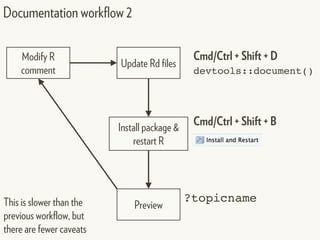
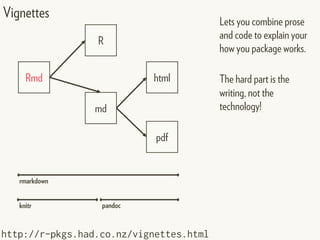
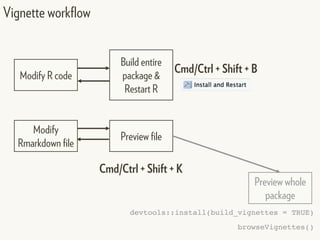
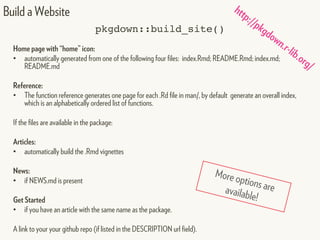
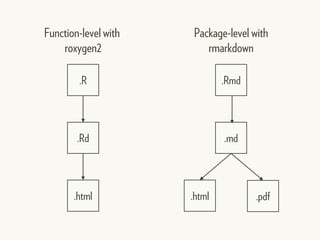
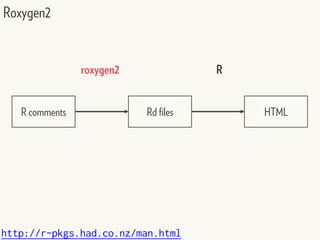
This document provides an overview of creating package documentation using roxygen2 and R markdown. It discusses writing function documentation with roxygen2 comments, previewing documentation locally, and creating package documentation like vignettes using R markdown. The document demonstrates the documentation workflow and encourages documenting other objects like data and classes. It also introduces creating package websites using pkgdown to showcase the package documentation.


.
* This is a list
* This is another item
```R
# Some R code
f <- function() x + 1
```
## This is a secondary heading
You can also do `inline code`, numbered lists and quotes and
more.
Basic markdown formatting](https://image.slidesharecdn.com/rpackagedevelopmentcreatepackagedocumentation-isabellagollini-181126113617/85/R-package-development-create-package-documentation-isabella-gollini-4-320.jpg)





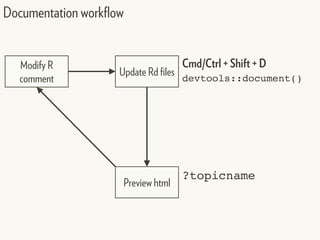

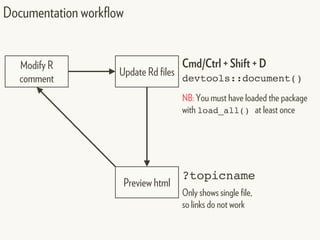
![[hadcol]
Change working directory/project to:
https://github.com/forwards/hadcol](https://image.slidesharecdn.com/rpackagedevelopmentcreatepackagedocumentation-isabellagollini-181126113617/85/R-package-development-create-package-documentation-isabella-gollini-15-320.jpg)
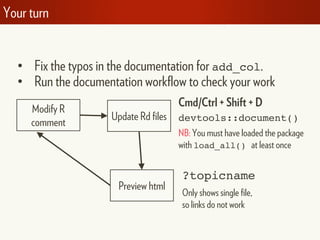



![# Activate by running
# use_roxygen_md()
**bold**, _italic_, `code`
* [func()]
* [pkg::func()]
* [link text][func()]
* [link text][pkg::func()]
Use markdown for formatting](https://image.slidesharecdn.com/rpackagedevelopmentcreatepackagedocumentation-isabellagollini-181126113617/85/R-package-development-create-package-documentation-isabella-gollini-22-320.jpg)