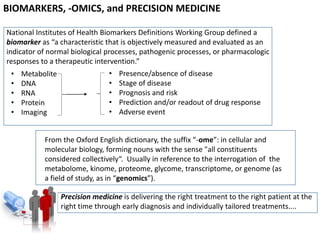
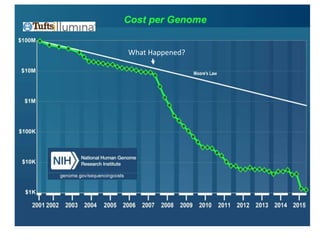
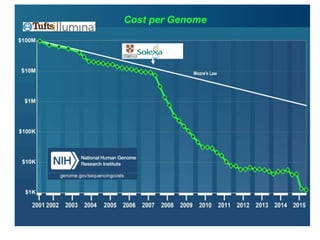
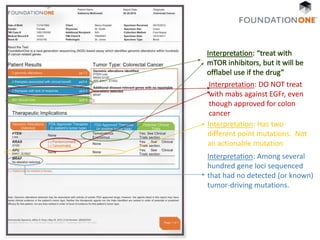
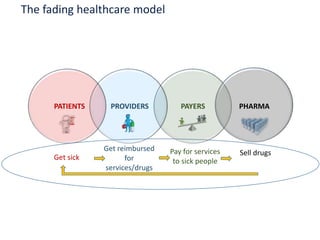
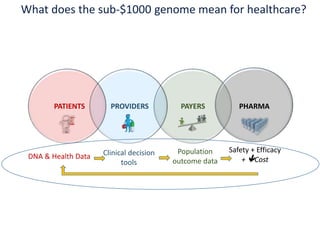
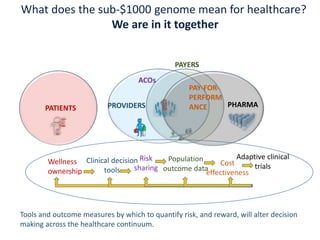
The document discusses the intersection of precision medicine, biomarkers, and healthcare policy. It describes how biomarkers and -omics data can be used for precision medicine to improve diagnostic accuracy, deliver targeted therapies, and stratify patient populations. However, clinical validation of biomarkers now requires large datasets and years of studies due to regulatory and payer requirements. This has reduced incentives for diagnostic innovation. The document also discusses challenges around clinical interpretation of complex multi-omic tests, evolving medical training and workflows, and disconnects between patent and reimbursement policies.