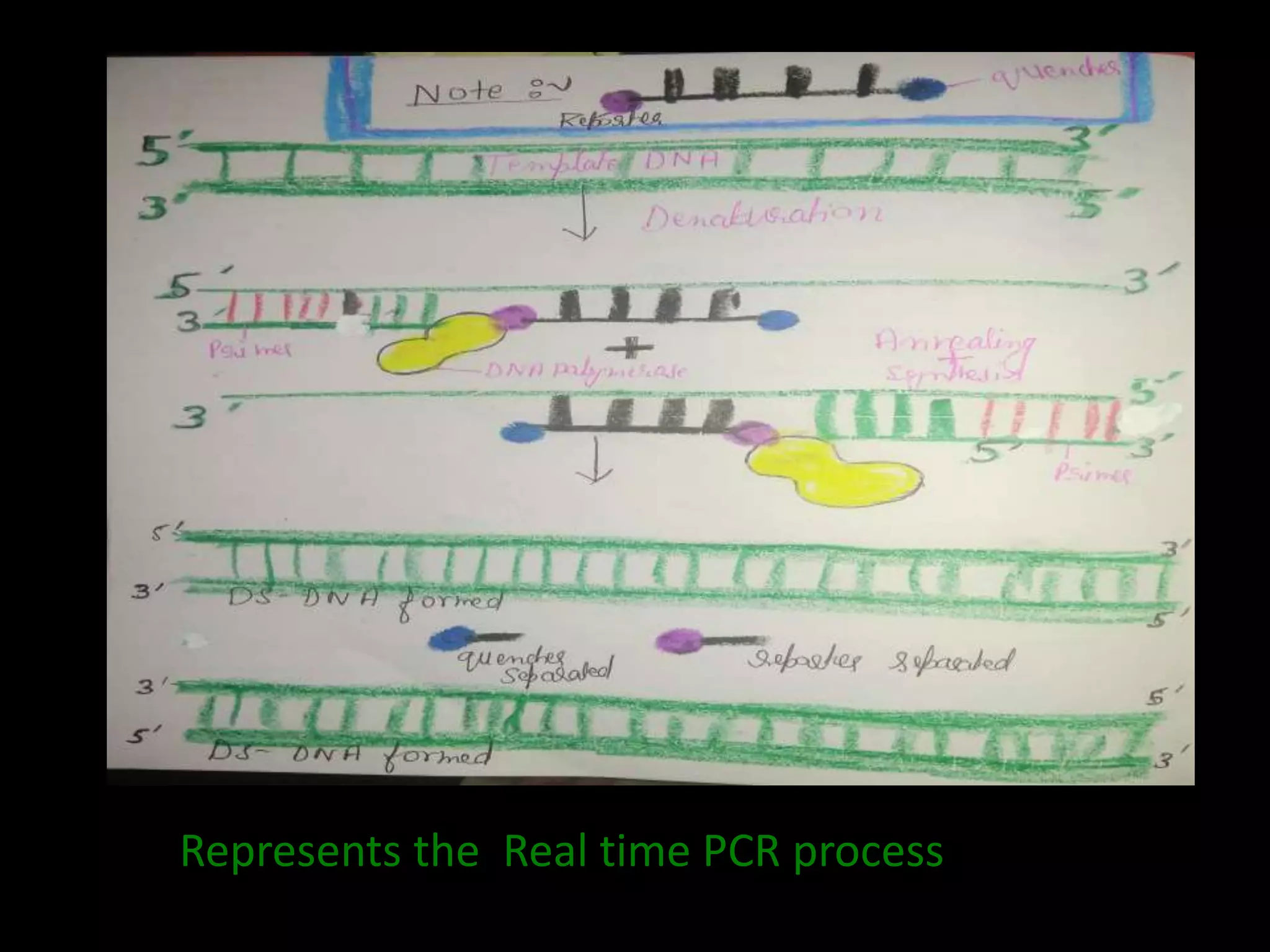
The document presents an overview of Real-Time PCR (RT-PCR), discussing its various types, applications, and advantages over traditional PCR methods. RT-PCR offers rapid, precise, and quantitative results by measuring PCR amplification during the early phases of the reaction. It highlights the significance of components like SYBR Green dye and TaqMan probes in fluorescence detection and discusses the advantages of high sensitivity, dynamic range, and resolution in RT-PCR.