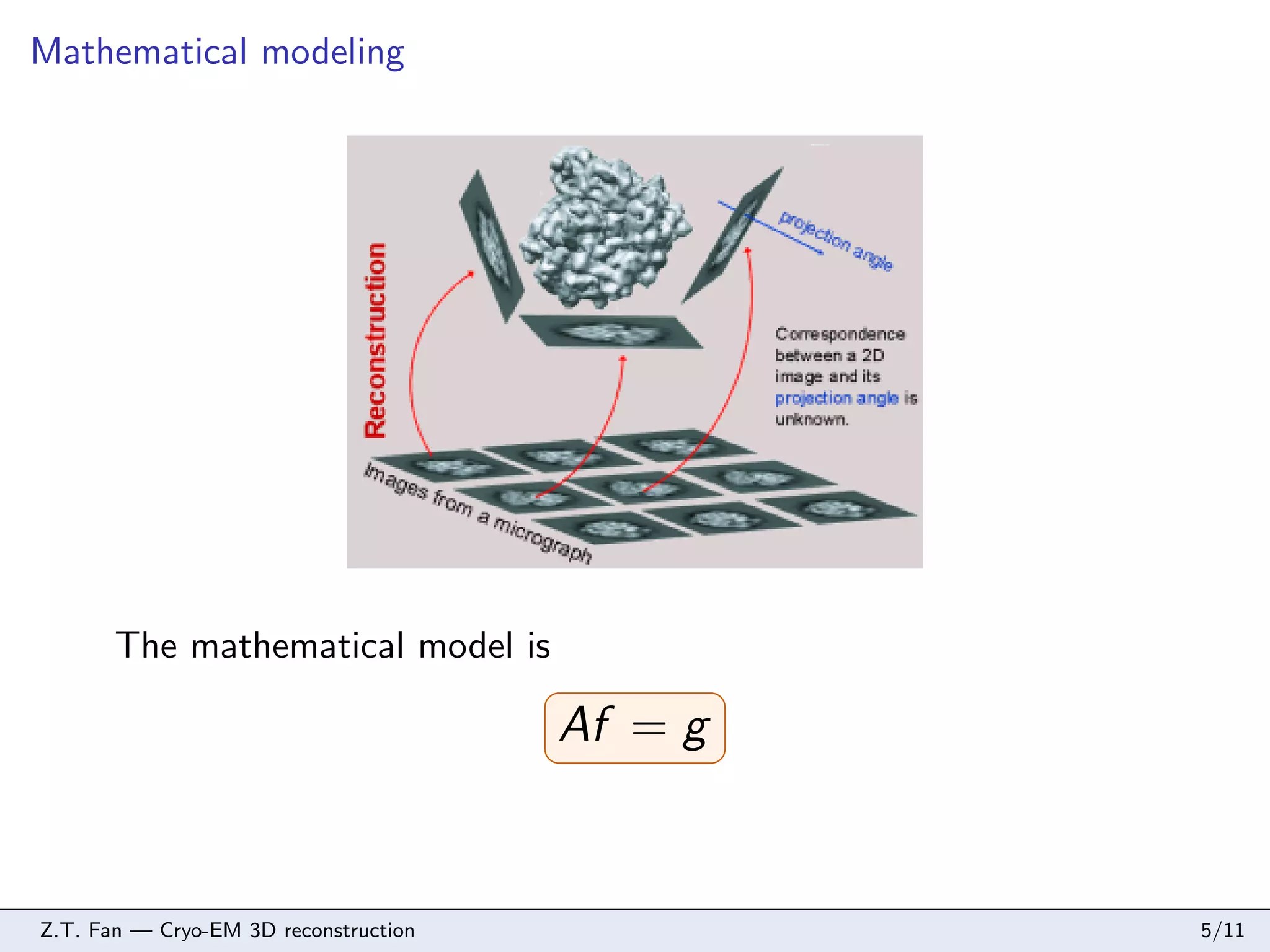
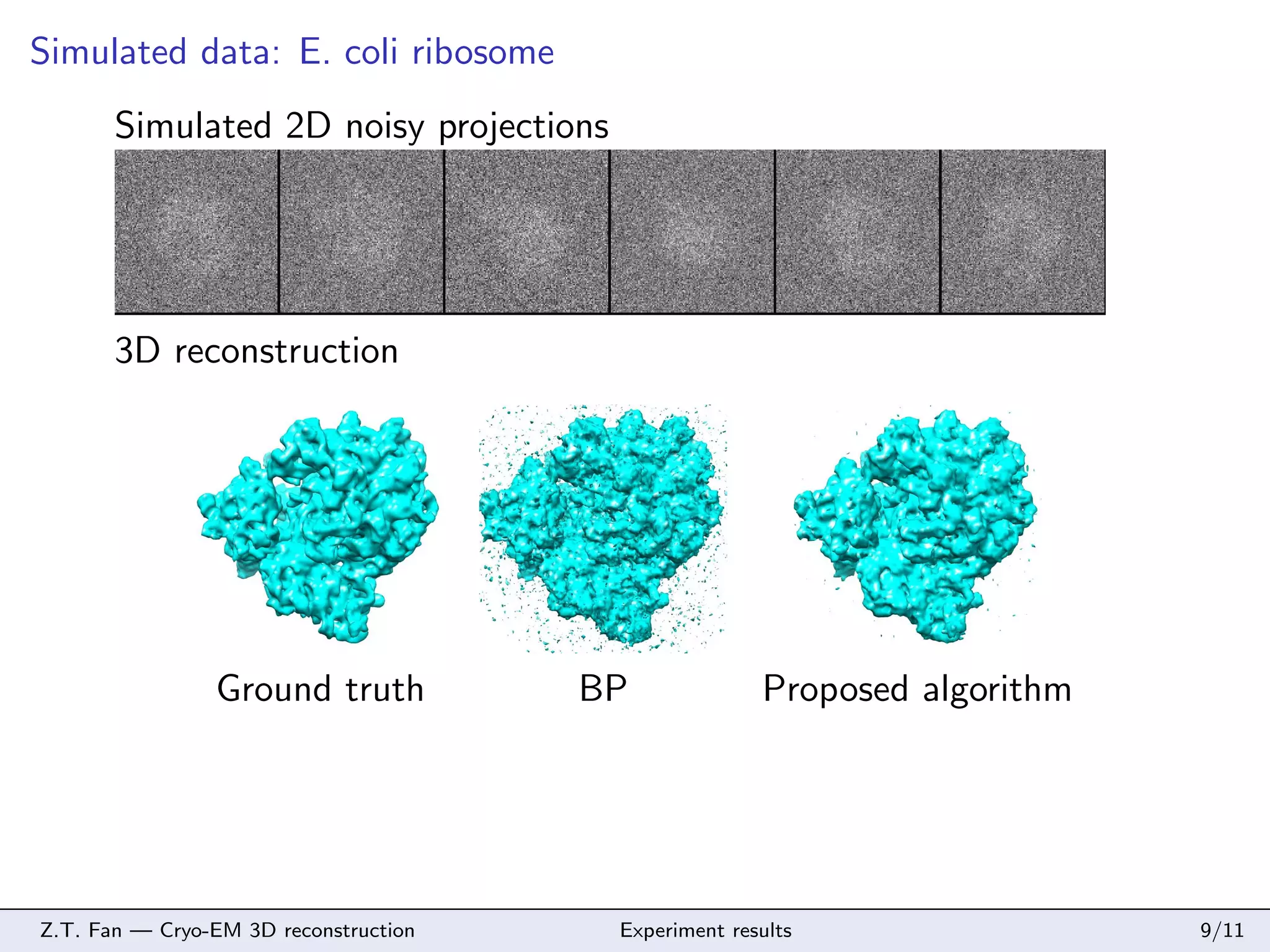
This document discusses cryo-electron microscopy (cryo-EM) 3D reconstruction techniques. It describes the cryo-EM imaging process and challenges in reconstructing 3D structures from 2D projection images, including large noise and data size. The document proposes a memory-saving algorithm using tight wavelet frames for cryo-EM 3D reconstruction that formulates the reconstruction as a sparse representation problem solved with soft-thresholding and gradient descent. Simulation results on an E. coli ribosome and experimental results on an adenovirus demonstrate the proposed algorithm can reconstruct 3D structures from noisy projection data.