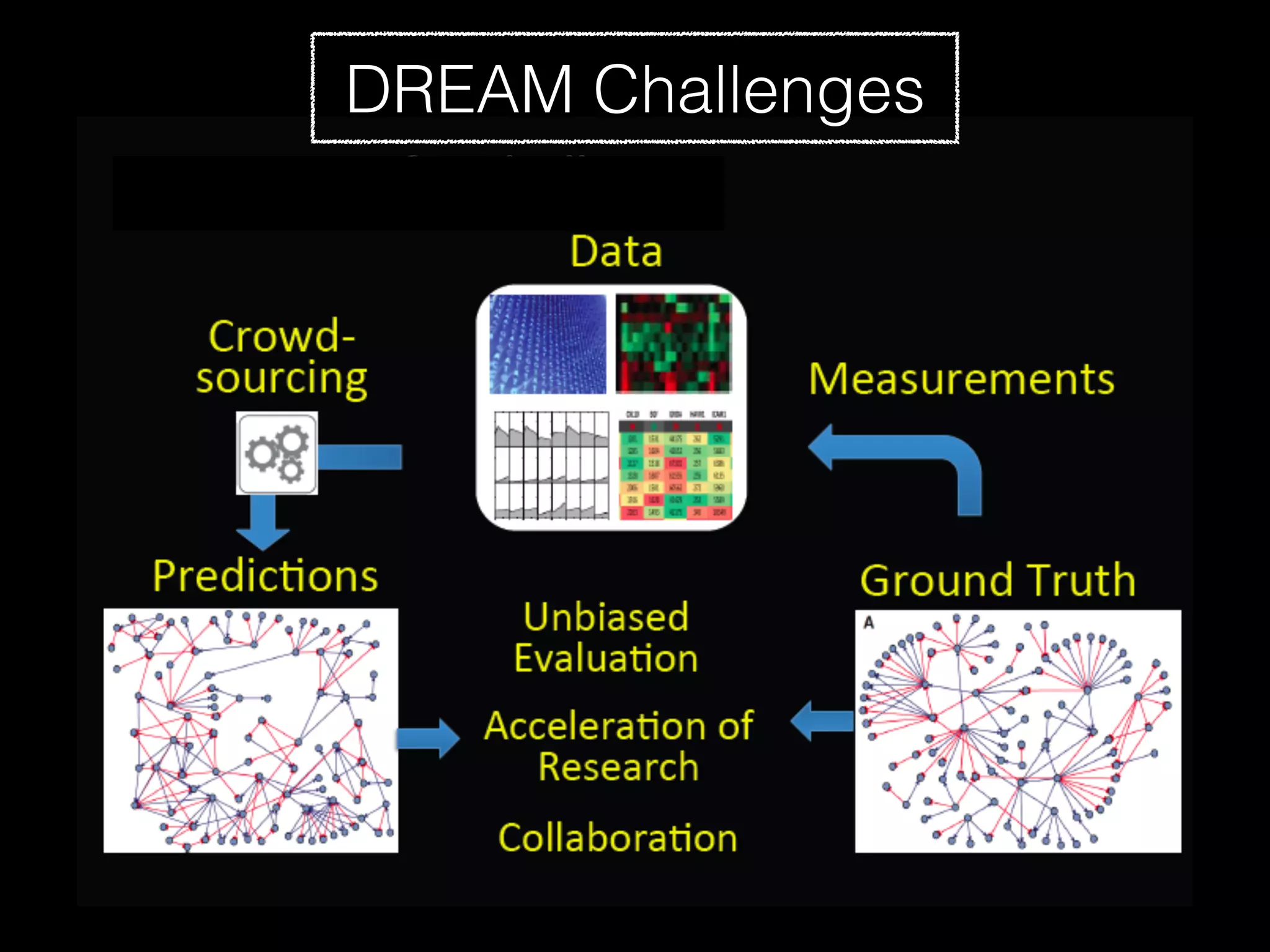
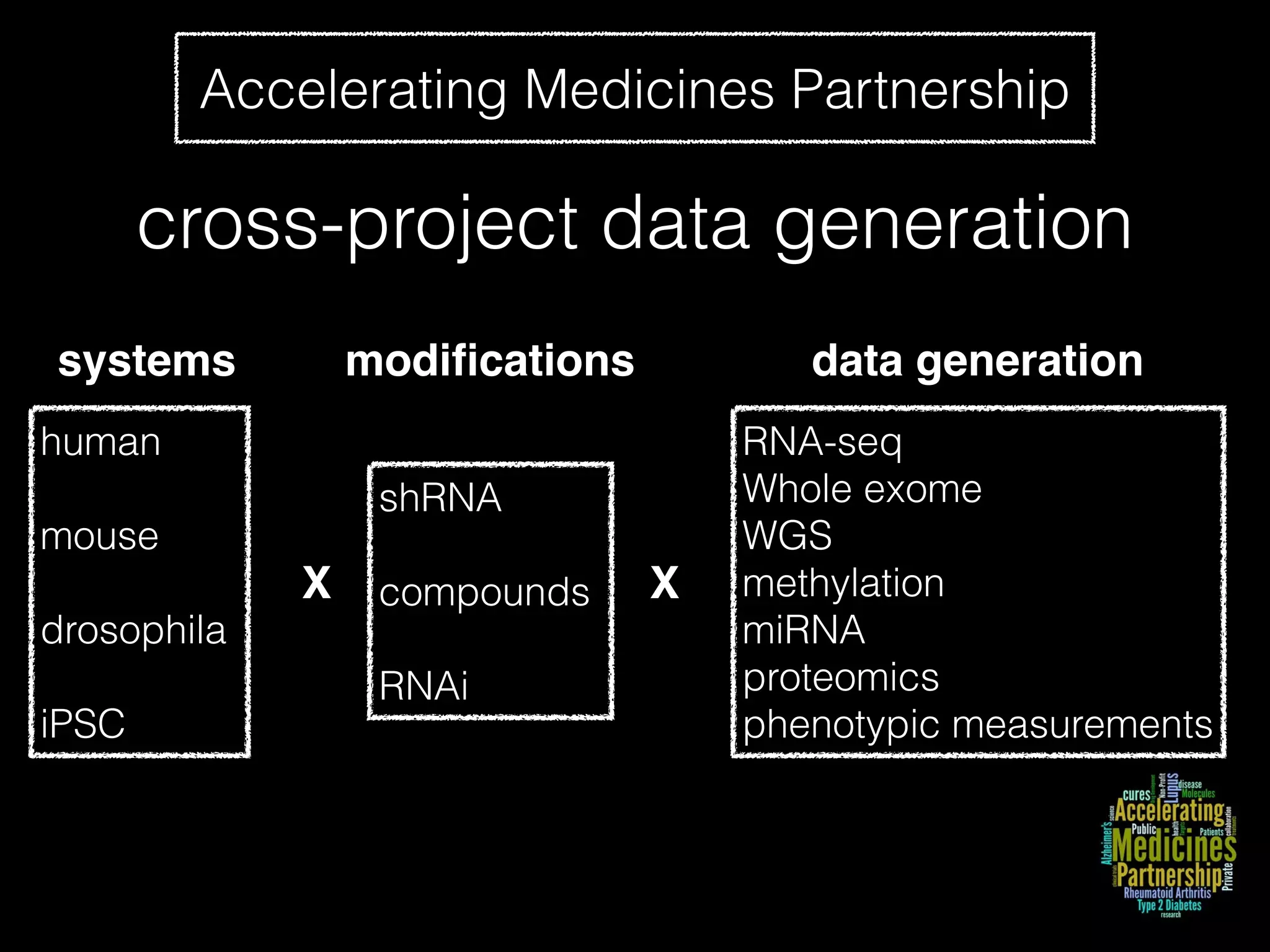
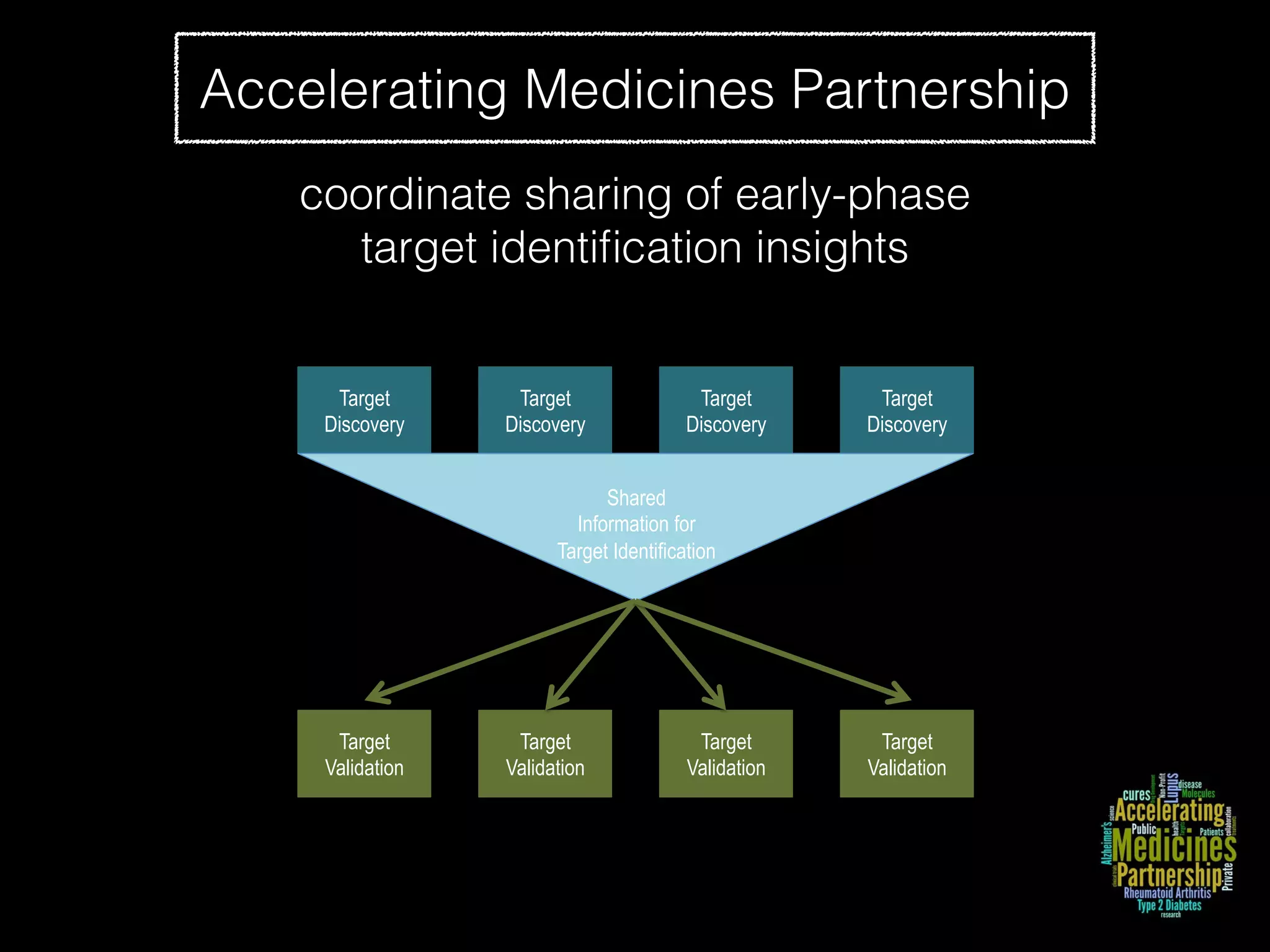
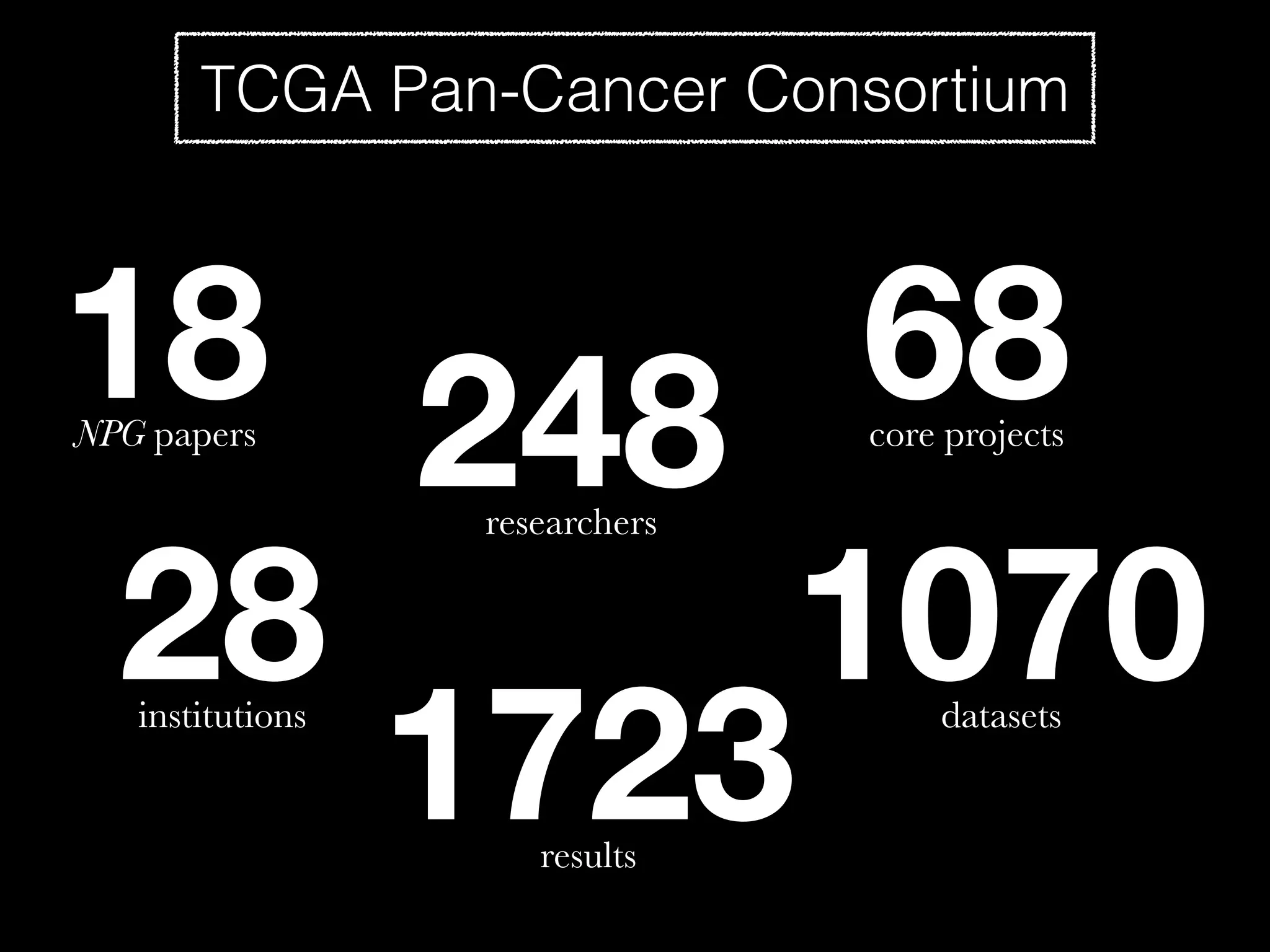
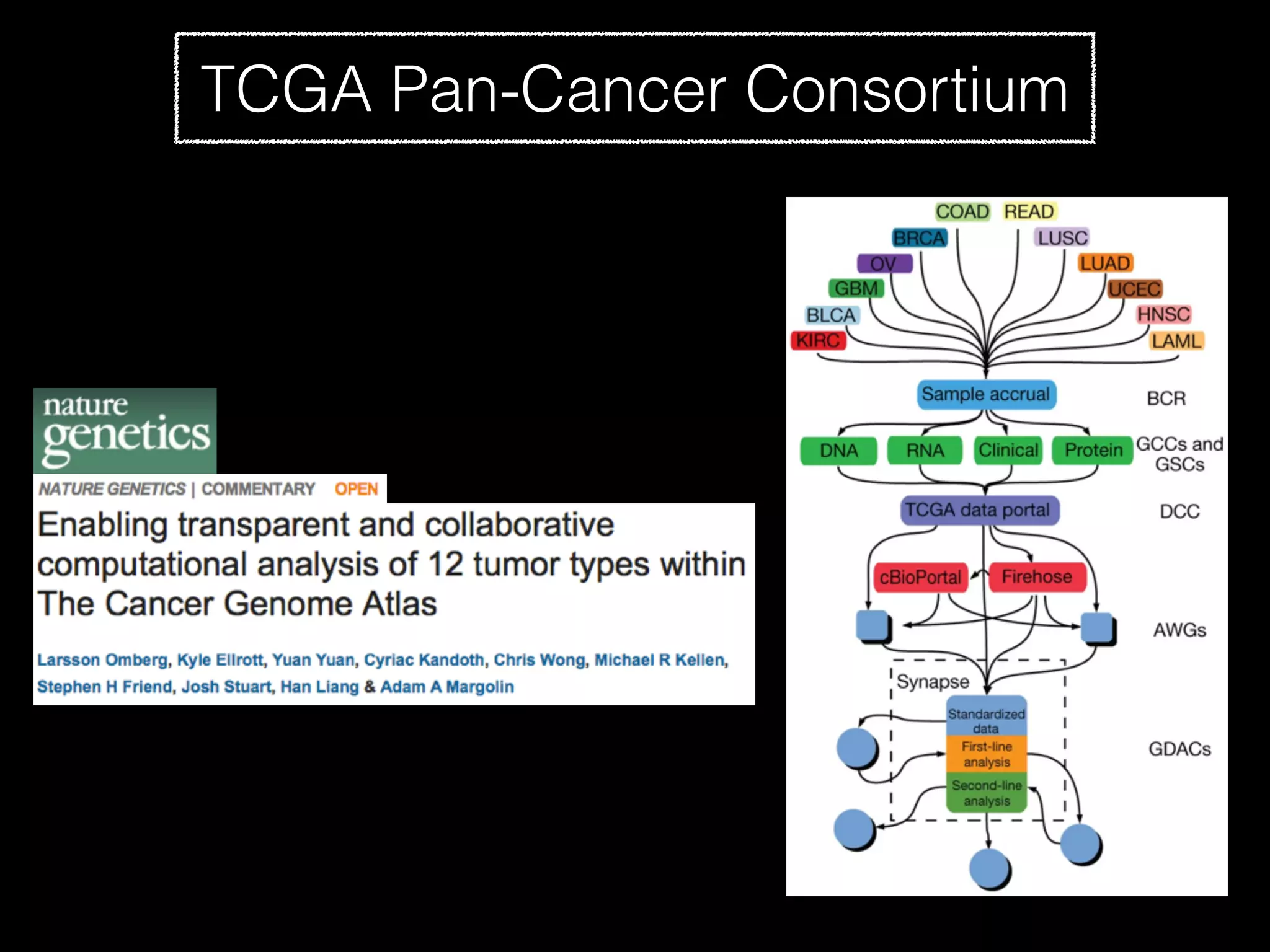
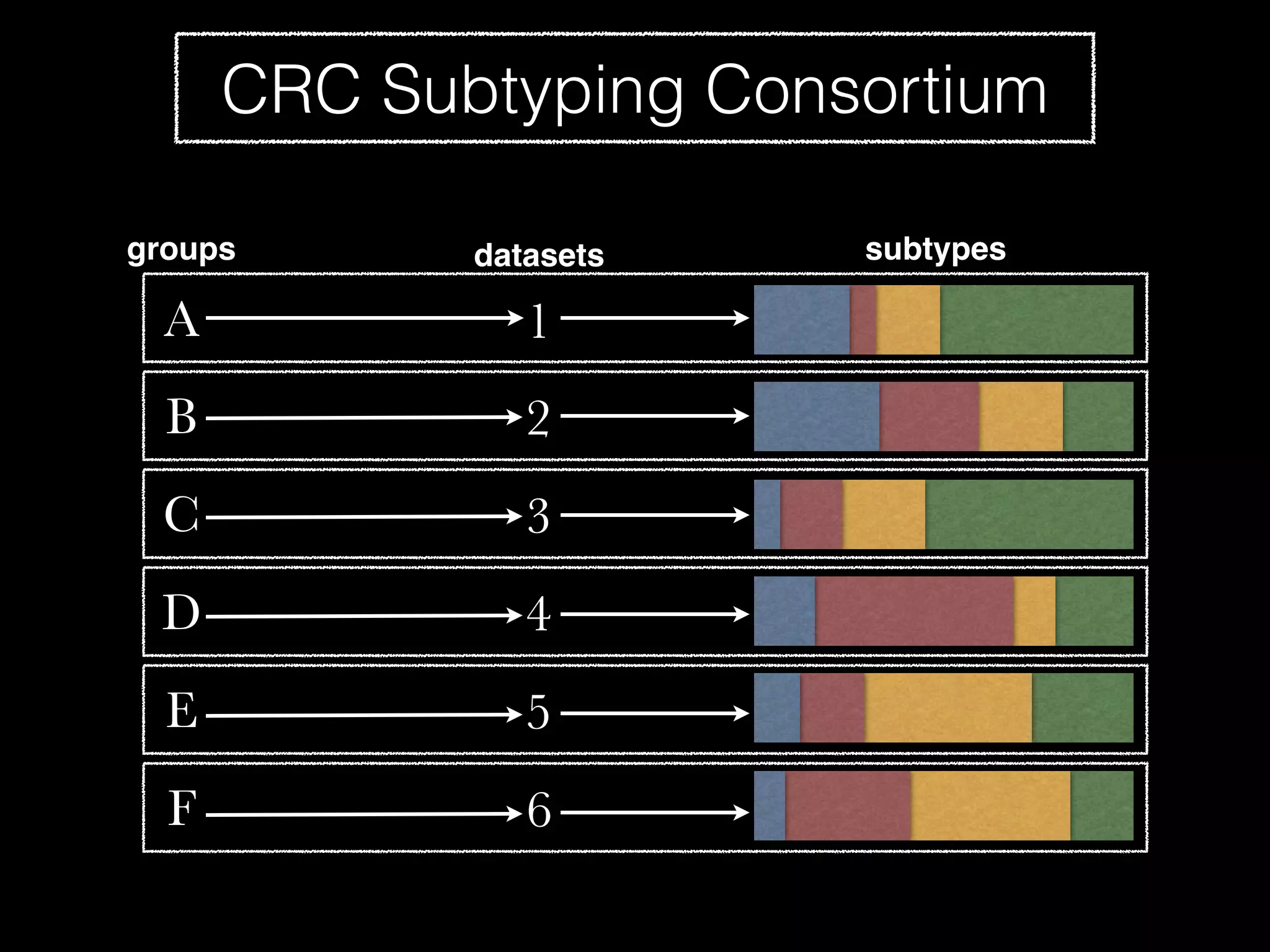
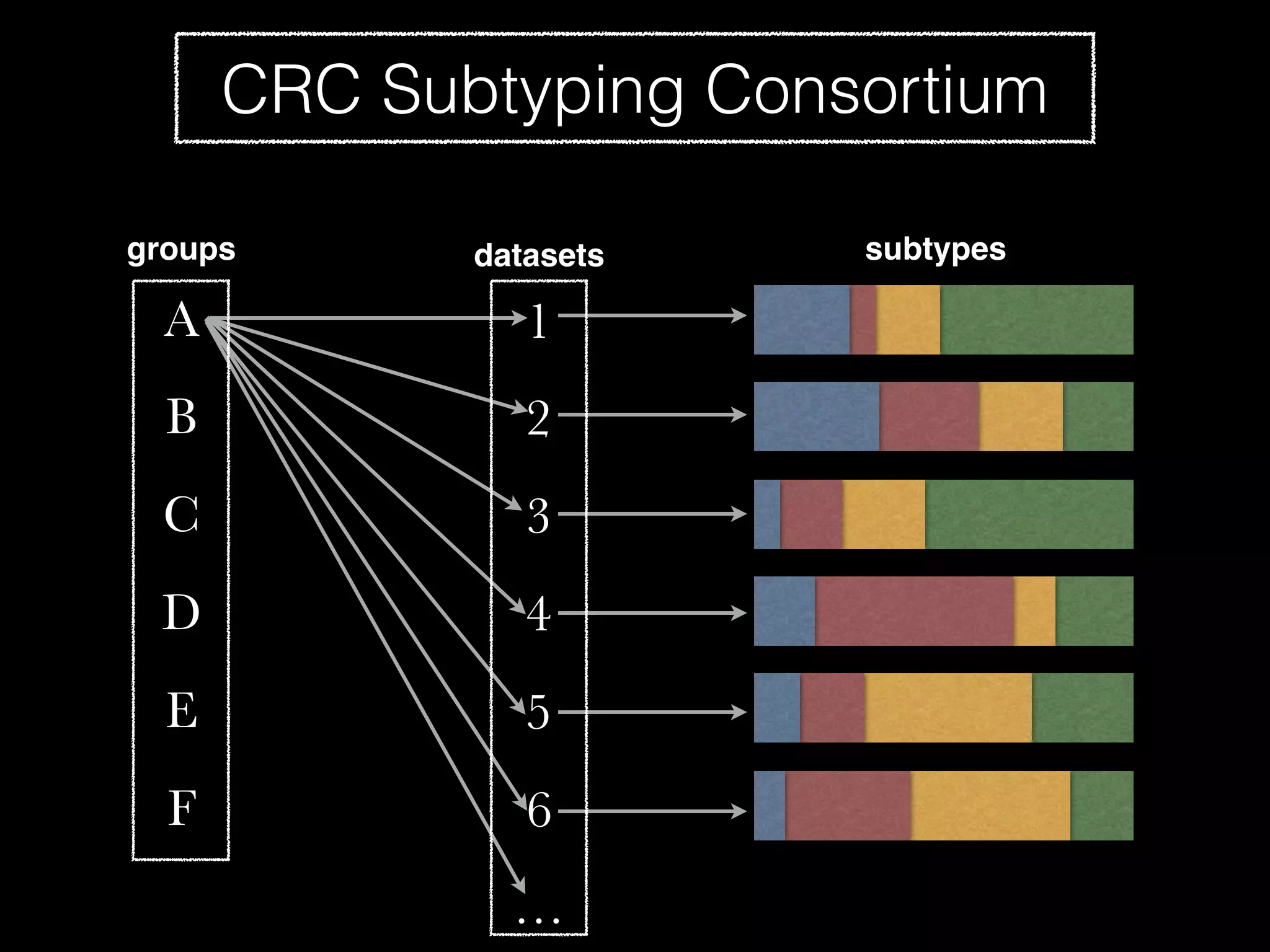
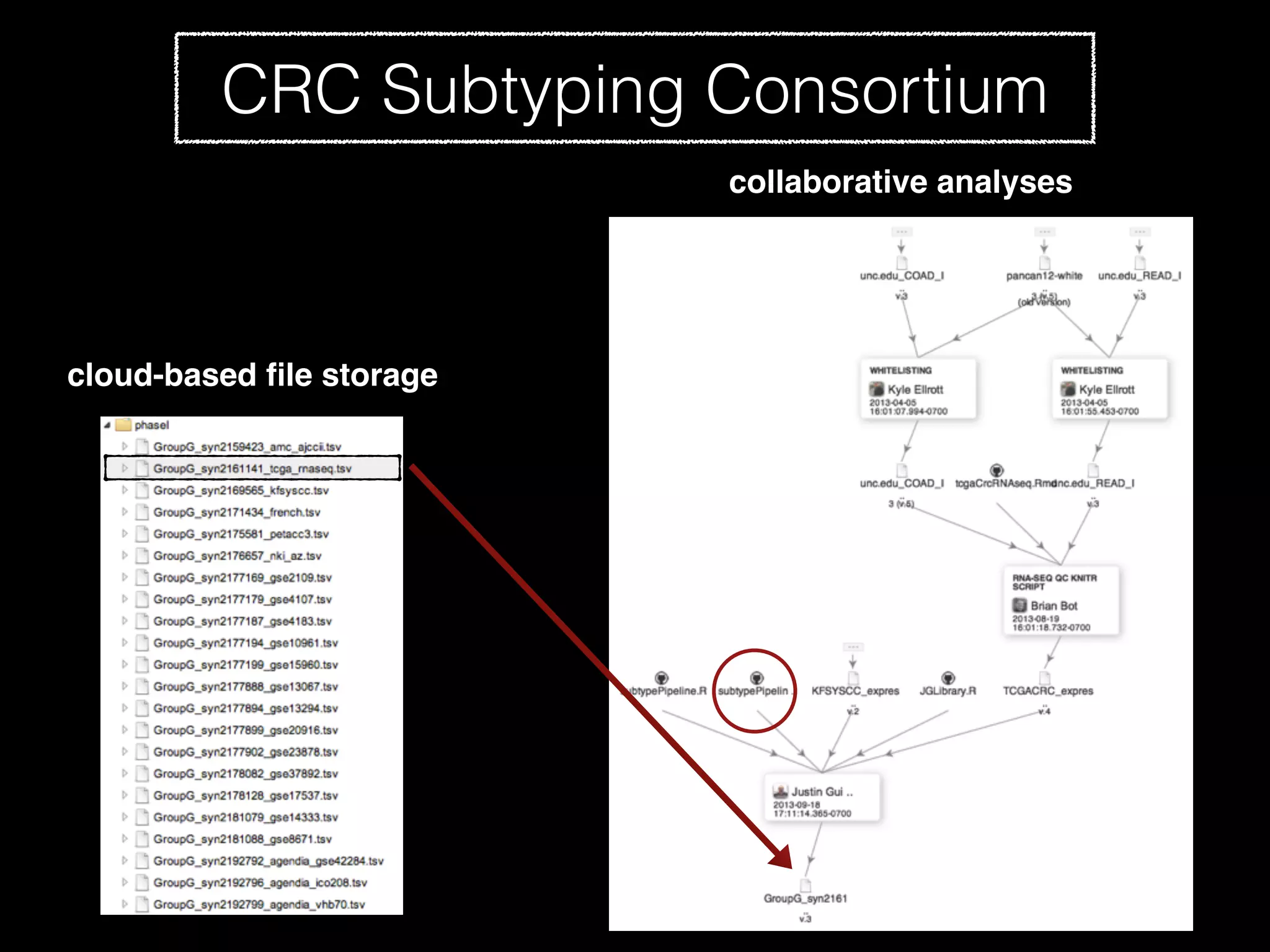
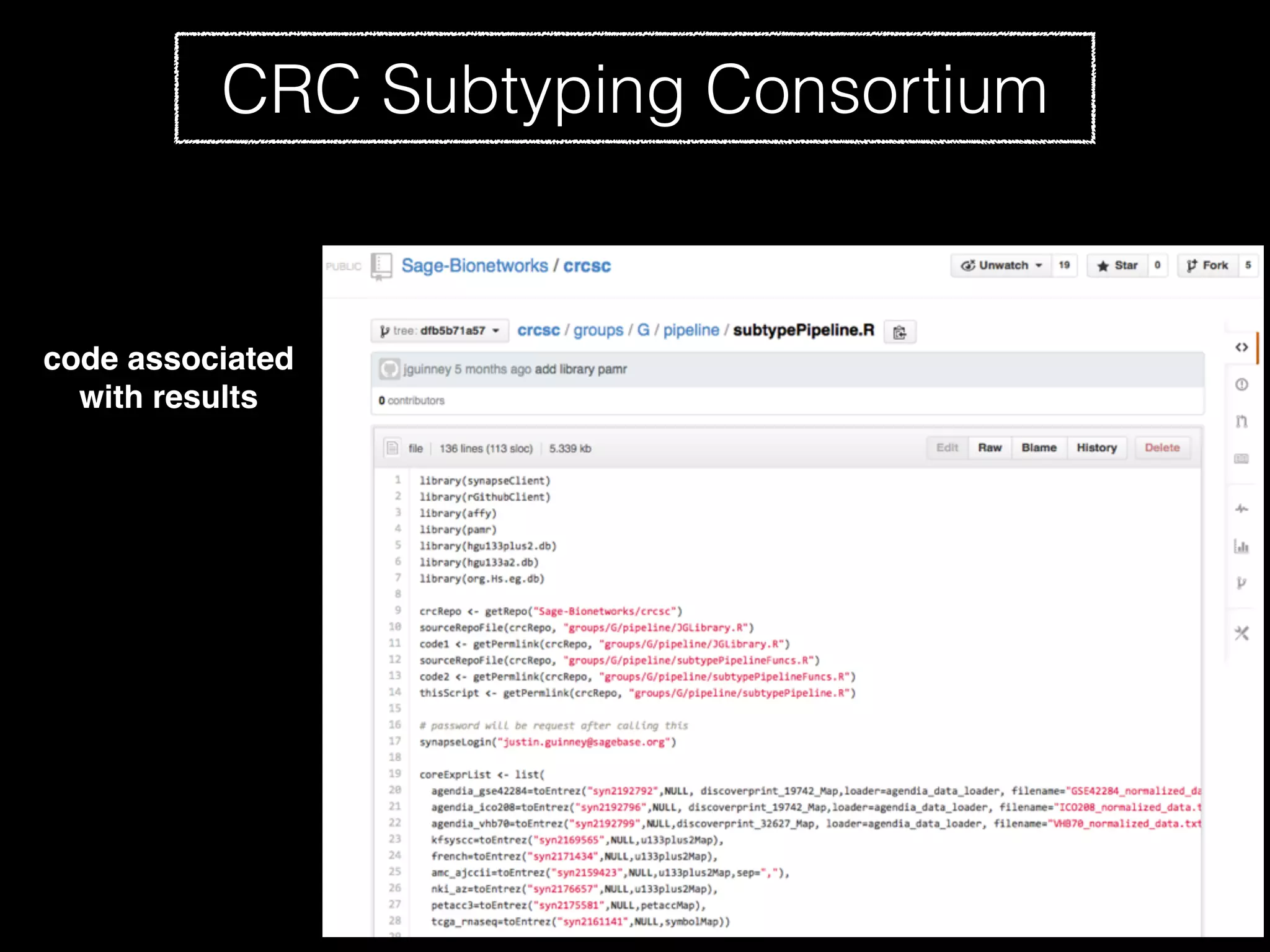
Sage Bionetworks is a non-profit organization that aims to make biomedical research more open and collaborative. It develops the Synapse platform to enable sharing of data, code, results and communication among researchers. Sage supports various research communities and projects through Synapse, including the CRC Subtyping Consortium, TCGA Pan-Cancer Consortium and Accelerating Medicines Partnership. These projects involve collaborative analysis of complex datasets toward goals like cancer subtyping and target identification for diseases.