that virtual photons are transferred between a ‘donor’ (‘D’) and ‘acceptor’ fluorophore (‘A’), resulting in acceptor fluorescence when the donor is excited, which can be quantified by spectrophotometry Annealing ofAnnealing of fluorophorefluorophore--labelledlabelled 55’’ and 3and 3’’ RNA substrates toRNA substrates to an RNA bridge (thean RNA bridge (the ““guide RNAguide RNA””) anchors the donor and) anchors the donor and acceptor fluorophores in close proximityacceptor fluorophores in close proximity Consequently, transfer of virtual photons results in FRETConsequently, transfer of virtual photons results in FRET After ligation and denaturation (to disruptAfter ligation and denaturation (to disrupt unligatedunligated dsRNAdsRNA),), FRET emission is measured byFRET emission is measured by spectrophotometryspectrophotometry:: In the presence of active REL1, ligation occurs, resulting in denaturation-resistant FRET In the absence of active REL1, the dsRNA dissociates with denaturation, abrogating FRET Thus, inhibitors of REL1 will abrogate FRET as compared with a control '' (( ! #! # RNA oligos annealed resolved on aRNA oligos annealed resolved on a 20%20% acrylamideacrylamide/ 5%/ 5% glycerol /1 x TBE nonglycerol /1 x TBE non--denaturing geldenaturing gel 55’’ and 3and 3’’ RNA substrates annealed to bridge show efficientRNA substrates annealed to bridge show efficient FRETFRET Ligation by REL1 results in denaturationLigation by REL1 results in denaturation--resistant FRETresistant FRET Absence ofAbsence of ligationligation abrogates FRETabrogates FRET Furthermore, absence of theFurthermore, absence of the gRNAgRNA bridge precludes annealingbridge precludes annealing and thus FRETand thus FRET )) ** ++ A)A) rREL1rREL1 InductionInduction B)B) Assay conditions (buffer pH)Assay conditions (buffer pH) Induction of soluble protein improved by optimising [ITPG]Induction of soluble protein improved by optimising [ITPG] Induction of soluble protein further improved by inclusion ofInduction of soluble protein further improved by inclusion of heat shock prior to IPTG facilitating correct protein foldingheat shock prior to IPTG facilitating correct protein folding Ligation and annealing efficiency optimised with respect toLigation and annealing efficiency optimised with respect to assay buffer pHassay buffer pH pH 8.0pH 8.0 most conducive to annealing and ligationmost conducive to annealing and ligation ,, -- '' ! !! ! # $ %# $ % '( $'( $ RNA editing by uridine insertion/deletion is essential for mitochondrial gene expression in all trypanosomatids pathogens. Editing is catalyzed by multiprotein complexes, the editosomes. A key catalytic component of editosomes is RNA editing ligase 1 (REL1). REL1 has been validated as a drug target in T. brucei in vitro and in vivo (Schnaufer et al., Science 2001). The 1.2 crystal structure of REL1 revealed a deep ATP binding pocket with numerous opportunities for specific binding of small molecule inhibitors (Deng, Schnaufer et al., J Mol Biol 2004). There are no close REL1 homologs in the mammalian host and thus this enzyme represents a target of potentially high specificity. Computational screening efforts based on molecular dynamics simulations resulted in the identification of several REL1 inhibitors with IC50 values in the single-digit µM range (Amaro, Schnaufer et al., PNAS 2008; Durrant, Hall et al., PLoS NTD 2010). Here we describe the development of a high throughput screening (HTS) compatible, fluorescence-based REL1 assay. The assay was optimised with respect to reaction chemistry; fluorophores; rREL 1 production enzyme kinetics culminating in higher sensitivity; better dynamic range higher throughput .. /0/0 Assay statistically validated as suitable forAssay statistically validated as suitable for high throughput screeninghigh throughput screening Utilising two independent methods of calculating the amount ofUtilising two independent methods of calculating the amount of ligatedligated FRET product theFRET product the [ATP] substrate corresponding to Km[ATP] substrate corresponding to Km = 2.5= 2.5µµMM Km = [ substrate ] @Km = [ substrate ] @ ½½ x V maxx V max B)B) Adapting reaction conditions for actual high throughput screenAdapting reaction conditions for actual high throughput screen A)A) Determination of [ATP] substrate concentration @ Km (=Determination of [ATP] substrate concentration @ Km (= ½½ xx VmaxVmax)) A compoundA compound independently validated as possessing REL 1independently validated as possessing REL 1 inhibitory properties demonstrates similarinhibitory properties demonstrates similar ligationligation inhibition ininhibition in the context of this assaythe context of this assay Furthermore, this can be demonstrated byFurthermore, this can be demonstrated by non denaturing PAGEnon denaturing PAGE To economise on use of L1 and determineTo economise on use of L1 and determine ligationligation time yieldingtime yielding linear order conditionslinear order conditions ligationsligations were performed @ variouswere performed @ various [ATP]. In conclusion it was found that:[ATP]. In conclusion it was found that: 5 minute5 minute ligationsligations using 50ng of L1 @ 10using 50ng of L1 @ 10µµM ATP yielded rapidM ATP yielded rapid sensitive reactions compatible with linear order conditions, i.esensitive reactions compatible with linear order conditions, i.e.. affording dynamic range for determining putative inhibition ofaffording dynamic range for determining putative inhibition of rRELrREL 1 by small compound1 by small compound ligandsligands 11 (( Plus L1 Minus L1 0 5000 10000 15000 20000 25000 30000 35000 40000 45000 50000 55000 60000 65000 70000 75000 80000 85000 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 2 !3!# Z = 1 - ( 3 s + 3 c) (µs - µc) Z = 1 - ( 16397 + 1079) (70119 - 5909) Z = 1 - 0.3 = 0.7 !!! 45 6 -7 50ng #003-1 + 10µM ATP - L1 + 10 min 0 5000 10000 15000 20000 25000 30000 35000 40000 45000 50000 55000 60000 65000 70000 0 5 10 15 20 25 30 35 40 45 50 55 60 65 70 75 80 85 8 !3!# Gain = 1453 C)C) Verification of standard reaction kineticsVerification of standard reaction kinetics As depicted there is a clear correlation between reactionAs depicted there is a clear correlation between reaction kinetics underkinetics under standard operating conditionsstandard operating conditions and FRET signaland FRET signal evinced fromevinced from non denaturing PAGEnon denaturing PAGE Oligo markers ! # ! # 55’’ # 2# 2 33’’ # 1# 1 DualDual 200ng 2000ng - L1 - gR#1 Den: 200x #139 ND 200ng 2000ng - L1 - gR#1 (+ 200ng) (+ 200ng) 99 // ! #! # 0 10 20 30 40 50 60 70 80 90 100 L5 D L30 D Min L1 D Min gR#1 D L5 ND L30 ND Min L1 ND Min gR#1 ND :9; Gel #11_Cy5 Densitometry #114:FRET Signal magnitude by FRET photometry compared with (gel) Cy5Signal magnitude by FRET photometry compared with (gel) Cy5 densitometrydensitometry There is aThere is a manifest and unambiguous relationship betweenmanifest and unambiguous relationship between signal magnitude associated with gel products and those on 96signal magnitude associated with gel products and those on 96 well plateswell plates No L1 0uM ATP 2.5uM ATP 5uM ATP 10uM ATP 0.0 10000.0 20000.0 30000.0 40000.0 50000.0 60000.0 70000.0 80000.0 0.0 5.0 10.0 15.0 20.0 25.0 30.0 35.0 40.0 45.0 !3 Gain = 1453 (1)(1) (1)(1) (2)(2) (2)(2) 0.5 1.0 1.5 2.0 0 20 40 60 80 100 0 2 1';.)= µµµµ8 :8* IC50 = 9.3µM +/- 0.673µM R2 = 0.9904 0 10000 20000 30000 40000 50000 60000 70000 Tris 3.5 Tris 5.0 Tris 5.5 Tris 6.0 Tris 6.5 Tris 7.0 Hepes 8.0 ; + 4 ;: ? !3 Plus 50ng #002-5 Minus L1 Gain = 1531 9-) %-' .-1 )-9 %-% 1-@ %- + L1/ - L1 S/BS/B View publication statsView publication stats
